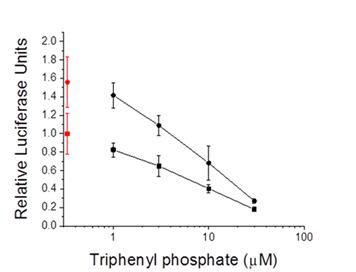Grantee Research Project Results
Final Report: Multi-Sensor Reporter Cell Technology to Assess Hazard Involving Endocrine Signaling Pathways
EPA Grant Number: R835165Title: Multi-Sensor Reporter Cell Technology to Assess Hazard Involving Endocrine Signaling Pathways
Investigators: LeBlanc, Gerald A.
Institution: North Carolina State University
EPA Project Officer: Aja, Hayley
Project Period: March 1, 2012 through February 28, 2016 (Extended to February 28, 2017)
Project Amount: $950,507
RFA: Developing High-Throughput Assays for Predictive Modeling of Reproductive and Developmental Toxicity Modulated Through the Endocrine System or Pertinent Pathways in Humans and Species Relevant to Ecological Risk Assessment (2011) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
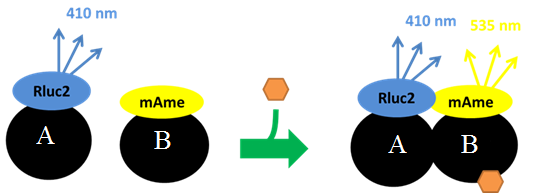
The high-throughput evaluation of toxicity pathways is an emerging paradigm for future chemical toxicity characterizations. The development of assays that evaluate several endpoints along a defined signaling pathway or multiple-pathways within a network could provide insight into: 1) the identification of the most susceptible molecular targets of toxicant action along a cellular pathway/network, 2) the combined actions of different chemicals along a common pathway/network, and 3) multiple actions of a single chemical at different sites within a pathway/network resulting in enhanced toxic outcome. We have exploited bioluminescence resonance energy transfer (BRET) technology (Figure 1) to examine the impact of chemicals on defined cellular signaling pathways. The following pathways were evaluated:
- Human PPARa signaling pathway. This ligand-activated pathway regulates aspects of glucose and lipid metabolism. PPARa activation favors glucose retention and lipid metabolism.
- Human PPARg signaling pathway. This ligand-activated pathway also regulates aspects of glucose and lipid metabolism. PPARg activation favors glucose metabolism and lipid retention. Together, the PPARa and PPARg signaling pathways constitute a major portion of the PPAR signaling network.
- Daphnid MfR signaling pathway. This ligand-activated pathway regulates aspects of development and reproduction in arthropods.
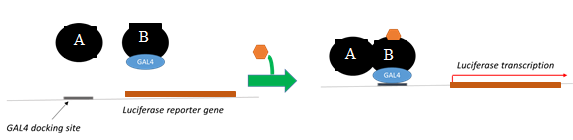
The overall goals of the program were to: 1) establish concordance between conventional reporter gene assays (Figure 2) and BRET assays, and 2) quantify the "gain of information" associated with the simultaneous use of reporter gene and BRET assays.
Summary/Accomplishments (Outputs/Outcomes):
Assay Development
BRET assays
BRET assays were developed to quantify the dimerization of receptor protein subunits and recruitment of co-activators in response to agonist binding. The premise was that many nuclear receptors function as dimers and the dimers recruit co-activator proteins all in response to the binding of an agonist (e.g., hormone, xenobiotic). Agonist-activated receptor assembly was measured by fusing one receptor protein subunit with the luciferase protein, Renilla luciferase 2, and fusing the other protein subunit or co-activator protein with fluorophores that are susceptible to excitation by the photons emitted by the luciferase protein. These fusion constructs were expressed in cells; cells were treated with putative agonist; and, assembly allowed to occur. Light-producing substrate for the Renilla luciferase 2 (DeepBlueSea) was then added and light emission by the fluorophores was then measured. Light emission would be expected to occur only if the protein fused to the fluorophore was bound to the protein containing the Renilla luciferase 2 as close proximity (<10 nm) of the photon source and fluorophore is required for energy transfer to occur. The ratio of the light emitted by the fluorophores, EBFP2 (475 nm) or mAmetrine (535 nm), divided by the light emitted by the Renilla luciferase 2 (410 nm) with corrections for background emissions is referred to as the BRET ratio. BRET ratios were measured in intact cells in real time as a measure of ligand-stimulate protein dimerization (Figure 1).
Figure 1 BRET assay used to measure ligand-stimulated dimerization. Protein A is fused to the Renilla luciferase 2 enzyme which metabolizes an added substrate that emits photons at 410 nm. Protein B is fused to a fluorophore (e.g., mAmetrine) which is excited at 410 nm and then emits photons at 535 nm. The ratio of light emitted at 535 and 410 nm is the BRET ratio used as a measure of protein-protein binding.
