Grantee Research Project Results
2020 Progress Report: Multiplexed human BrainSphere Developmental Neurotoxicity Test for Six Key Events of Neural Development
EPA Grant Number: R839505Title: Multiplexed human BrainSphere Developmental Neurotoxicity Test for Six Key Events of Neural Development
Investigators: Lena, Smirnova , Thomas, Hartung , David, Gracias , Cynthia, Berlinicke , Romero, J. Carolina
Current Investigators: Lena, Smirnova , Thomas, Hartung , David, Gracias , Cynthia, Berlinicke
Institution: The Johns Hopkins University
EPA Project Officer: Lasat, Mitch
Project Period: August 1, 2019 through July 31, 2022 (Extended to July 31, 2024)
Project Period Covered by this Report: August 1, 2019 through July 31,2020
Project Amount: $849,277
RFA: Advancing Actionable Alternatives to Vertebrate Animal Testing for Chemical Safety Assessment (2018) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
1. Design an iPS cell line with fluorescently-tagged fusion glial fibrillary acidic protein (GFAP), proteolipid protein 1 (PLP), synaptophysin (SYP), and b-III-tubulin using CRISPR/CAS9 gene-editing technology.
2. Generate 3D multi-fluorescent BrainSpheres from the iPS line created in objective 1 and characterize them in terms of neural differentiation and fusion-protein fidelity.
3. Expose the multi-fluorescent BrainSpheres to five chemicals for the 6-in-1 BrainSphere assay optimization; establish the 3D electrophysiological recording and high content imaging (HCI) workflow.
4. Establish test strategy workflow for the toxicity screening using the 6-in-1 BrainSphere assay and test further 30 compounds. Develop SOPs.
Progress Summary:
For the generation of the iPSC lines containing the four knock-in (KI) fluorescent tags for neural markers (b-III-tubulin, GFAP, SYP and PLP/MBP), we first generated the gRNA and reporter constructs (Objective 1). In order to maintain TUBB3, GFAP and MBP expression and avoid altering the expressed protein, which could affect function, we tethered together the TUBB3 and GFAP ORF and the fluorescent protein gene with a P2A self-cleaving peptide. In this configuration, both endogenous and fluorescent protein should be produced from one ORF while retaining their respective cellular localization and functionality. For PLP and SYP, we want to measure the function of the specific proteins in order to assess synapse formation and myelination, respectively, thus generated constructs to express fluorescent fusions of these proteins. We decided to generate the multi-reporters in two different cell lines (doners) instead of one as proposed originally in order to demonstrate reproducibility of the later toxicity testing: NIBSC8 (female) and CS2PFYiCTR-nx.x (male) lines are QCed, banks prepared. Both cell lines were authenticated by Short Tandem Repeat profiling and pluripotency assessed by immunofluorescence and flow cytometry. Next, we confirmed and characterized the efficient differentiation of these two lines into neuroprogenitors and subsequently into BrainSpheres. In addition, we generated preliminary data on toxicity testing using the NIBSC8 line. We exposed the BrainSpheres to the panel of test compounds (chlorpyrifos, MeHg, cuprizone, lead, cadmium, chromium, arsenic and mixture of cadmium and chromium) to identify subtoxic doses of chemicals, and performed some functional assays (neurite outgrowth, oxidative stress, mitochondrial health), which will set the concentrations to be used for imaging-based assays and electrophysiology later in the project.
The first construct available was myelin basic protein (MBP)-P2A-GFP. The homologous sequence together with the reporter gene was successfully inserted in the NIBSC-8 iPSC cell line (Fig 1A). As this was the first gene edited, we optimized the transfection conditions to increase the efficiency, which is usually quite low for iPSC. The optimzed transfection protocol will then be applied to the remaining reporters. Subsequently, three heterozygous clones were isolated and used to generate 3D BrainSpheres and verify functionality of the reporter (Fig 1B). Currently, colonies from single cells are being screened to isolate homozygous clones for the MBP-GFP reporter. Currently, we are performing cryosections and flow cytometry to demonstrate colocalization of MBP endogenous protein and GFP tag.
Next, we generated a PLP-GFP fusion construct and transfected it into two iPSC lines (Fig 1C). The optimized transfection protocol led to great transfection efficiency. Clones generated from single cells were amplified followed by DNA isolation and PCR analysis. We got a KI efficiency of about 20-30% (Fig 1D-E). All clones were banked and 6 of each cell line expanded for further characterization and validation (in progress). After selection of our best clone in each cell line, we will proceed to edit the next gene (all constructs already generated), repeat the same validation protocol and continue in the same way for the other two genes.
