Grantee Research Project Results
2013 Progress Report: Model of Toxicant Response in Engineered Liver
EPA Grant Number: R834998Title: Model of Toxicant Response in Engineered Liver
Investigators: Rajagopalan, Padmavathy , Murali, T. M. , Ehrich, Marion
Institution: Virginia Tech
EPA Project Officer: Chung, Serena
Project Period: June 1, 2011 through May 30, 2015
Project Period Covered by this Report: June 1, 2013 through May 31,2014
Project Amount: $750,000
RFA: Computational Toxicology: Biologically-Based Multi-Scale Modeling (2010) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
Objective 1: Use 3D liver mimics as a platform for testing the hepatotoxicity of prototypic hepatotoxicants.
Objective 2: Compute biological process linkage networks that anchor phenotypes triggered by hepatotoxicants to gene expression profiles.
Objective 3: Discover the effects on hepatic cells of combinations of the toxicants examined in Objectives 1 and 2. Prioritize toxicity pathways for further proteomic and metabolomic analyses.
Progress Summary:
Our overall approach has the following components. We have assembled 3D organotypic models composed of primary rat hepatocytes, liver sinusoidal endothelial cells (LSECs), and Kupffer cells separated by a polyelectrolyte layer that mimics the Space of Disse (SoD). The polymeric SoD is assembled using Type 1 collagen and hyaluronic acid. Figure 1 is a schematic of the assemble of 3D liver models used in these studies.
We have completed the experimental work on acetaminophen (APAP) induced hepatotoxicity using conventional 2D cell cultures and the organotypic models assembled in the Rajagopalan group.
We are in the process of conducting transcriptomics. We are integrating the transcriptional data with comprehensive molecular interaction networks and functional annotation databases to compute biological process linkage networks.
We expect that this approach will establish a combined experimental and computational pipeline for toxicity testing and risk assessment centered on 3D liver mimics and biological process linkage networks. Our approach allows perturbations in toxicity pathways to be traced across multiple scales: from regulatory proteins and transcriptional perturbations to communication between different hepatic cell types. Unique aspects of our approach include modeling the cell’s response to combinations of toxic chemicals and in more than one hepatic cell type.
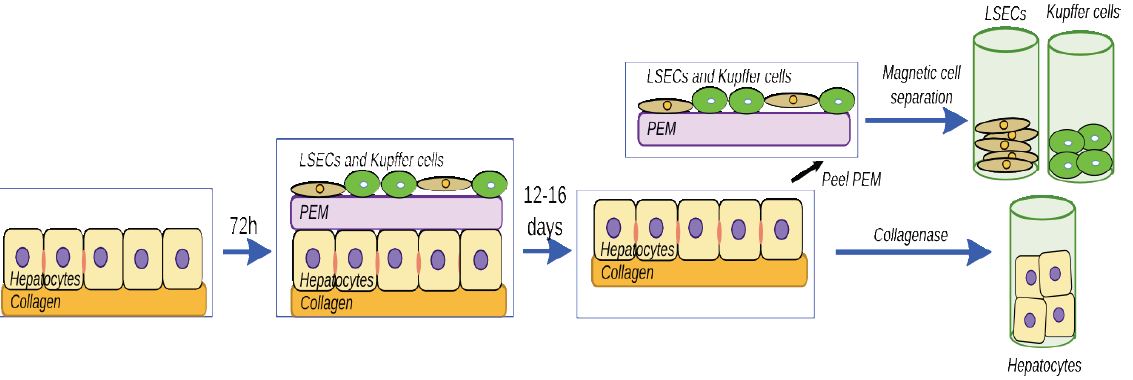
Figure 1. Process for assembling liver models and separating cells at the end of the culture period.
Key Results from Acetaminophen (APAP) Mediated Toxicity on 2D and 3D Hepatic Models
We have completed investigating the effects of APAP toxicity using 3D liver models that contain primary rat hepatocytes, liver sinusoidal endothelial cells (LSECs), Kupffer cells (KCs) and a polymeric Space of Disse (SoD).