Reporter gene assays
Reporter gene assays were developed as a standard to compare BRET assays for accuracy and precision. In addition, BRET and reporter gene assays were used together to evaluate "gain-of -information" associated with the use of both assays. The luciferase gene equipped with upstream GAL4 docking sites was used as a transcriptional reporter. The reporter gene was expressed in cells along with the receptor protein subunits, one of which was fused with the GAL4 binding motif. Agonist-mediated transcriptional activation was measured by the production of light following addition of luciferase substrate (Figure 2).
Figure 2 Luciferase reporter gene transcription assay depicts transcription of the reporter gene (luciferase) by agonist-stimulated receptor dimer.
BRET proof of concept
BRET assays were constructed to assess agonist-stimulated assembly of the human PPARa:RXRa:SRC1 and PPARg:RXRa:SRC1 receptors. For these assays, the RXRa subunit was fused to the Renilla luciferase 2 protein. This allowed for the use of this subunit in any BRET assay involving RXRa dimer partners (e.g., TR:RXR, PXR:RXR, VDR:RXR). PPAR a or g was fused to the fluorophore EBFP2 and SRC1 was fused to mAmetrine. Assays were performed in HepG2 cells.
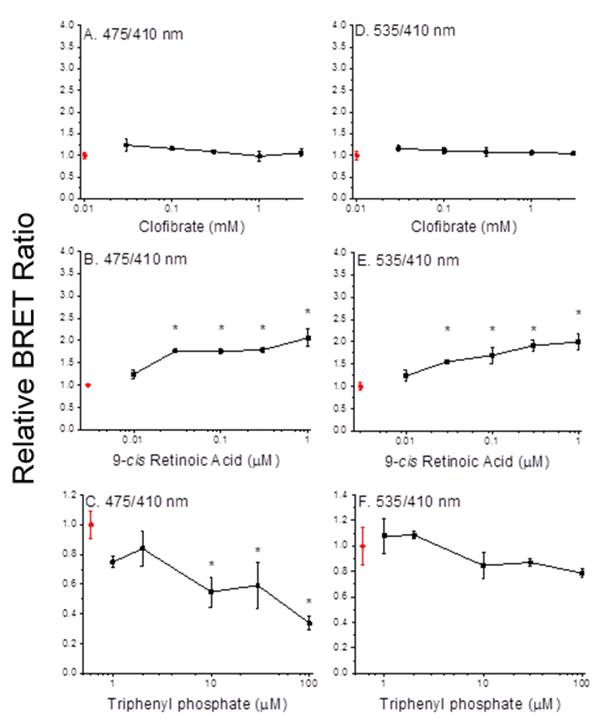
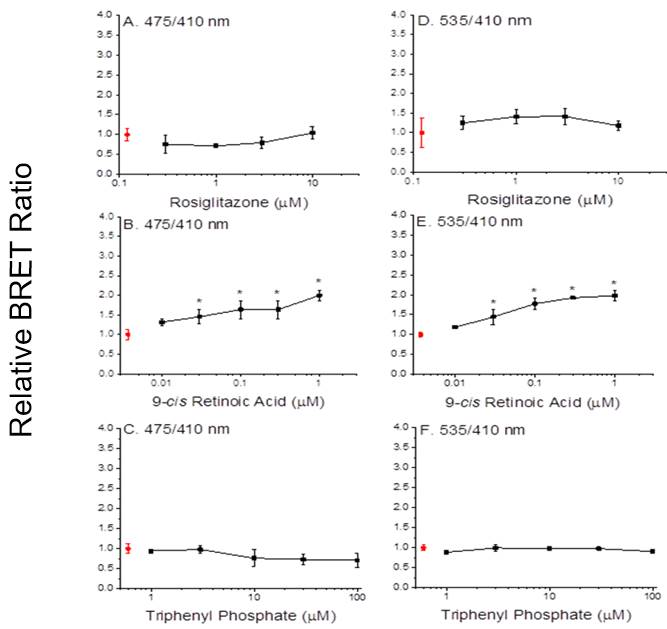
Assembly of both PPARa:RXRa:SRC1 and PPARg:RXRa:SRC1 receptors were detected by BRET following treatment of cells with the RXRa ligand 9-cis retinoic acid (Figures 3B, 3E, 4B, and 4E). However, neither the PPARa ligand clofibrate (Figures 3A and 3D) nor the PPARg ligand rosiglitazone (Figures 4A and 4D) stimulated multimerization of the receptor complexes. In contrast, ligands to either the RXRa or PPAR subunits stimulated transcription in the reporter gene assay. The reason for absence for an increase in the BRET ratio in response to PPAR ligands is unknown, but may indicate that a sub-population of the receptors assemble in a ligand-independent fashion and it is this sub-population that is activated by the PPAR ligands. Regardless, results demonstrated that the BRET has utility in screening chemicals for activity towards the RXRa subunit but not the PPAR subunit.
