Grantee Research Project Results
Final Report: Center for Integrative Research on Childhood Leukemia and the Environment
EPA Grant Number: R836159Center: Center for Integrative Research on Childhood Leukemia and the Environment - 2015
Center Director: Metayer, Catherine
Title: Center for Integrative Research on Childhood Leukemia and the Environment
Investigators: Metayer, Catherine
Institution: University of California - Berkeley
EPA Project Officer: Callan, Richard
Project Period: September 1, 2015 through August 31, 2019 (Extended to August 31, 2020)
Project Amount: $2,599,999
RFA: Children's Environmental Health and Disease Prevention Research Centers (2014) RFA Text | Recipients Lists
Research Category: Children's Health , Human Health
Objective:
Core A: The main objectives of Core A are to provide oversight, coordination, and integration of the Center for Integrative Research on Childhood Leukemia and the Environment (CIRCLE) activities; establish/manage an External Advisory Committee (EAC); coordinate community engagement, appoint and collaborate with the Pediatric Health Specialist and support the Career Development Investigator within the structure of the Center. Listed below are the detailed specific aims we pursued to achieve these objectives.
Core B - Community Outreach and Translation Core (COTC)
Core C – Mouse Model Core Facility: We tested the hypothesis that pre-natal exposures to chemicals that have been suspected of impacting the health of human children would alter immune development and accelerate the development of leukemia in mouse models of childhood leukemia. We also examined whether persistent effects on DNA methylation could underlie possible observed effects. Core C completed exposures of pregnant mice and analyses of the offspring in response to tobacco in the form of third hand smoke, to the PAH dibenz[a,h] anthracene, and to the insecticides Permethrin and Cypermethrin.
Project 1: The potential interplay between in utero chemical exposures and the immune status of both the mother and the child in the development of childhood acute lymphoblastic leukemia (ALL) has not been studied. We hypothesize that specific in utero chemical exposures will impact maternal and neonatal immune status and increase the risk of childhood ALL. To test this hypothesis, we will leverage existing independent resources from two NIH-funded studies: the California Childhood Leukemia Study (CCLS, a population-based case-control study), and the California Mother-Child Birth Cohort (CA Birth Cohort, a cohort of children diagnosed with leukemia, healthy children without leukemia, and their mothers).
Project 2: We employed exposomics to characterize endogenous and exogenous chemical exposures in archived neonatal blood spots (ANBS) and maternal-pregnancy serum to discover potential in utero risk factors for childhood acute lymphoblastic leukemia (ALL). The untargeted omics methods employed were metabolomics, to characterize small molecules, and Cys34 adductomics, to characterize exposures to reactive electrophiles bound to human serum albumin (HSA). We also performed targeted analyses of small molecules and Cys34 adducts that represent biomarkers of exposure to factors that have been previously associated with childhood ALL.
Project 3: In an effort to understand the rising leukemia rates in children, we propose to assess the interplay between environmental chemicals, immune status, and DNA methylation in childhood ALL development, taking into account a comprehensive constitutive genetic assessment. We leveraged existing resources from two NIH-funded studies, the California Childhood Leukemia Study (CCLS, a population-based case-control study), and the California Mother-Child Birth Cohort (CA Birth Cohort, a cohort of children diagnosed with leukemia, healthy children without leukemia, and their mothers). Additionally, we tested the effect of in utero chemical exposures on DNA methylation in pre-B cells (our target cell population), as well as on leukemogenesis, using an engineered mouse model for the first time.
Summary/Accomplishments (Outputs/Outcomes):
Administrative Core A:
Aims 1 and 2. Assist the Center Director, Dr. Catherine Metayer, in the coordination and integration of the CIRCLE’s components and activities. Promote and assess productivity, effectiveness, and appropriateness of CIRCLE activities, and facilitate scientific opportunities and areas for collaboration among all CIRCLE members.
Core A has supported all administrative- and research-related activities in CIRCLE, including working closely with campus post-award unit to monitor the complex book-keeping of 12 chartstrings and 7 subawardees split between NIEHS and EPA. The IRB renewal protocol for CIRCLE (#15-09-2232) was approved by the California State Committee for Protection of Human Subjects (CPHS) and the UC Berkeley CPHS received approval for an inter-institutional reliance agreement to the CA State CPHS. The CIRCLE website was created in Year 1 and updated regularly (http://circle.berkeley.edu/). The Internal Advisory Committee met every two months to provide updates on all Cores and Projects, and to facilitate scientific interaction between CIRCLE investigators. Minutes were taken to document these discussions.