Cultures Investigated:
i. Hepatocyte monolayer (HM)
ii. Collagen sandwich (CS)
iii. 2D co-culture of hepatocytes and LSECS (2DHL)
iv. 3D liver model with hepatocytes-polymeric SoD-LSECs (3DHL)
v. 3D liver model with hepatocytes-polymeric SoD-LSECs and KCs (3DHLK)
Timepoints at which measurements were taken:
i. APAP administration on day 4, measurements conducted 24h later.
ii. APAP administration on day 6, measurements conducted 24h and 48h later.
We have completed the following experimental work:
i. Conducted viability measurements using the MTT assay
ii. Measured the ratios of aspartate amino transferase/ alanine amino transferase (AST/ALT)
iii. Measured depletion of glutathione
iv. Conducted CYP2E1 western blots
v. Measured albumin secretion
vi. Measured urea secretion
Key Highlights of APAP-treated 2D and 3D cultures
Conventional 2D cultures such as hepatocyte monolayers (HMs) rapidly dedifferentiate and therefore do not exhibit sensitivity to this drug beyond 24h-36h. Although, collagen sandwich (CS) cultures are more stable, they do not appear to be sensitive to APAP at low concentrations. A dose response study for APAP from 10-40 mM was conducted for HM, CS 2DHL, 3DHL and 3DHLK. Only the 3DHL and 3DHLK models exhibited cell death at a concentration of 10 mM (in comparison to untreated samples) (Figure 2). In contrast, HM, CS and 2D cultures exhibited cell death only at a 40 mM concentration of APAP. These data demonstrate that the 3DHL and 3DHLK models are sensitive to APAP at concentrations (10 mM) relevant to rodent and human hepatotoxicity.
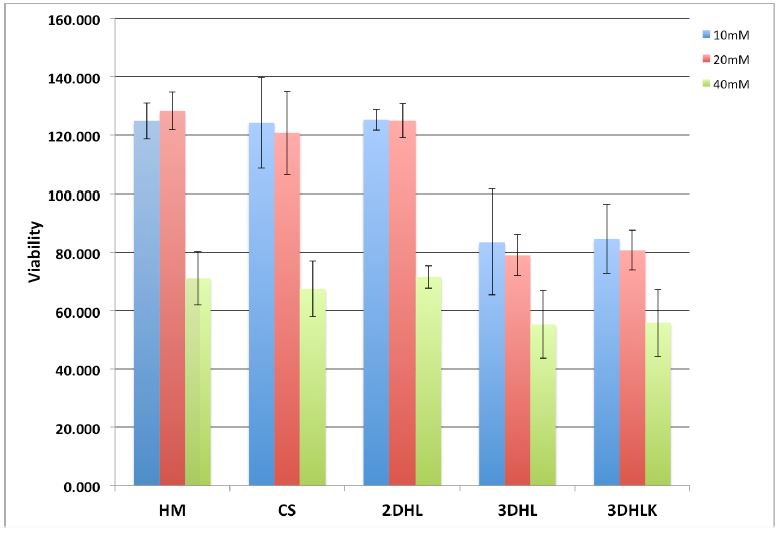
Figure 2. Viability data on 2D and 3D liver cultures. APAP administration on day 4, measurements conducted 24h later.
We measured the concentrations of alanine aminotransferase (ALT) and aspartate aminotransferase (AST) enzymes in spent culture medium in APAP treated samples. These enzymes are markers of hepatocellular death. An increase of 2-fold or higher in AST/ALT ratios is considered a marker of significant hepatotoxicity. The AST/ALT ratio is < 2.0 for 2D cultures (HM, CS, 2DHL) up to a concentration of 20 mM. In contrast, the 3D cultures exhibit AST/ALT ratios >2.0 at 20 mM. At 40 mM, these ratios range between 4.0-4.5 in 3DHL and 3DHLK cultures, respectively. These trends clearly indicate that the 3D liver models exhibit a significantly greater response to a hepatotoxicant such as APAP (Figure 3).