The limited utility of the BRET assays using these nuclear receptor family members prompted us to evaluate a receptor of another protein family, the bHLH-PAS proteins. The selected receptor was the methyl farnesoate receptor (MfR). This receptor consists of two subunits Met and SRC. The receptor is involved in various aspects of reproductive maturation in crustaceans.
Our first objective was to determine whether the endogenous agonist, methyl farnesoate, stimulated dimerization of the receptor subunits. Indeed, methyl farnesoate stimulated a robust dimerization of the Met and SRC subunits (Figures 5C and 6B). Using BRET, we also demonstrated that the Met subunits form a multimer in cells and this multimer is dissociated in response to methyl farnesoate (Figures 5B and 6A). Thus, Met-Met dissociation and Met-SRC association proved to be discernible steps in the adverse outcome pathway. The BRET assay and the reporter gene assay provided comparable concentration:response curves with methyl farnesoate indicating comparable sensitivity between the two assays (Figure 6C). These results demonstrated that the BRET assay provided a reliable measure of agonist-stimulated activation of the MfR and provided proof of concept for the use of this assay with the MfR and likely other receptors.
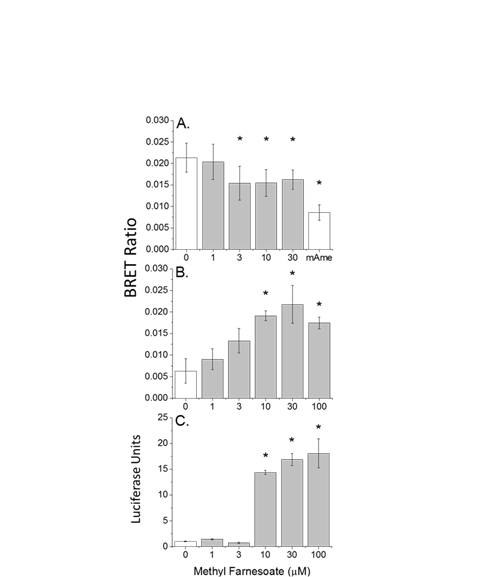
Figure 3. Dimerization of PPARa and RXRa (A-C) and recruitment of SRC1 (D-F) with increasing concentrations of clofibrate (A, D), 9-cis retinoic acid (B, E) and triphenyl phosphate (C, F). Data points and error bars represent the mean and standard deviation, respectively (n=3) and are normalized to the control BRET ratio. Control values (no ligand) are presented in red. An asterisk denotes a significant difference from the control value (p<0.05).
Figure 4. Dimerization of PPARg and RXRa (A-C) and recruitment of SRC1 (D-F) with increasing concentrations of rosiglitazone (A, D), 9-cis retinoic acid (B,E) and triphenyl phosphate (C,F). Data points and error bars represent the mean and standard deviation, respectively (n=3) and are normalized to the control BRET ratio. Control values (no ligand) are presented in red. An asterisk denotes a significant difference from the control value (p<0.05).
BRET in the characterization of PPAR receptor:chemical interactions
Triphenyl phosphate The organophosphate flame retardant triphenyl phosphate has been implicated in the activation of PPARg:RXRa:SRC1. BRET assays were used to evaluate the interaction of this chemical with both PPARa:RXRa:SRC1 and PPARg:RXRa:SRC1. Triphenyl phosphate reduced the BRET ratio associated with the dimerization of PPARa and RXRa along with the recruitment of SRC1 to the complex (Figures 3C and 3F). These results suggest that triphenyl phosphate stimulated the dissociation of the spontaneous PPARa:RXRa:SRC1 aggregates discussed earlier. Reporter gene assays performed in the absence and presence of the PPARa agonist oleic acid also demonstrated that triphenyl phosphate inhibited this receptor complex (Figure 7). Reporter gene assays performed with individual PPARa or RXRa subunits fused to the GAL4 domain revealed that triphenyl phosphate is capable of binding to and suppressing the activity both subunits (Figure 8).
