Grantee Research Project Results
2007 Progress Report: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
EPA Grant Number: R833010Title: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
Investigators: Hashsham, Syed , Tiedje, James M. , Tarabara, Volodymyr
Institution: Michigan State University
EPA Project Officer: Page, Angela
Project Period: September 1, 2006 through August 31, 2009 (Extended to August 31, 2011)
Project Period Covered by this Report: September 1, 2006 through August 31,2007
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The objective of this research is to develop and validate a highly parallel, sensitive, specific, and quantitative biochip combining the principles of polymerase chain reaction (PCR) and microarrays for the simultaneous detection of 20 waterborne pathogens. Establishing highly parallel and specific methods are essential to reduce the health risk from microbial pathogens present in source and drinking waters.
Progress Summary:
In this report, we summarize the efforts and results achieved in the following four areas: 1) design and validation of quantitative PCR assays targeting multiple virulence genes and potential fecal indicator markers; 2) integrating the commercially available BioTrove plate with conventional PCR systems; 3) progress of sample concentration with cross flow microfiltration; and 4) the development, characterization, and validation of nano-particles for detection of viable pathogens.
1. Design and validation of quantitative PCR assays targeting VMGs and fecal indicators: We proposed to employ a high-throughput on-chip PCR technique that can economically analyze numerous samples for multiple VMG and fecal indicator assays in a sensitive, quantitative, and specific manner. The commercially available OpenArray™ (developed by BioTrove, Woburn, MA) has been selected to validate and survey designed assays. OpenArrays are of the size of a traditional glass slide (26 mm x 76 mm) and consist of an array of over 3,072 real time PCR reaction chambers (33 nl). The chambers are arranged into 48 sub-arrays (a different sample can be placed on each sub-array) consisting of 64 wells (a different primer set per well). Initially, this instrument will serve as an efficient tool for high throughput and low cost development and validation of real time PCR assays targeting virulence and marker genes. As described below, we are also exploring methods to integrate OpenArray plates with conventional PCR thermocyclers for economic pathogen detection in the water industry.
Validation studies (using the BioTrove system) have been performed on primer sets targeting 220 virulence and 41 potential fecal indicator markers representing 30 different organisms. Multiple assays were designed per organism to increase specificity. Primers were designed by collecting all related sequences available on public databases for a given gene target. Conserved regions targeting all homologues of the target gene were used for candidate primer sets. A subset of the assays was validated with type strains spiked into complex samples, and surveyed further with environmental samples (as described below). Validation of the designed primer sets demonstrated a detection limit of 1 to 10 genomic copies per reaction well. A high rate of specificity and reproducibility was observed with the designed assays with approximately 90 percent of designed assays only showing amplification when target was present. We have also examined approaches that increase reliability of primer design and reduce the extent of validation necessary when developing large-scale QPCR assays. Results demonstrated that sequence characteristics (GC content, Tm, amplicons length, etc.) of primers, amplicons, genome size, amplification conditions, and the matrix in which the target is present played a large factor on primer success. A manuscript describing this research has been submitted (December 2007) for publication to Applied and Environmental Microbiology.
1.2. Survey of assays with environmental samples: Approximately 85 environmental samples have been collected to screen for designed fecal indicator marker genes and a subset of validated virulence genes. Samples were collected from the following: raw, primary, secondary, and tertiary effluent from five separate wastewater treatment plants in Michigan, river water upstream and downstream of the tertiary waste outfall, fecal matter from various hosts (horse, cow, chicken, sheep, and swine) from multiple farms, and multiple samples from surface waters in close proximity to manure storage facilities. Detection of pathogens in these environmental samples will be further verified with a secondary means of identification to ensure specificity of assays. Amplification profiles based on type of environmental sample (surface water, soil, waste-water, fecal contamination) have been observed. Future activities will include a clustering analysis (based on amplified assays) to identify host specific contamination profiles.
2. Integrating the commercially available BioTrove plate with conventional PCR systems: The OpenArray plate used with a conventional PCR thermocycler offers an ideal economical tool to screen for pathogens with high specificity and sensitivity in the water and wastewater industry. The low volume (33 nL) reaction chambers on the OpenArray reduces the cost from $3.50 (for conventional 25 uL real time PCR) to $0.23 per assay. Conventional PCR thermocyclers are increasingly inexpensive and can be purchased for $990 (Antarus Biotech, Huntsville, AL). Thus, we are currently examining a means to integrate OpenArray plates with conventional PCR thermocyclers by fitting with a flat temperature block. A low cost means of SYBR excitation using a blue light electroluminescent film is currently being examined for end-point detection of PCR amplification with standard computer scanners.
3. Progress on sample processing and concentration with crossflow microfiltration: We proposed to develop a crossflow filtration system for the fast and reliable recovery of pathogens from large water samples. A bench-scale crossflow system - a prototype for the smaller scale, portable filtration unit - has been constructed and tested under a range of hydrodynamic conditions using feed water samples seeded with bacteria (E. Coli) and virus surrogate colloids (SiO2). Both crossflow flux and permeation rate were found to have a dramatic influence on the membrane fouling. While it was possible to eliminate the deposition of microbes on the membrane surface when the membrane was operated in subcritical flux conditions, the permeation rates typical for this regime were unacceptably low for recovery and detection applications.

