Grantee Research Project Results
2009 Progress Report: Bioavailability of Metallic Nanoparticles and Heavy Metals in Landfills
EPA Grant Number: R833893Title: Bioavailability of Metallic Nanoparticles and Heavy Metals in Landfills
Investigators: Hu, Zhiqiang , Elias, Dwayne A. , Wall, Judy D.
Current Investigators: Hu, Zhiqiang , Wall, Judy D. , Elias, Dwayne A.
Institution: University of Missouri - Columbia
EPA Project Officer: Aja, Hayley
Project Period: April 1, 2009 through March 31, 2012
Project Period Covered by this Report: April 1, 2009 through March 31,2010
Project Amount: $399,262
RFA: Exploratory Research: Nanotechnology Research Grants Investigating Fate, Transport, Transformation, and Exposure of Engineered Nanomaterials: A Joint Research Solicitation - EPA, NSF, & DOE (2007) RFA Text | Recipients Lists
Research Category: Nanotechnology , Safer Chemicals
Objective:
The objectives of this research are to determine bioavailability of nanoparticles and heavy metal species in bioreactor landfills as compared to traditional municipal solid waste landfills; and to elucidate the mechanisms governing bioavailability as well as the mode of antimicrobial action by nanoparticles. In order to accomplish these objectives, three hypotheses will be tested: 1) nanoparticles that can leach into the water from landfill runoff are bioavailable; 2) nanoparticles enhance heavy metal bioavailability and leachate toxicity; and 3) the bioavailability of nanoparticles and heavy metals is higher in bioreactor landfills than in traditional landfills.
Progress Summary:
In this report, we summarize the efforts and results achieved in the following three areas: 1) bioavailability of nanoparticles inferred from biochemical methane potential (BMP) tests; 2) Determination of the changes of microbial community structure and species abundance using quantitative polymerase chain reactions (qPCR); and 3) and lab-scale landfill reactor set-up and operation.
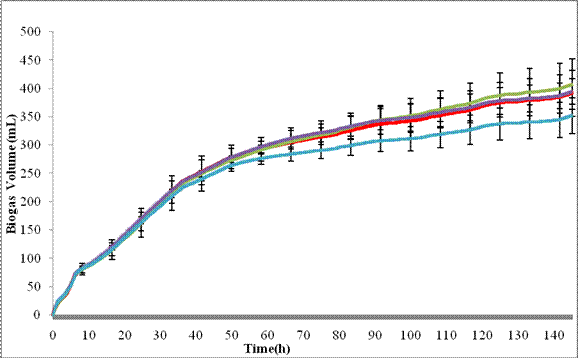
1. Bioavailability inferred from biochemical methane potential (BMP) tests. The BMP rates are being monitored using real-time anaerobic flow modules in the presence and absence of nanoparticles. In recent work, we evaluated the impact of AgNPs on anaerobic digestion and compared the results with those from Ag+ ions. A modified biochemical methane potential test was used to infer anaerobic microbial activity and nanosilver toxicity. The biogas production tests have proved to be very sensitive for the detection of metal toxicity to sludge through the evaluation of biogas generation rate and cumulative gas volume. Digested sludge was taken from a two-stage mesophilic digester (run at 20-30 days of hydraulic retention time) at the Columbia Waste Water Treatment Plant in Columbia, MO. Stock suspensions of 0.7 mM silver nanoparticles were made from silver nitrate (AgNO3, EM Science) by adding sodium borohydride (NaBH4, Sigma) with polyvinyl alcohol (PVA) as a capping agent. A sixteen-bottle AER-200 Respirometer (CHALLENGE Technology, AK) was used for all batch anaerobic digestion experiments. Each 500 mL glass bottle (with septa) was fed with 300 ml digested sludge. The sludge was exposed to AgNPs at three different concentrations: 17.5 µM (1.9 mg/L), 87.5 µM (9.5 mg/L) and 175 µM (18.9 mg/L) along with the control. The cumulative biogas production with and without AgNP treatment was automatically monitored and recorded (Figure 1). Based on the biogas production curves, there was no significant difference of biogas production between the AgNP treated sludge and the control when the concentrations of AgNPs were 87.5 µM (9.5 mg/L) and below. However, AgNPs at the concentration of 175 µM (18.9 mg/L) in the sludge appeared to have the lower cumulative gas volume. More research work to determine the volume of biogas produced from lab-scale landfill reactors using real-time anaerobic flow modules (Challenge Respirometer System) is underway.