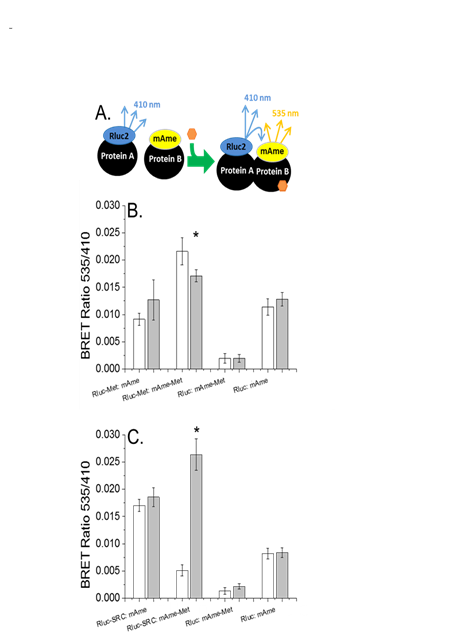
Figure 5. Multimerization of Met and its association with SRC1 a) Diagramatic representation of BRET assays used in these experiments. Protein A fused to the photon donor Rluc2 and protein B fused to the fluorophore m-Ametrine are expressed in cells. Photons (410 nm) emitted by the Rluc2 substrate are emitted. Upon addition of agonist (orange hexagon) the proteins dimerize. Now, photons emitted by the Rluc2 substrate excite the closely associated and fixed fluorophore and photon are emitted at 535 nm providing an elevated BRET ratio (535 nm emission/410 nm emission). When assessing ligand-mediated dissociation, an elevated BRET ratio exists in the absence of agonist and decreases with agonist addition. b) Spontaneous multimerization of Met with ligand-dependent dissociation White bars denote BRET ratios in the absence of methyl farnesoate. Gray bars represent BRET ratios from cells treated with 30 µM methyl farnesoate. c) Methyl farnesoate-dependent dimerization of the MfR subunits White bars denote BRET ratios in the absence of methyl farnesoate. Gray bars represent BRET ratios from cells treated with 100 µM methyl farnesoate. All data are presented as the mean and standard deviation (n=3). An asterisk denotes a significant difference between the control and methyl farnesoate treatment (p<0.05, Student's t test).
Figure 6. Concentration-responses for methyl farnesoate-dependent MfR subunit interactions a) Rluc2-Met and mAmetrine-Met spontaneous multimerization and dissociation with methyl farnesoate addition. The treatment denoted as "mAme" consisted of cells expressing Rluc2-Met and free mAmetrine. This treatment provides the BRET ratio that is Met multimerization independent (e.g., baseline BRET ratio). b) Methyl farnesoate-stimulated dimerization of Rluc2-SRC and mAmetrine-Met. c) Luciferase reporter gene activation in cells expressing Met and SRC-Gal4 with increasing concentrations of methyl farnesoate. All data are presented as mean and standard deviation (n=3). An asterisk denotes a significant difference from the 0 µM methyl farnesoate treatment (ANOVA).
Figure 7. PPARa:RXRa-Gal4 driven reporter gene activity in cells treated with triphenyl phosphate alone (circles) or in combination with the PPARaligand oleic acid (30 µM) (squares). Control (0 µM triphenyl phosphate) values are presented in red. Data are presented as the mean and standard deviation (n=3).
BRET assays performed with PPARg:RXRa:SRC1 and triphenyl phosphate produced decidedly different results. Triphenyl phosphate neither reduced nor increased the BRET ratio associated with PPARg:RXRa:SRC1 (Figures 4C and 4F). These results are consistent with those observed with PPARg agonists (e.g., rosiglitazone, Figures 4A and 4D). Indeed, reporter assays revealed that triphenyl phosphate is a potent agonist to the PPARg subunit (Figure 9). Results thus indicate that triphenyl phosphate acts as an antagonist when associated with the RXRa subunit but acts as a strong agonist when associated with the PPARg subunit. The net effect of this dual activity appears to be weak agonistic activity towards the receptor complex.
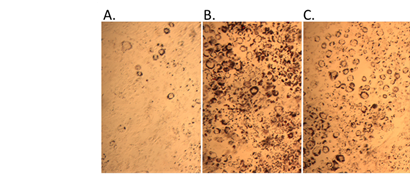
An agonist to the PPARg:RXRa:SRC1 receptor complex would be predicted to increase the differentiation of pre-adipocytes to adipocytes with commensurate increases in adipocyte lipid accumulation. Exposure of mouse 3T3-L1 cells to triphenyl phosphate did increase pre-adipocyte differentiation and lipid accumulation (Figure 10).