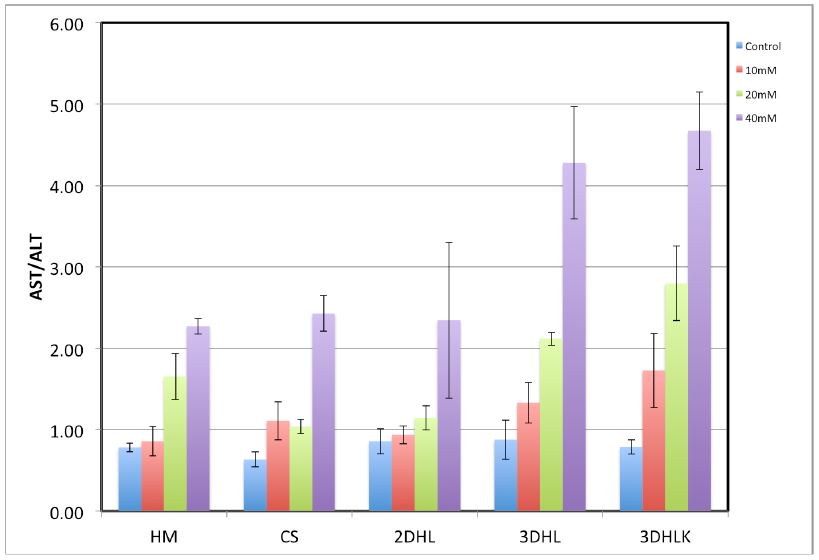
Figure 3. AST/ALT ratios in 2D and 3D liver cultures. APAP administration on day 4, measurements conducted 24h later.
Conducting toxicity studies in medium and high throughput assays.
We have assembled an innovative liver “organotypic culture model” (OCM) assembled in 96w microplates (denoted henceforth as μOCM) that will enable the simultaneous measurements of individual and cocktails of chemicals. Rajagopalan has assembled hepatic μOCMs cultured up to a 5-day period and tested the cytotoxicity of acetaminophen (APAP). In this experiment, hepatic μOCMs were assembled comprised of hepatocytes and liver sinusoidal endothelial cells (LSECs), denoted as 3DHL in (Figure 4). Approximately, 75,000 primary rat hepatocytes were seeded in each well. The biopolymeric COL/HS membrane was placed 72h later followed by approximately 3125 primary rat LSECs. Hepatocyte monolayers (HMs) also were cultured on the same 96w plate to serve as controls. APAP was administered at a high concentration (40 mM) on day 4 in culture with MTT measurements conducted 24h later. In (Figure 1), we demonstrate that hepatic μOCMs are sensitive when they are exposed to a 40 mM concentration of APAP whereas HM cultures do not exhibit significant cell death. The cell viability in HMs was 83.6% after APAP administration, whereas, in the 3DHL hepatic μOCMs, the viability dropped to approximately 36%. These experiments serve as preliminary proof of concept investigations that clearly demonstrate that hepatic μOCMs can be assembled and that they are sensitive to APAP. This encouraging result demonstrates the potential of hepatic μOCMs in toxicant testing.
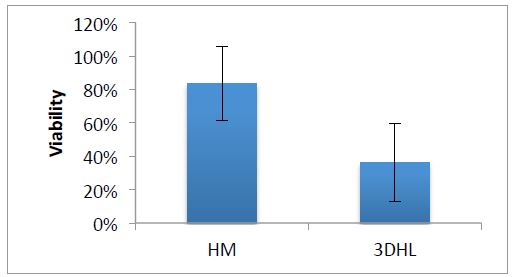
Figure 4. Viabilities of HM and hepatic μOCMs upon APAP administration.