Figure 1. Cumulative gas production in the digested sludge samples exposed to different concentrations of AgNPs verses time. The biogas volume was automatically recorded every 10 minutes by the respirometer. The red, purple, green and blue lines represent the control, samples treated with 17.5 µM, 87.5 µM and 175 µM AgNPs, respectively. Error bars represent one standard deviation of triplicate measurements.
2. Determination of microbial community structure and species abundance. Methanogens typically include two families of acetoclastic methanogens that transform acetate to methane, Methanosaeta and Methanosarcina, and three classes of hydrogenotrophic methanogens that utilize hydrogen and carbon dioxide to form methane, Methanobacteriales, Methanococci and Methanomicrobiales. Protocols for qPCR to determine the relative abundance of methanogenic species in two types of landfill reactors have been developed. PCR primers and TaqMan probes were designed based on known nucleotide sequences of acetoclastic and hydrogenotrophic methanogens. Total genomic DNA was extracted from 1.5 mL well-mixed anaerobic sludge (or leachate), using the MoBio Ultra Clean TM Soil DNA isolation kit (MoBio Laboratories, Inc., CA). Real time quantitative PCR assays were performed using ABI 7500 Real Time PCR System and 7500 SDS system software version 1.4. All the qPCR internal standards were prepared from 16S rRNA gene clones of the known methanogens or SRB.
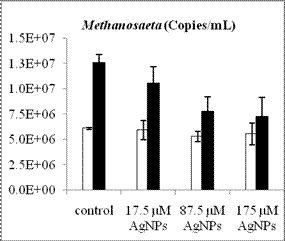
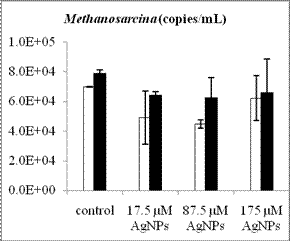
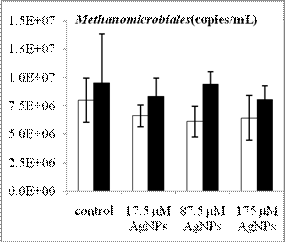
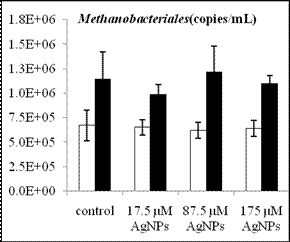
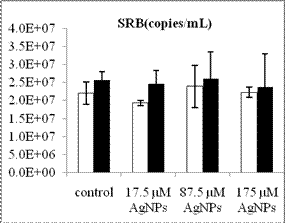
By using real-time quantitative PCR assays we have determined the abundance of methanogens and SRB before and after anaerobic digestion for the four treatments (control, 17.5 µM AgNPs, 87.5 µM AgNPs and 175 µM AgNPs). The cell number of acetoclastic methanogens inferred from their gene copy numbers generally increased after digestion, regardless of AgNP treatment (Figure 2). Compared to Methanosarcina with gene copy numbers in the order of 104, Methanosaeta species had a relatively higher numbers, with gene copy numbers in the range of 106-107. Hence, Methanosaeta species was the dominant acetoclastic methanogens in the digested sludge. While the overall populations of methanogens and SRB increased after anaerobic degradation of glucose, the population of the dominant acetoclastic methanogen Methanosaeta was decreased as the nanosilver concentration increased. Future work will include leachate sample collection, DNA extraction and qPCR analysis to determine the changes of anaerobic microbial community structure and species abundance during anaerobic digestion in lab-scale landfills.
 |
 |
| (a) | (b) |
 |
 |
| (c) | (d) |
 |
|
| (e) |
Figure 2. Population changes of methanogens and SRB inferred from the gene copy number before (white bars) and after (black bars) anaerobic digestion. The representative acetoclastic methanogens include Methanosaeta (a) and Methanosarcina (b) and the hydrogenotrophic methanogens include Methanomicrobiales (c) and Methanobacteriales (d). Error bars represent one standard error of the mean (n =3).