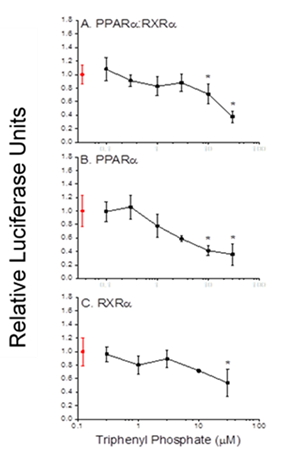
Figure 8. Gal4-driven luciferase reporter gene activity following treatment of cells with triphenyl phosphate. A. Activity associated with the PPARa:RXRa-Gal4 heterodimer. B. Activity associated with the PPARa-Gal4 subunit. C. Activity associated with the RXRa-Gal4 subunit. Control (0 µM triphenyl phosphate) values are presented in red. Data are presented as the mean and standard deviation (n=3) and are normalized to the control values. An asterisk denotes a significant (p<0.05) difference from the control.
Other organophosphates Having observed the inhibitory effect of triphenyl phosphate on the PPARa receptor complex, several additional organophosphate compounds were evaluated along with known PPARa:RXRa agonists for their effects on PPARa signaling using both the BRET assay and the reporter gene assays.
Results with known agonist yielded similar results as reported above. RXRa ligands (9-cis retinoic acid and LGD 1069) activated the BRET and reporter gene assays with similar potency (Table 1). PPARa agonists (clofibrate and Wy-14,643) activated the reporter gene assay only (Table 1). All of the organophosphate compounds were inhibitory in the reporter gene assay and all but one compound was inhibitory in the BRET assay (Table 1).
Resulted supported previous observations that BRET assays can detect PPAR receptor antagonist. However, the assay was somewhat less sensitivity in detecting the inhibitory effects of the organophosphate compounds as compared to reporter gene assays. Again, it must be noted that for both BRET and the reporter gene assays are designed to detect receptor activation. At this time we do not understand the significance of a loss of activity as measured with the organophosphates in these assays.
| Compound | Lowest Observed Effect Concentration (μM) | |
|---|---|---|
| BRET | Reporter Gene Assay | |
| 9-cis retinoic acid | 0.03 | 0.10 |
| LGD1069 | ≤0.01 | ≤0.01 |
| Clofibrate | >3.0 | ≤0.10 |
| Wy-14,643 | >30 | 1.0 |
| 2-Ethylhexyl diphenyl phosphate | inhibition 300 | inhibition 3.0 |
| Tri-o-tolyl phosphate | inhibition 30 | inhibition 30 |
| Tri-n-butyl phosphate | inhibition 300 | inhibition 10 |
| Tris(2-butoxyethyl) phosphate | ≥300 | inhibition 100 |
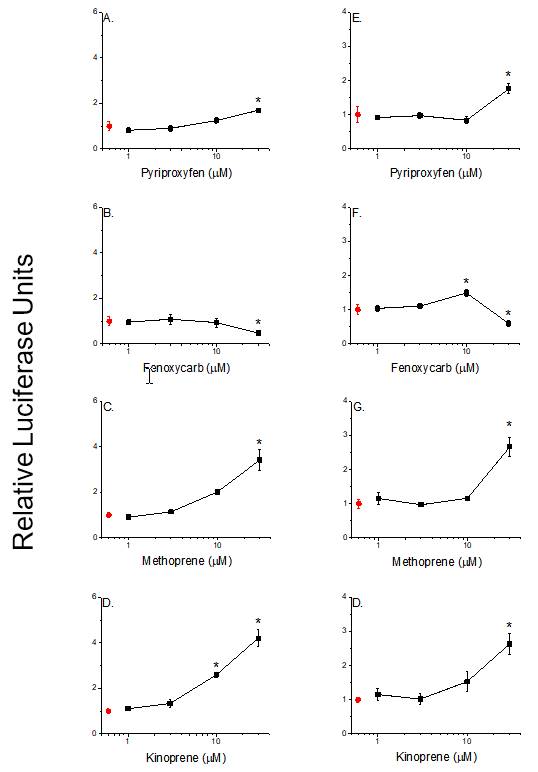
Insect growth regulators Insect growth regulators (IGRs) are a class of insecticides designed to disrupt processes that are specific to insect physiology. We noted that the IGR pyriproxyfen is structurally similar to the potent PPARg agonist rosiglitazone. Accordingly, we hypothesized that pyriproxyfen and related IGRs may serve as good candidate chemicals to assess activation of the PPARg:RXRa:SRC1 receptor complex and its detection using BRET. Reporter gene assays revealed that all of the IGRs evaluated—pyriproxyfen, fenoxycarb, methoprene, and kinoprene—transactivated the PPARg receptor complex (Figure 11). More detailed analysis revealed that all of the IGRs interacted with the PPARg subunit while some also interacted with the RXRa subunit. Activation, however, was dependent upon association of the IGRs specifically with the PPARg subunit. None of the IGRs stimulated receptor subunit dimerization as detected with the BRET assay.
These results are consistent with our prior observations that agonist of the PPAR subunit do not stimulate receptor assembly but rather must activate existing assembled protein complexes. Results further indicate that the BRET assay is not universally applicable for use with all receptor complexes.