 |
|
|
Figure 2: Total interfacial energy between 6 membranes (5 MPF-coated and 1 commercial) and a surrogate virus
|
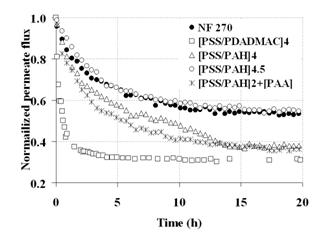
We evaluated coated membranes with respect to 1) their antifouling properties and 2) regenerability. To aid the rational design of antifouling coatings, we employed XDLVO theory as a tool to predict the interactions of microbes with surfaces of given charge and hydrophilicity. A priori XDLVO analysis was used to design films that exhibit repulsive interactions with microbes (Figure 2). Based on the XDLVO predictions, the optimal composition of antifouling coating films was identified. The membranes coated with these optimal films were then tested in a crossflow filtration experiments. Films consisting of 4.5 or 4 bilayers of poly(allylamine hydrochloride) (PAH) and poly(styrene sulfonate) (PSS) were shown to mitigate fouling of membranes by surrogate viruses (Figure 3) and bacteria (not shown) effectively. Permeation rates observed in bench-scale filtration tests were comparable with those observed using ultrafiltraion membranes treated with blocking agents – the system currently used to concentrate and recover viruses.
 |
|
Figure 3: Normalized permeate flux in experiments on the pre-concentration of surrogate viruses
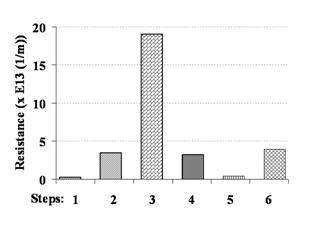
Figure 4: Hydraulic resistance for deconstruction and regeneration for the [PSS/PAH]4 membrane fouled by surrogate virus when backflushing was used to recover viruses from the membrane surface. Steps: (1) UF 50 kDa support; (2) after UF support was modified by MPFs; (3) after the membrane filtered surrogate viruses; (4) after the membrane was backflushed with water for 1 h; (5) after membrane was soaked in pH 10 buffer for 10 min; (6) after MPFs were reapplied to the UF support.
The low fouling rates ensured faster concentrations and minimized irrecoverable loss of microbes to the membrane surface. Another unique advantage of PMF- based membrane modification is the possibility of removing the coating by a simple soaking in a regeneration solution. This allows for the potentially complete recovery of pathogens from feed samples. While the procedure for the removal of coatings has been established for as-prepared PMF films, it was not known if PMFs fouled by biocolloids would be amenable to removal. In our study we demonstrated that removal of the fouled films with adhered colloids was incomplete (Figure 4) when standard regeneration solutions were used. Employing backflushing as a step in the removal sequence was found to enable a complete removal of the coat and associated/deposited deposited particles.
Future activities will include bench scale testing of developed antifouling films for the detection of the expanded range of pathogens. The designed prototype crossflow system will be used to evaluate coated membranes for the concentration of pathogens from natural water matrices.
4. Nanoparticles for detection of viable pathogens: The inability to differentiate between viable and non-viable organisms with PCR based methods necessitates a complementary monitoring device. The unique characteristics exhibited with Dye-doped nanoparticles (DDNPs) such as small size, strong fluorescent signals, photostability and large surface areas provide a sensitive and quantitative means of detecting viable organisms. In our experiments, Rubpy dye doped nanoparticles were synthesized by the reverse microemulsion method, and fluorescence was measured for quantification. This method was used to quantify Escherichia coli EDL933 and Pseudomonas aeruginosa and compared with results obtained using routine methods. In addition, E. coli K12 was induced into a VBNC state and used to test this method. DDNPs displayed quantification of a single cell in a 1 ml water sample. The quantification results correlated well with a routine plate counting method (more than 0.997). For the E. coli samples collected at different growth times, growth was observed after just 2 hr and were quite consistent with those from real-time PCR, plate counting, and absorbance measurement. VBNC induced E. coli K12 was detected and quantified with a detection limit of a single cell in a 1 mL sample. Data demonstrates that the nanoparticle bioassay was sensitive, rapid and rapid. Future studies will explore the simultaneous use DDNPs with multiple dyes (e.g., Texas Red, fluorescein isothiocyanate, tetramethylrhodamine isothiocyanate, tetracarboxy aluminum phthalocyanine) for parallel detection of multiple-targets. A manuscript describing this study is nearly complete and will be submitted by summer 2008.
Journal Articles:
No journal articles submitted with this report: View all 16 publications for this projectSupplemental Keywords:
drinking water, genetics, innovative technology, bacteria, biology, measurement methods, human health, water sample concentration, Water, Scientific Discipline, RFA, Drinking Water, Environmental Engineering, Environmental Chemistry, Environmental Monitoring, biomarkers, drinking water contaminants, nanotechnology, polymerase chain reaction, analytical methods, monitoring, pathogens, biochip,, RFA, Scientific Discipline, Water, Environmental Chemistry, Drinking Water, Environmental Monitoring, analytical methods, monitoring, pathogens, polymerase chain reaction, drinking water contaminants, nanotechnology, biochip, drinking water monitoringProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
