Grantee Research Project Results
2008 Progress Report: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
EPA Grant Number: R833010Title: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
Investigators: Hashsham, Syed , Tiedje, James M. , Tarabara, Volodymyr
Institution: Michigan State University
EPA Project Officer: Page, Angela
Project Period: September 1, 2006 through August 31, 2009 (Extended to August 31, 2011)
Project Period Covered by this Report: September 1, 2007 through August 31,2008
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The main objectives of this research are to i) develop and validate a highly parallel, sensitive, specific, and quantitative biochip for simultaneous detection of 20 waterborne pathogens, ii) evaluate the use of nanaoparticles to address the issue of viability, and iii) develop an efficient sample concentration scheme for fast and efficient recovery of waterborne pathogens. The progress related to these objectives made during the period September 1, 2007 to August 31, 2008 are described below.
Progress Summary:
1.1 Design and validation of quantitative PCR assays targeting VMGs and fecal indicators
Earlier we reported the validation of the on-chip PCR system for 20 waterborne pathogens. This work is now published (Stetfeld et al., 2008). We also evaluated selected data for virulence and marker genes for their melting characteristics in order to provide higher level of confidence in the idnetities (Pozhitkov et al., 2008). In year 2, additional experiments focusing on high throughput screening of potential indicator markers (41 potential fecal indicator markers representing 30 different organisms) and antibiotic resistance genes were conducted. It is well known that the outcome of a marker search approach can be expedited by high throughput screening both in terms of potential markers and environet and host-specific samples. From the analysis of approximately 100 samples collected and analyzed so far, we can state that a correlation exists between the prevalance of antibiotic resistence genes and potential indicator markers (Figure 1). We have also analyzed the data by clustering algorithms to evaluate if certain markers are host-specific.
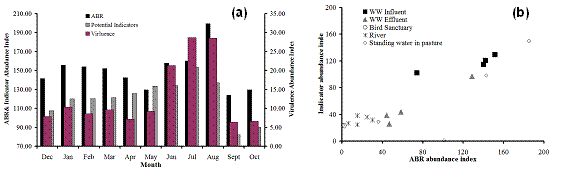
Figure 1. (a) Temporal occurrence of virulence and marker genes, potential indicator markers, and antibiotic resistance genes in various surface water samples, (b) Correlation between the indices of antibiotic resistance gene occurrence and potential indicator marker gene index. Abundance index was calculated as “(26 x Number of amplified genes) – (sum of Ct for amplified genes)”.
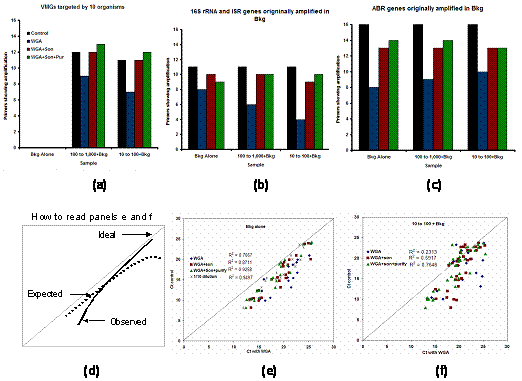
In its present form, this index is not intended for use beyond the comparison made here. We have also evaluated the efficacy of multiple displacement amplification (MDA). MDA is one of several techniques for amplifying small amounts of genomic DNA or whole genomes. In our case, without using the MDA, it would not have been possible to use many of the environmental samples on the on-chip PCR amplification platform (because of low DNA yield from these samples). It was critical, therfore, to quantitatively assess the bias in MDA and how it impacts the conclusion about abundance and detection. The MDA experiment was performed with control mixtures, with and without sonication and purification. The conclusion from these studies is that sonication significantly improved the performance of MDA yielding amplification results closer to the controls. Purification after sonication improved the performance further but to a lesser extent compared to sonication alone (Figure 2 a-c). The surprising result from this experiment was that concentration dependent bias was somewhat more pronounced for genes at higher abundance (lower Cts) than the genes at low abundance (longer Cts; Figure 2 d-e). Since this is such an important and peculiar observation, we are repeating the experiments to confirm it and designing some additional experiments to answer why this is the case. Our expectation was that the gene targets at lower abundance would show greater bias (deviation from the ideal 1:1 behavior) in their copy number when the DNA mixture containing them is subjected to whole genome amplification.
Figure 2. (a-c): Impact of sonication and purification on whole genome amplification (using MDA); (d-f): Bias as measured by change in Ct values on the on-chip PCR assay.
1.2 Dye-doped nanoparticle based detection of viability
To accomplish this objective, reverse microemulsion method was used for synthesis of Rubpy dye doped nanoparticles (NPs) and combined with an antibody-based assay. These dye doped NPs with antibodies were then used to track the temporal growth of Escherichia coli EDL933 and Pseudomonas aeruginosa. The presumption was that growth is the most reliable measure of viability. Hence methods that can document growth much earlier (e.g., after 1 cell grows into 100s of cells) than say observation of colonies due to larger growth (e.g., when 1 cell grows into millions of cells) will be preferred. The NP/antibody-based assay itself takes about 20 min and its sensitivity is comparable to antibody-based methods (~1 cell per ml of water). Using this assay, the growth of E. coli cells could be observed after 2 hr. Simultaneous measurement with real-time PCR, plate count, and absorbance indicated that NP-based method gave the indication of viability earlier than these methods. But it also required more hands on time. Potential of the NP-based assay was explored to assess the viable but non-culturable state of E. coli K12. The manuscript related to this work is in circulation among the co-authors and submission is planned for Applied and Environmental Microbiology. This task is complete and no further work is planned in this area during year 3.