Figure 9. Gal4-driven luciferase reporter gene activity following treatment of cells with triphenyl phosphate. A. Activity associated with the PPARg:RXRa-Gal4 heterodimer. B. Activity associated with the PPARg-Gal4 subunit. Control (0 µM triphenyl phosphate) values are presented in red. Data are presented as the mean and standard deviation (n=3) and are normalized to the control values. An asterisk denotes a significant (p<0.05) difference from the control.
Figure 10. Pre-adipocyte differentiation and lipid accumulation with triphenyl phosphate treatment. A. control, B. 2 µM rosiglitazone (positive control), C. 10 µM triphenyl phosphate.
Results indicated that the BRET assay is effective in detecting receptor antagonist implying antagonist-mediated receptor dissociation. However, these phenomenon is poorly understood and requires further study. Results also support our previous observations that agonist that bind to the PPAR subunit do not stimulate receptor assembly. This places the BRET assay at a significant disadvantage relative to reporter gene assays for the detection of receptor agonists.
Figure 11. PPARg:RXRa transactivation by IGRs as indicated by increased transcription of a PPRE-driven luciferase reporter gene after 24 (A-D) and 48 hour incubations (E-H). Data are presented as means with error bars depicting standard deviations (n=3). An asterisk denotes a significant difference from the control (p<0.05, One-Way ANOVA, Tukey's Multiple Comparison Test).
BRET in the characterization of MfR receptor:chemical interactions
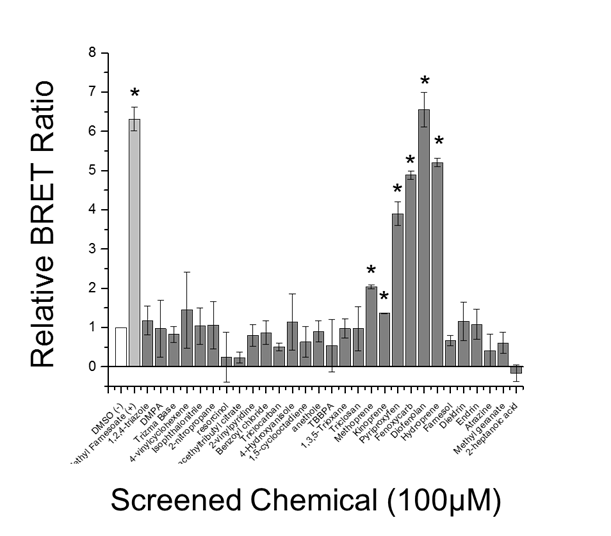
Thirty chemicals were evaluated for activity towards the MfR using the BRET assay to assess accuracy and precision relative to the reporter gene assay. Chemicals included the positive control, methyl farnesoate, IGRs, common-use biocides, and EPA-designated high production volume chemicals. Methyl farnesoate and the six IGRs were detected as MfR-active in the BRET assay (Fig. 12). All of the other chemicals screened negative in the assay. Similarly, methyl farnesoate and five of the six IGRs screened positive in the reporter gene assay and all other chemicals were negative (Fig, 13). Concentration-response analyses revealed that the BRET assay was typically more sensitive than the reporter gene assay.
Figure 12. Chemical screen for MfR activation using BRET assay White bar indicates DMSO vehicle control, light grey bar indicates the positive positive control (methyl farnesoate), and dark grey bars indicate screened chemicals. Data are presented as means with standard deviations (n = 3) and an asterisk denotes a significant increase in Met-SRC dimerization compared to DMSO control (p<0.05).
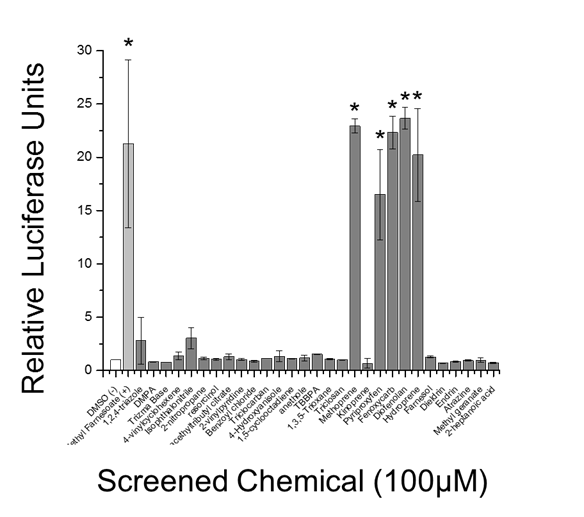
Figure 13. Chemical screen for MfR activation using luciferase reporter gene assay White bar indicates DMSO vehicle control, light grey bar indicates the positive control (methyl farnesoate), and dark grey bars indicate screened chemicals. Data are presented as means with standard deviations (n = 3) and an asterisk denotes significant increase in Met-SRC dimerization compared to DMSO control (p<0.05).