Core A managed the selection and acquisition of the biospecimens needed for all 3 Projects, and ensure sufficient overlap for future integration of the data generated on cytokines (Project 1), exposome and protein adducts (Project 2), and genetic/epigenetic (Project 3). The Center Director and co-Director were in regular contact with the California Biobank Program Coordinator (Robin Cooley) at the Genetic Disease Program to obtain the archived newborn blood specimens (ANBS) and maternal pregnancy blood samples. We have received all the ANBS from the CCLS participants. Regarding the CA mother-child cohort, we obtained maternal pregnancy blood and platelet samples for 137 cases and ANBS for their corresponding children, as well as maternal and child specimens for 411 controls. The initial linkage between the maternal and child biobanks yielded less samples than expected for cases (which was 200 mother/child pairs for cases). The linkage that was updated towards the end of the study period, will provide additional resources for future follow-up studies.
Aim 3. Organize CIRCLE activities, invitation of consultants, meetings, and focus groups. Core A supported the activities of the Community Outreach and Translation Core B, specifically the planning of meetings and calls with the Community Advisory Group (See Details in Core B report). The Center Director, the Pediatric Health Specialist, and other CIRCLE investigators worked with Core B leader Mark Miller on various outreach activities to the clinical audiences, general public, and Latinx communities that are the most affected by childhood leukemia. The CIRCLE Director who was also the Chair of the Childhood Leukemia International Consortium (CLIC) provided opportunities for Dr. Miller to continue to work on research translation with an international group of childhood leukemia researchers. Dr. Metayer organized the 2017 CLIC meeting with the Texas Children Hospital, in a joint meeting on Childhood Cancer and Health Disparity. We also partnered with the National Cancer institute and other federal agencies to update their messaging on childhood leukemia prevention. Core A also supported the attendance of senior and junior CIRCLE investigators to the Children’s Center annual meetings. The CIRCLE Director presented CIRCLE work at many venues over the past years as listed in the Executive Summary.
Aim 4. Organize and manage the External Advisory Committee. We formed an External Advisory Committee (EAC) with expertise that complemented research performed in CIRCLE. The EAC Members were Rodney Dietert from Cornell University (immunotoxicology), Kari (Christine ) Nadeau from Stanford University (pediatrics, immunology, and epigenetics), Raphael Gottardo from the Fred Hutchinson Cancer Research Center (Biostatistician with expertise in vaccine and infectious disease), Sandy (Alexander) Borowsky from UC Davis (pathology and mouse modeling), Daniel Nomura from UC Berkeley (metabolomics), and Jill Johnston from the University of Southern California (research translation and outreach). We held two EAC meetings in 2017 and 2019. The feedback from the EAC Members was overwhelmingly positive for all Cores and Projects, as well as for the integration of all components. Detailed reports were provided after the meeting.
Aim 5. Support the professional advancement of a new Career Development Investigator. Core A supported the professional advancement of Dr. Todd Whitehead, the Career Development Investigator providing exposure to all scientific and administrative components of research. Of note, Dr. Whitehead was successfully awarded two research grants during his tenure in CIRCLE. Core A also supported the mentoring of several undergraduate and graduate students and post-doctoral fellows in all Projects and Cores, who have launched their research careers in prestigious academic and private institutions.
Core B - Community Outreach and Translation Core (COTC)
Aim 1: Develop curricula for health care professionals communicating current research findings about childhood leukemia and the environment. COTC staff along with key researchers have presented at many regional and national medical conferences, via webinars. Key concepts related to children’s environmental health and childhood cancer risk have been disseminated via the “Story of Health” e-book. A special focus was in adapting these materials specifically for nursing, public health nursing, and nurse practitioner/midwife audiences.
The COTC developed an organizing principle to discuss risk factors for childhood leukemia in the context of the spectrum of risks identified with particular chemicals or classes of chemical. We have used this to promote the concept that we as a society should add a focus on prevention of childhood cancer to the current focus on treatment. This idea forms the basis of multiple talks by Drs. Miller and Metayer, the Story of Health chapter on childhood leukemia, a video developed in conjunction with “Little Things Matter” called “Cause or Cure” (https://wspehsu.ucsf.edu/projects/childhood-leukemia-environment/), and peer reviewed journal publications.
Dr. Miller attended a meeting at the National Cancer Institute (NCI), along with Drs. Metayer and Wiemels, to enhance collaborations between the Childhood Leukemia International Consortium (CLIC) and International Consortium of Child Cancer Cohorts (I4C) and presented a talk on prevention which was a key agenda item. A work group formed at this meeting developed a revision of the NCI children’s cancer fact sheet that now includes discussion of environmental risk factors and is posted on the NCI’s website (https://www.cancer.gov/types/childhood-cancers/child-adolescent-cancers-fact-sheet).
The COTC worked with NGOs to promote children’s environmental health literacy (with particular focus on risk factors for childhood leukemia) such as Physicians for Social Responsibility, Collaborative for Health and the Environment, and Cancer Free Economy Network.
