Grantee Research Project Results
2021 Progress Report: Multiplexed human BrainSphere Developmental Neurotoxicity Test for Six Key Events of Neural Development
EPA Grant Number: R839505Title: Multiplexed human BrainSphere Developmental Neurotoxicity Test for Six Key Events of Neural Development
Investigators: Lena, Smirnova , Thomas, Hartung , David, Gracias , Cynthia, Berlinicke
Institution: The Johns Hopkins University
EPA Project Officer: Callan, Richard
Project Period: August 1, 2019 through July 31, 2022 (Extended to July 31, 2024)
Project Period Covered by this Report: August 1, 2020 through July 31,2021
Project Amount: $849,277
RFA: Advancing Actionable Alternatives to Vertebrate Animal Testing for Chemical Safety Assessment (2018) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
- Design an iPS cell line with fluorescently tagged fusion glial fibrillary acidic protein (GFAP), proteolipid protein 1 (PLP), synaptophysin (SYP), and b-III-tubulin using CRISPR/CAS9 gene-editing technology.
- Generate 3D multi-fluorescent BrainSpheres from the iPSC line created in objective 1 and characterize them in terms of neural differentiation and fusion-protein fidelity.
- Expose the multi-fluorescent BrainSpheres to five chemicals for the 6-in-1 BrainSphere assay optimization; establish the 3D electrophysiological recording and high content imaging (HCI) workflow.
- Establish test strategy workflow for the toxicity screening using the 6-in-1 BrainSphere assay and test further 30 compounds. Develop SOPs
Progress Summary:
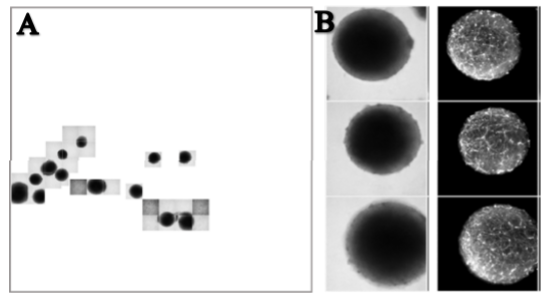
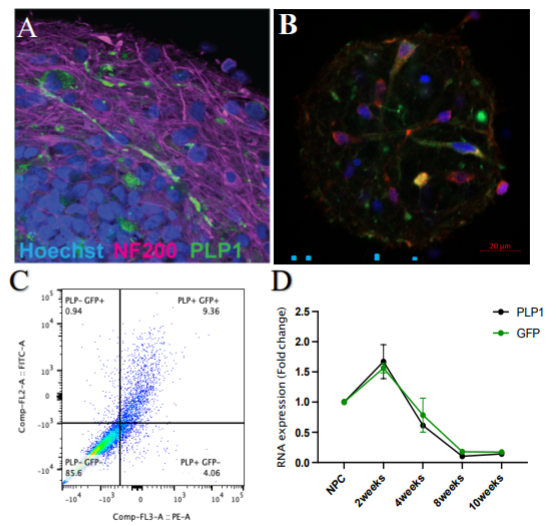
Objective 1-2: This year, we continued to characterize and validate the PLP-GFP clones in both cell lines (NIBSC8 and CS2PFYiCTR-nx.x). Six clones were selected for each cell line to perform screening. After chromosomal aberration screening, sequencing, and evaluation of off-target effects, we continued with three clones for NIBSC8 and 2 for CS2PFYiCTR-nx.x. These clones were differentiated into BrainSpheres and immunofluorescence and flow cytometry analysis showed the presence of the green tag in oligodendrocytes (Fig 1A, B, C). We observed colocalization of PLP-GFP and oligodendrocyte marker, MBP (Fig 1B). Gene expression showed a strong correlation between PLP1 and GFP levels at different time points (2, 4, 6, and 8 weeks Fig1D). This data further shows specificity of the inserted tag to the PLP1 gene. We established a standard operating procedure of the transfection and QC steps (manuscript in preparation). After selection of one PLP1-GFP clone in each cell line we proceeded to the next marker - Synaptophysin (editing SYN-BFP and SYN-GFP). First transfections failed since initial gRNA selected for SYP was not efficient and we had to re-design and re-clone the gRNA vector. The new gRNA was very efficient, and we were able to incorporate SYP-BFP in NIBSC8 containing PLP-GFP as well as SYP-GFP in NIBSC8 Wildtype. Validation and QC of the clones has been done: the selected clones had homozygous insertion of fluorescent tag and do not have chromosomal aberrations or off-target effects. We differentiated SYN-GFP and PLP1-GFP/SYN-BFP iPSC into BrainSpheres to assay tag expression similarly as we did for PLP1. In parallel upon transfection of iPSC with CRISPR/Cas9 system targeting GFAP and TUBB3 genes, we observed also a very low transfection efficiency and had to redesign and re-clone gRNAs (2 per gene). CS2PFYiCTR cell line containing PLP1-GFP tag was then transfected with TUBB3-RFP. The new gRNA for TUBB3 performed very efficiently and we are currently growing colonies from single cells to validate and QC positive clones.
Figure 1: Characterization of PLP-GFP expression in the BrainSpheres. A. Endogenous expression of GFP (green) and axons stained with neuronal marker NF200 (purple), nuclei (blue). B. Co-localization of endogenous PLP (green) and oligodendrocyte marker, MBP (red). C. Flow cytometry dot plot showing colocalization between endogenous GFP expression and PLP1+ cells. D. PLP1 and GFP gene expression during BrainSpheres development (NPC – neuroprogenitor cells).