In the characterization of MfR receptor:chemical interactions, thirty chemicals were evaluated for activity towards the MfR using the BRET assay to assess accuracy and precision relative to the reporter gene assay. Chemicals included the positive control, methyl farnesoate, IGRs, common-use biocides, and EPA-designated high production volume chemicals. Methyl farnesoate and the six IGRs were detected as MfR-active in the BRET assay (Figure 12). All of the other chemicals screened negative in the assay. Similarly, methyl farnesoate and five of the six IGRs screened positive in the reporter gene assay and all other chemicals were negative (Figure 13). Concentration-response analyses revealed that the BRET assay was typically more sensitive than the reporter gene assay.
Conclusions:
These results provide evidence to support the following conclusions with respect to use with the MfR:
- The BRET assay is highly accurate in detecting active chemicals.
- The BRET assay has a negligible incidence of false positives.
- The BRET assay has greater sensitivity than the reporter gene assay.
- The BRET assay is less expensive, requires shorter time duration, and is more conducive to high-throughput applications than reporter gene assays.
Additional study is warranted to identify which receptor complexes are conducive to the BRET assay (e.g., MfR) and which are not (e.g., PPAR).
Overall, our results demonstrate that agonist-mediated receptor assembly represents an early event in some nuclear receptor adverse outcome pathways. For relevant receptors, this endpoint can serve as a viable alternative to the gene activation endpoint as measured using reporter gene assays for the high-throughput screening of chemicals. Advantages to the BRET assay include; a) short chemical exposure duration time, b) real-time, in-situ endpoint measurement, c) reduced cost. Critical to further development of the BRET toxicity screening assay are: a) identifying those receptors in which agonist-mediate receptor assembly is relevant, and b) characterizing the significant of receptor dissociation as a measure of antagonist action.
References:
- LeBlanc G.A. and Medlock E.K. 2015. Males on demand: the environmental-neuro-endocrine control of male sex determination in daphnids. FEBS J. 282:4080-4093.
- Wang Y., Kwon G., An L., Holmes CN, Haeba M., and LeBlanc G.A. 2016. Differential interaction of the flame retardant triphenyl phosphate within the PPAR signaling network. MOJ Toxicology. 2(3):00039.
- Kakaley E.K., Wang H., and LeBlanc G.A. 2017. Agonist-mediated assembly of the crustacean methyl farnesoate receptor. Scientific Reports. 7:45071/DOI:10.1038/srep45071.
- Kakaley E.K. Eytcheson S., and LeBlanc G.A. Ligand-mediated receptor assembly as an endpoint for high-throughput chemical toxicity screening. Environ. Sci. Tech. (under review).
- LeBlanc G.A. 2012. Novel modes of endocrine action and disruption. Environmental Endocrine Disruptor Testing and Evaluation: Current and Future Developments. 6th Society of Environmental Toxicology and Chemistry Europe Special Science Symposium. Brussels, Belgium.
- LeBlanc G.A., Wang Y.H.., Holmes C.N., Kwon G., and Medlock E.K. 2013. A transgenerational endocrine pathway in Crustacea. Carolinas Society of Environmental Toxicology and Chemistry Meeting. Raleigh, NC.
- Medlock E.K. and LeBlanc G.A. 2013. Reproductive and developmental signaling pathway in crustaceans. Society of Environmental Toxicology and Chemistry North America 34th Annual Meeting. Nashville, TN.
- Holmes C.N. and LeBlanc G.A. 2013. Integrating synergy into additive models of chemical mixtures toxicity. Carolina SETAC Annual Meeting, Raleigh, NC.
- LeBlanc G.A. 2013. Emerging targets of endocrine disruption: the xenocrine system. Society of Environmental Toxicology and Chemistry North America Focused Topic Meeting. Endocrine Disruption: Chemical Testing, Risk Assessment Approaches, and Implications. US Environmental Protection Agency, NC.
- LeBlanc G.A. 2013. Sex on demand: environmental sex determination in crustacean. University of North Carolina Wilmington, NC.
- LeBlanc G.A. 2013. Endocrine disruption beyond the estrogen, androgen, and thyroid signaling pathways. Webinar presented by the U.S. Environmental Protection Agency, Washington, DC.
- LeBlanc G.A. 2013. Multi-sensor reporter cell technology to assess hazard involving endocrine signaling pathways. U.S. Environmental Protection Agency, Washington, DC.