3. Lab-scale landfill reactor set-up and operation. We have collected waste materials from the Columbia Sanitary Landfill (Columbia, MO.) The waste composition was shown in Table 1. The water content of the landfill waste was 9.7% before packing. All the waste was shredded into small size (<1 inch). Prior to filling of the 9 L glass reactor (Figure 3), a 2-inch thickness of pea stone (gravel, 2 kg per landfill reactor) will be placed at the bottom of the reactor to retain small particles from leaching out. After a total of 3 kg landfill waste with or without nanoparticle treatment and aliquots (1000 ml) of anaerobic sludge (served as inocula, COD =10 000 mg/L) were added in each reactor, the waste mixture was covered with gravel. Leachate in some of the landfill reactors were collected for chemical analysis on a biweekly basis. The biogas production are being recorded using Challenge Respirometric System.
Table 1 Composition of landfill solid waste
|
Weight (kg) |
Ratio |
|
|
Metal |
0.301 |
0.91% |
|
Paper |
4.65 |
14.06% |
|
Rock |
5.85 |
17.69% |
|
Wood and Shredding |
1.575 |
4.76% |
|
Soil with organics |
12.15 |
36.74% |
|
Fruits and vegetable |
5.173 |
15.64% |
|
Plastic bags |
3.375 |
10.20% |
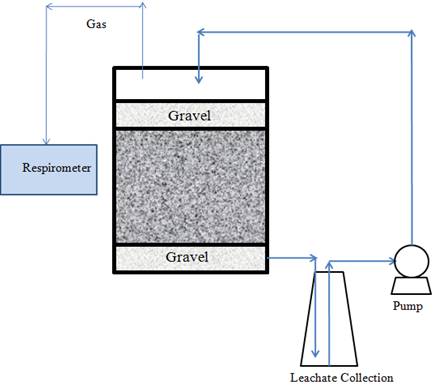
Figure 3. A schematic of lab-scale landfill reactor with leachate recirculation
A total of 6 lab-scale landfill reactors including conventional (without leachate recirculation) and bioreactor (with leachate recirculation) have been set up and operated. These reactors were labeled as following: conventional landfill (control, #1), conventional landfill (treated with 10 mg/L of nanosilver, #4), bioreactor landfill (control, #6), bioreactor landfill (treated with 10 mg/L of nanosilver, #2), bioreactor landfill (treated with 10 mg/L of nanosilver, #3, same as #2), bioreactor landfill (treated with 1 mg/L of nanosilver, #5). Leachates collected from these reactors at the beginning of operation have been analyzed and the result is listed in Table 2. The chemical oxygen demand (COD) ranged from 16975 to 18100 ppm in all reactors. The concentrations of NH4+-N ranged from 46.00 to 50.95 ppm while the concentrations of NO3-N varied from 0 to 95.98 ppm.
Table 2. Chemical analysis of leachate from landfill solid waste
|
Reactor # |
pH |
COD (mg/L) |
NH4+-N (mg/L) |
NO3--N (mg/L) |
NO2--N (mg/L) |
|
1 |
5.96 |
16975 |
46.26 |
28.46 |
0 |
|
2 |
6.21 |
17287.5 |
46.96 |
17.20 |
0 |
|
3 |
5.95 |
17287.5 |
46.00 |
2.73 |
0.39 |
|
4 |
5.96 |
17475 |
49.39 |
95.98 |
0.15 |
|
5 |
6.17 |
18100 |
49.65 |
0 |
0.23 |
|
6 |
5.96 |
17662.5 |
50.95 |
60.61 |
0 |
Total genomic DNA was extracted from 1.5 mL well-mixed leachate from each sample, using the MoBio Ultra Clean TM Soil DNA isolation kit (MoBio Laboratories, Inc., Carlsbad, CA). Genomic DNA extracted was quantified using Nanodrop-standard method at 260 nm, and purity was determined by measuring the 260/280 nm absorbance ratio. The extracted DNA samples were stored at -20 oC before use. Future work will continue to monitor cumulative gas production and leachate composition in each reactor.
A paper entitled, Impact of Nanosilver on Anaerobic Sludge Digestion and Methanogenic Population Dynamics, was submitted to Environmental Science & Technology.
Journal Articles on this Report : 1 Displayed | Download in RIS Format
| Other project views: | All 4 publications | 4 publications in selected types | All 4 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Yang Y, Xu M, Wall JD, Hu Z. Nanosilver impact on methanogenesis and biogas production from municipal solid waste. Waste Management 2012;32(5):816-825. |
R833893 (2009) R833893 (Final) |
Exit Exit Exit |
Supplemental Keywords:
bioavailability, cellular, heavy metals, analytical, absorption, adsorption, bioaccumulation, gene expression, growth, mechanisms, metabolism, toxins, nanotechnology, microbiology, biotechnologyProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.