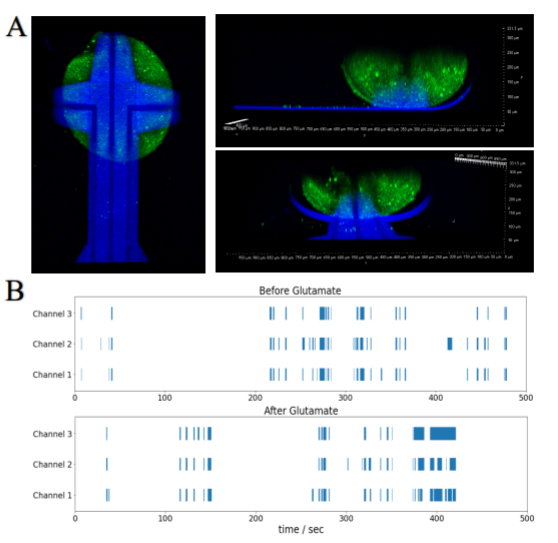
Objective 3 Part 1: This year, we focused on characterization of BrainSpheres captured in the self-folding shell electrodes. Live and “healthy” BrainSphere organoids enveloped inside shell electrodes, and the transparency of the micro 3D devices exhibit the potential for future optical stimulation (Fig 2A). Successful recordings of field potential of the enveloped brain organoids inside the 3D shell electrodes were achieved with the electrodes on the 3 leaflets of the device. First, we recorded the activity of brain organoids on our shell electrodes in their 2D format. We noticed the organoid grew spread on the 2D surface instead of maintaining its 3D architecture when seeded on 2D MEAs. For 3D MEAs, the BrainSpheres were seeded to the center of the shell electrodes before self-folding, then enveloped within the leaflets, thus a shell was formed. 3 days later, we observed the firing of the BrainSpheres with the 3D shell electrodes. We successfully recorded 3D field potentials for 4 weeks, with the 3D spatial electrophysiology activities recorded across each leaflet, collecting 3D spatiotemporal field potential information of the BrainSpheres. We stimulated BrainSpheres captured in the 3D MEA shell with 50 μM glutamate as an excitatory chemical stimulus. The raster plot (Fig. 2B) before and during the glutamate stimulus showed enhanced electrical bursting during stimulus.
Figure 2: 3D electrophysiological recording. A. Confocal images of a BrainSphere enveloped inside shell electrodes, Green: Ca2+ Imaging (Fluo-4), Blue: SU-8 shell electrodes gripper. B. Raster plot showing enhanced electrical bursting of the BrainSphere after Glutamate stimulation.
Figure 3: Preliminary scanning of the BrainSpheres. A. First pass brightfield. B. Fluorescent confocal of selected fields.
Objective 3 Part 2: This year, we have begun to establish and optimize the highcontent imaging and image analysis pipeline we will use to quantify the effects of toxicants on BrainSpheres. To establish the high-throughput imaging process, we have been utilizing a feature of the automated, fluorescence-based imaging system, Cellomics CellInsight CX7 LZR, called EurekaScan. This feature allows the identification of regions of interest within the well, being imaged based on a preliminary low-resolution scan. From the initial scan, the fields that are of interest are then re-imaged more extensively. For this project, we have used brightfield for the initial scan, allowing us to determine where in the well the BrainSpheres reside, then as a second pass we obtain fluorescent confocal stacks only of these fields (Fig. 3A-B). In this way, we can automatically image multiple BrainSpheres per condition. We have also begun to develope the algorithms by which we will quantify the various metrics that are potentially affected by toxicants. Preliminary algorithms were generated to (1) quantify the average and total length of neurite outgrowth (this was initially performed using fixed beta-III-tubulin stained BrainSpheres) and (2) identify PLP-GFP endogenous expression within a BrainSphere (this is being developed using live cell imaging of BrainSpheres generated from PLP reporter lines). We are currently using this pipeline to perform plate uniformity assays and establish a baseline of the timing and extent of expression of each of our markers as well as to determine the well-by-well variation of our different quantitative measures.
Objective 3 Part 3: We exposed BrainSpheres to model toxicants to derive preliminary data and effective concentrations to be used in the HCI high throughput screening pipeline. Compounds tested: chlorpyrifos, cuprizone, ceramide, selected flame retardants (TBBPA, TPHP, IPP, BDE-47), heavy metals and their mixture (Pb, As, Cd, Cr), PAHs.
Comparison of actual accomplishments with the goals and objectives (outputs/outcomes): Although we are still behind the original schedule with respect to cell line generation due to later hire and COVID and some further delays due to inefficient gRNA activity and lab supply shortage, we made substantial progress and are planning to finish QC and differentiation to BrainSpheres of three from four tags by the end of the 2021. Through the troubleshooting process, our lab gained substantial experience in CRISPR/Cas9 gene-editing, which is summarized in a methodological publication and SOP. We are generating two multicolor iPSC lines instead of one line as proposed originally.
Future Activities:
Finalize cell line generation and QC. Establish HCI pipeline, establish electrophysiology recording upon chemical stimuli and toxicant treatment.
Journal Articles on this Report : 1 Displayed | Download in RIS Format
| Other project views: | All 9 publications | 9 publications in selected types | All 9 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Modafferi S, Zhong X, Kleensang A, Murata Y, Fagiani F, Pamies D, Hogberg HT, Calabrese V, Lachman H, Hartung T, Smirnova L. Gene–Environment Interactions in Developmental Neurotoxicity:a Case Study of Synergy between Chlorpyrifos and CHD8 Knockout in Human BrainSpheres. Environmental Health Perspectives 2021;129(7):077001. |
R839505 (2021) |
|
Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.