- Holmes C.N. and LeBlanc G.A. 2014. Insect growth-regulating insecticides differentially activates the PPAR signaling network. Society of Toxicology 53th Annual Meeting, Phoenix, Az.
- Medlock E.K. and LeBlanc G.A. 2014. Activation and disruption of a crustacean reproductive signaling pathway. Society of Environmental Toxicology and Chemistry 35th Annual Meeting, Vancouver BC, Canada.
- Medlock E.K. and LeBlanc G.A. 2014. Activation of a sex determination signaling cascade in crustaceans. Carolinas Society of Environmental Toxicology and Chemistry Meeting, Clemson, SC.
- Medlock E.K. and LeBlanc G.A. 2015 Deciphering an environmental-endocrine signaling pathway for the identification of efficacious screening endpoints. Society of Environmental Toxicology and Chemistry 36th Annual Meeting. Salt Lake City, UT.
- Holmes C.N., Kwon G., Wang Y., An L., and LeBlanc G.A. 2015. Bioluminescence energy resonance transfer: an innovative technology for evaluating chemical interactions with nuclear receptors. Society of Toxicology Future Tox III Meeting, Arlington, VA.
- Holmes C.N., Kwon G., Wang Y., An L., and LeBlanc G.A. 2015. Ligand-dependent receptor assembly as an endpoint for the high-throughput screening of chemicals for endocrine activity. Society of Environmental Toxicology and Chemistry 36th Annual Meeting, Salt Lake City, UT.
- Holmes C.N. and LeBlanc G.A. 2015. Insect growth regulating insecticides differentially activate the PPAR signaling network. Society of Toxicology 54th Annual Meeting, San Diego, CA.
- LeBlanc G.A., An L., Eytcheson S., Holmes C.N., Kwon G., Wang Y., and Medlock E.K. 2015. High-throughput assays to assess interactive effects of chemicals on endocrine signaling pathways. Environmental Toxicology and Chemistry Focus Topic Meeting: Endocrine Disruption, U.S. Environmental Protection Agency, Research Triangle Park, NC.
- Medlock E.K. and LeBlanc G.A. 2015. Males on demand: the environmental-neuro-endocrine control of male sex determination in daphnids. Poscast, FEBS Journal website.
- Medlock E.K. and LeBlanc G.A. 2015. Bioluminescence resonance energy transfer (BRET) as an approach for high-throughput screening of chemicals for toxicity. North Carolina State University Graduate Student Symposium, Raleigh, NC.
Journal Articles on this Report : 7 Displayed | Download in RIS Format
| Other project views: | All 26 publications | 7 publications in selected types | All 7 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Adoteye K, Banks HT, Cross K, Eytcheson S, Flores KB, LeBlanc GA, Nguyen T, Ross C, Smith E, Stemkovski M, Stokely S. Statistical validation of structured population models for Daphnia magna. Mathematical Biosciences 2015;266:73-84. |
R835165 (Final) |
Exit |
|
|
Adoteye K, Banks HT, Flores KB, LeBlanc GA. Estimation of time-varying mortality rates using continuous models for Daphnia magna. Applied Mathematics Letters 2015;44:12-16. |
R835165 (Final) |
Exit |
|
|
Kakaley EK, Wang HY, LeBlanc GA. Agonist-mediated assembly of the crustacean methyl farnesoate receptor. Scientific Reports 2017;7:45071 (9 pp.). |
R835165 (Final) |
Exit Exit Exit |
|
|
LeBlanc GA, Medlock EK. Males on demand: the environmental-neuro-endocrine control of male sex determination in daphnids. The FEBS Journal 2015;282(21):4080-4093. |
R835165 (2015) R835165 (Final) |
Exit Exit Exit |
|
|
Medlock Kakaley EK, Eytcheson SA, LeBlanc GA. Ligand-mediated receptor assembly as an endpoint for high-throughput chemical toxicity screening. Environmental Science & Technology 2017;51(16):9327-9333. |
R835165 (Final) |
Exit Exit Exit |
|
|
Rutter E, Banks H, LeBlanc G, Flores K. Continuous Structured Population Models for Daphnia magna. BULLETIN OF MATHEMATICAL BIOLOGY 2017;79:2627-2648 |
R835165 (Final) |
Exit |
|
|
Wang Y, Kwon G, An L, Holmes CN, Haeba M, LeBlanc GA. Differential interactions of the flame retardant triphenyl phosphate within the PPAR signaling network. MOJ Toxicology 2016;2(3):00039 (11 pp.). |
R835165 (Final) |
Exit Exit Exit |
Supplemental Keywords:
endocrine, high-throughput screening, cell-based assayProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
- 2015 Progress Report
- 2014 Progress Report
- 2013 Progress Report
- 2012 Progress Report
- Original Abstract
7 journal articles for this project