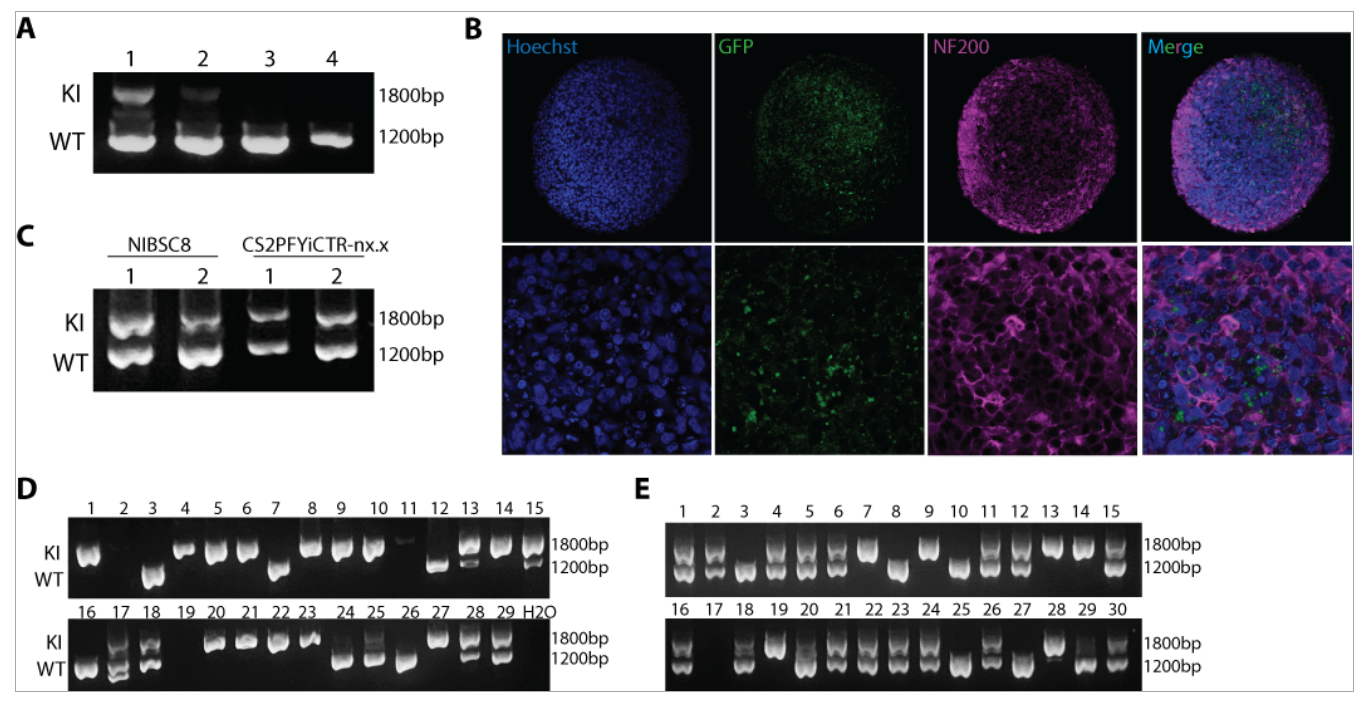
Figure 1. CRISPR/CAS9 gene-editing of MBP-GFP reporter and PLP-GFP fusion protein in iPSCs. A. Agarose gel showing KI efficiency of MBP-GFP in NIBSC8 iPSCs using four transfection protocols, with the first more efficient (DNA isolated from cell pool after transfection). B. Representative confocal images of MBP-GFP heterozygous clones showing GFP expression (green) and neurofilament protein (NF200, magenta) in 8 week-BrainsSpheres (20X and 63X magnification). C. Agarose gel showing KI efficiency of PLP-GFP in NIBSC8 and CS2PFYiCTR-nx.x iPSCs using two transfection protocols. D-E. Agarose gel showing isogenic clones for CS2PFYiCTR-nx.x and NIBSC8 iPSC, respectively. Double bands showing heterozygous insertion, while individual band at 1800 bp suggests homozygous insertion; individual 1200bp band is wild type without insertion.
With regard to the building of a 3D shell electrode microelectrode arrays (MEAs) for BrainSpheres (Part of Objective 3), we have further optimized a self-foldable MEAs gripper and achieved the controllable folding of the grippers with different size and folding angles, which allow us to catch BrainSpheres with different diameter. We sandwiched gold wires and PEDOT detecting electrodes to the gripper, the PEDOT pillars of the electrodes enable the soft touch between the BrainSphere and electrodes. At the same time, we have established a self-built electrophysiology MEA recording system and we built up the protocol to connect the MEAs grippers to recording system. In parallel, we used traditional 2D MEAs to perform functional analysis of the neural differentiation of the lines over time.
Comparison of actual accomplishments with the goals and objectives (outputs/outcomes): We have made consistent progress with regard to the milestone initially proposed. We are slightly behind the schedule due to delayed hiring process of the key personel (Dr. Romero started only in February 2020) and limited access to the laboratory during the pandemic. However, considering the current global situation, we managed to have reasonable progress in objective 1 by designing and cloning all the reporters and generation of cell lines (one line completed and is under QC, second line is at 80%); in objective 2 by charectarization and QC of two iPSC lines and BrainSpheres generated from those lines; and in objective 3 by conducting toxicity testing experiments of 7 test compounds using one of the lines (dose range finding and functionality experiments). During the initial phase of this project, we troubleshooted and optimized iPSC transfection, generation of homozygous lines as well as 3D electrophysiology.
Future Activities:
We have the gene-editing technique working very efficiently and all the constructs in place to generate the three remaining flag proteins during the next year. Due to pandemic, the original milestone to have the cell line by the mid of the second year, is now planned to be accomplished by the end of the second year. Thus, the main focus for the subsequent reporting period will be the generation, characterization and HCI of the multi-fluorescent BrainSpheres. We will apply the test compunds to those spheres to generate preliminary data, which will help to establish the screening workflow using the multi-fluorescent BrainSpheres.
Journal Articles on this Report : 1 Displayed | Download in RIS Format
| Other project views: | All 9 publications | 9 publications in selected types | All 9 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Chesnut M, Paschoud H, Repoind C, Smirnova L, Hartung T, Zurich MG, Hogberg HT, Pamies D. Human IPSC-Derived Model to Study Myelin Disruption. INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES 2021;22(17):9473. |
R839505 (2020) |
Exit |
Supplemental Keywords:
Biology, Chemicals, Health, Human, Impact, Regulatory, Risk, ToxicsProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.