1.3 Integrating the commercially available BioTrove plate with conventional PCR systems:
As indicated in our previous report, we have also been examining a means to integrate OpenArrayTM plates with conventional PCR thermocyclers by fitting with a flat temperature block. A low cost means of SYBR excitation using a blue light electroluminescent film is also being examined for end-point detection of PCR amplification with standard computer scanners. Both these possibilities were evaluated and no problems are seen in transporting the on-chip PCR system to conventional thermal cyclers with gel-doc based reading for end point assay. This option is best suited when presence/absence determination must be made of a large number of pathogens in a large number of samples. Limit of detection is same as for the routine RT-PCR (~ 10-100 copies of target DNA per sample).
1.4 Sample processing and concentration with crossflow microfiltration
Towards the development of a crossflow filtration system for fast and reliable recovery of pathogens from large water samples, a bench-scale crossflow system has been constructed and tested under a range of hydrodynamic conditions using feed water samples seeded with inorganic colloids (SiO2) and virus surrogates (P22, MS2). Both crossflow flux and permeation rate were found to have a dramatic influence on the membrane fouling. While it was possible to eliminate the deposition of microbes on the membrane surface when the membrane was operated in subcritical flux conditions, the permeation rates typical for this regime were unacceptably low for recovery and detection applications. In an effort to minimize irreversible deposition of pathogens on the membrane operated at desirable high permeation rates, we focused on the development of a concentrator that is operated at higher transmembrane pressures and is based on the separation by membranes coated with adhesive films. Specifically, we employed deposition of polyelectrolyte multilayers (PEMs) onto porous support surfaces to produce fouling-resistant membranes (Figure 3). By depositing alternate polyanion and polycation layers on the ultrafiltration membrane support, coatings with adjustable surface charge and hydrophilicity were prepared. By the judicious choice of the components (polyelectrolytes) of the PEM coat, a wide range of functionalities can be introduced no the membrane surface.
Figure 3: Virus adhesion to a regular membrane (top panel) and to a PEM membrane (lower panel). Possibility of regeneration of PEM membrane is also illustrated
Outcomes: (i) The impact of developing an on-chip real time PCR assay for most waterborne pathogens could be considerable both on safety and in developing/regulations. The assay could be used in various exploratory studies and in outbreaks to prioritize microbial contaminants for CCL. (ii) The impact of evaluating the bias in whole genome amplification is equally broad. Quantification of this bias is of great valut to EPA because of the need to share environmental DNA among a number of researchers without the concer for biased results. An environmental sample yielding only 100 ng of DNA could be used by 10s to 100s of researchers if MDA and other genomic DNA amplification techniques to not have significant biases. (iii) In scenarios where viability of an organism must be determined in very short time period (e.g., less than 2 hours), this NP based method will be useful. (iv) Reliable and efficienct recovery of microbial pathogens is key to all detection methods applicable to water industry. Narrowing the variability in recovery efficiency from sample to sample and decreasing the time of sample concentration will be the two outcomes from the tasks associated with sample processing. Both these outcomes will be helpful in minimizing the risk from waterborne pathogens.
Future Activities:
- Development and Validation of On-chip PCR for Waterborne Pathogens: Experiments related to the on-chip PCR system using the pathogens markers will focus on validation of the assay for environmental samples, and evaluate genotyping potential for selected pathogens. Identities of the amplified products will be confirmed by sequencing for better quality assurance.
- Simultaneous Screening of VMG, Potential Indicator Marker Genes, and ABR Genes: For the subtask related to MDA, experiments are underway to replicate the peculiar observation mentioned above (and shown in Figure 3) and evaluate if the MDA has a bias that is concentration or gene dependant. Future experiments in the area of lowering the cost of the on-chip PCR assay will focus on quantification of the gene targets using routine imaging techniques, data processing, and chip material.
- Membrane-based Sample Concentrator: We have been re-configuring the concentrator to be suitable for higher pressure/fluxes operation. The appropriate components (e.g. high pressure peristaltic pump, etc.) have been identified and purchased. The re-configured version of the concentrator will be evaluated. Virus indicators P22 and MS2 will be used as surrogates in evaluating the efficiencies of the proposed concentrator. A larger spectrum of pathogens will be tested in our studies once the methodology is established and fully tested with P22 and MS2. Four more coatings have been prepared and characterized in terms of their surface charge and hydrophilicity. In year 3, we will evaluate regenerability of these new coatings. We will also carry out bench scale testing of developed antifouling films for the detection of the expanded range of pathogens. Evaluation of the performance of the prototype concentrator under conditions of multiple pathogens in natural water matrices is also planned in year 3.
- Publications: Three manuscripts, focusing on i) the correlation of potential indicators with antibiotic resistance genes, ii) the quantitative evaluation of bias during whole genome amplification, and iii) membrane filtration for sample processing will also be written during Year 3.
Journal Articles on this Report : 2 Displayed | Download in RIS Format
| Other project views: | All 16 publications | 5 publications in selected types | All 3 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Pozhitkov AE, Rule RA, Stedtfeld RD, Hashsham SA, Noble PA. Concentration dependency of nonequilibrium thermal dissociation curves in complex target samples. Journal of Microbiological Methods 2008;74(2-3):82-88. |
R833010 (2008) R833010 (Final) |
Exit Exit Exit |
|
|
Stedtfeld RD, Baushke SW, Tourlousse DM, Miller SM, Stedtfeld TM, Gulari E, Tiedje JM, Hashsham SA. Development and experimental validation of a predictive threshold cycle equation for quantification of virulence and marker genes by high-throughput nanoliter-volume PCR on the OpenArray platform. Applied and Environmental Microbiology 2008;74(12):3831-3838. |
R833010 (2008) R833010 (Final) R831628 (2008) |
Exit Exit Exit |
Supplemental Keywords:
Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.