We have preliminary data that demonstrates that liver μOCMs can be assembled in 96w plates, they are sensitive to APAP and urea secretion can be assayed through conventional biochemical methods (Figure 5).
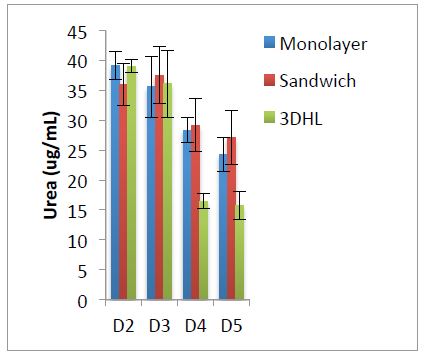
Figure 5. Urea secretion for hepatocyte cultures in 96w plates
Future Activities:
1. Complete transcriptomic measurements on APAP-treated 2D and 3D liver cultures.
2. Conduct computational analysis to determine APAP-perturbed pathways and networks of reactions.
3. Conduct experiments to determine hepatotoxicity of perfluoro octanoic acid (PFOA).
4. Complete the computational analysis of ToxCast™ data (collaboration with Dr. Imran Shah, USEPA).
Journal Articles on this Report : 10 Displayed | Download in RIS Format
| Other project views: | All 31 publications | 11 publications in selected types | All 10 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Bharadwaj A, Singh D, Ritz A, Tegge A, Poirel C, Kraivkivski P, Adames N, Luther K, Kale S, Peccoud J, Tyson T, Murali T. GraphSpace: stimulating interdisciplinary collaborations in network biology. BIOINFORMATICS 2017;33(19):3134-3436 |
R834998 (2013) |
Exit |
|
|
Larkin AL, Rodrigues RR, Murali TM, Rajagopalan P. Designing a multicellular organotypic 3D liver model with a detachable, nanoscale polymeric Space of Disse. Tissue Engineering Part C: Methods 2013;19(11):875-884. |
R834998 (2013) |
Exit |
|
|
Lasher CD, Rajagopalan P, Murali TM. Summarizing cellular responses as biological process networks. BMC Systems Biology 2013;7:68. |
R834998 (2013) |
Exit Exit |
|
|
Rajagopalan P, Kasif S, Murali TM. Systems biology characterization of engineered tissues. Annual Review of Biomedical Engineering 2013;15:55-70. |
R834998 (2013) |
Exit Exit |
|
|
Rita A, Poirel C, Tegge A, Sharp N, Simmons K, Powell A, Kale S, Murali T. Pathways on demand: automated reconstruction of human signaling networks. SYSTEMS BIOLOGY AND APPLICATIONS 2016;2(16002) |
R834998 (2013) |
Exit |
|
|
Sam SA, Teel J, Tegge AN, Bharadwaj A, Murali TM. XTalkDB: a database of signaling pathway crosstalk. Nucleic Acids Research 2017;45(D1):D432-D439. |
R834998 (2013) |
Exit Exit |
|
|
Tegge AN, Sharp N, Murali TM. XTALK: a path-based approach for identifying crosstalk between signaling pathways. Bioinformatics 2016;32(2):242-251. |
R834998 (2013) |
Exit Exit |
|
|
Tegge A, Rodrigues R, Larkin A, Murali T, Pagagopalan P. Transcriptomic Analysis of Hepatic Cells in Multicellular Organotypic Liver Models. NATURE 2018;8(11306) |
R834998 (2013) |
Exit |
|
|
Vu LT, Less RR, Rajagopalan P. The promise of organotypic hepatic and gastrointestinal models. Trends in Biotechnology 2014;32(8):406-413. |
R834998 (2013) |
Exit Exit Exit |
|
|
Vu L, Orbach S, Ray W, Cassin ME, Rajagopalan P, Helm R. The hepatocyte proteome in organotypic rat liver models and the influence of the local microenvironment. PROTEOME SCIENCE 2017;15(12) |
R834998 (2013) |
Exit |
Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