The Story Of Health (SOH) e-book has been an outstanding success. We have continued working with the Western States PEHSU, ATSDR, California EPA OEHHA, and other partners to develop and promote of the five chapters of the SOH multimedia eBook (on infertility and reproductive health). The fifth chapter was released in October 2019 on Lifecourse Exposures and Cognitive Decline. So far, SOH has had over 13,000 registrations for physician, nurse, and health educator continuing education (CE) credit representing approximately 17,000 hours of CE credits to be awarded by the CDC. Surveys of users confirm exceptionally high satisfaction rates for the program. We developed a low literacy version of the SOH, in an innovative comic book format in both English and Spanish, for use in our various programs including seminars for Promotoras de Salud, and parents and staff in early educational environments. Approximately 10,000 hard copies were distributed by our COTC as well as various other NGOs, Children’s Centers, etc. The COTC produced an innovative “Shadow Puppet Theater” short video titled “Love in the Time of Toxicants”. This provides an alternative format for introducing the ideas in the Story of Health materials for a general community audience. It is available in English and Spanish and has been well received in the US as well as Mexico (https://wspehsu.ucsf.edu/projects/improving-environmental-health-literacy-young-adults/).
Aim 2: Establish a community advisory board that will be representative of the various communities with whom we are partnering. The COTC has established a distinguished community advisory group that held regular meetings to provide helpful input to our program. Representatives included a health educator from the California Department of Public Health, a community activist from an environmental justice community along the Mexican border, a physician member of the Kaiser Hispanic physicians organization, a member of a non-profit that funds research in childhood leukemia as well as parent and community outreach (who is also a parent of a child with leukemia), a staff member from an NGO that targets education of the public on risk factors and preventive activities related to breast cancer, and a leader of another COTC the CERCH’s Children Center with extensive experience working with the Latino farmworker community. At times we had guest attendees who contributed their expertise, for example, a pediatric oncology nurse practitioner and a pediatric oncology fellow from Stanford. The Community Advisory Committee met annually, once in person and another time via internet, and provided helpful direction and feedback on our outreach and materials development and volunteered to help in a variety of concrete ways.
Aim 3: Develop novel training materials in collaboration with Commonweal, CIRCLE investigators, and the Administrative Core that will be used to conduct train-the-trainer sessions with the aim of introducing children’s environmental health into existing pre-marital and pre-family programs for such audiences as religious organizations, community and public health nursing, colleges, and others.
As noted above, we developed a low literacy version of the SOH, in a comic book format, for use in our various anticipated programs. This was released in May 2017 in online and print versions in both English and Spanish. These are being used in our outreach programs, particularly in our trainings for Promotores de Salud. We have produced an innovative “Shadow Puppet Theater” short video titled “Love in the Time of Toxicants”. This provides an alternative format for introducing the ideas in the Story of Health materials for a general community audience. It was shown at the American Public Health Associations annual meeting in 2017 and was very well received. It is available in English and Spanish and has been well received in the US as well as Mexico. The comic and video address preconceptual and prenatal exposure risks in a story format designed to appeal to young adults. Vickie Leonard of the COTC has developed two powerpoint slide sets (different lengths) for use in trainings. These were designed to be used (and are currently being used!) by promotores and others in their outreach and are available in English and Spanish versions. Comments from our targeted audiences for the low literacy Story of Health Materials (Love in the Time of Toxicants video and Carlos and Rosa Get Married comic book) are below. The materials are viewable here https://wspehsu.ucsf.edu/projects/improving-environmental-health-literacy-young-adults/ and at https://circle.berkeley.edu/translation-and-outreach/for-parents/.
Other materials developed by the COTC include two innovative microsites. One, Carlos and Maria Plan a Family, features a series of infographics on childhood exposures and risk for childhood leukemia. The other, Dirty Little Secrets about House Dust, examines household exposures of contaminants attached to dust particles and actions to limit children’s exposure (https://circle.berkeley.edu/translation-and-outreach/for-parents/). A series of five engaging infographics raising awareness (in English and Spanish) of risk factors for childhood leukemia were produced in the past year. The materials described above were released and are available in Spanish and specifically address the needs of the Latinx community. They have been used in numerous workshops presented to parents, promotores de salud (lay health workers), childcare workers, and at health fairs. Our materials are being used in programs along the border in Texas by promotora and medical and nursing student training sponsored by the El Paso based PEHSU.
Aim 4. Develop specific outreach approaches and materials for the Latino community in collaboration with Commonweal and identify and obtain both traditional and non-traditional media coverage of current childhood leukemia findings. The COTC in collaboration with CIRCLE researchers and the California Cancer Registry identified that the rate of childhood leukemia among Latinx children in California has been rising at a higher rate than among white children. CIRCLE research also identified preconception, prenatal, and early childhood as sensitive time periods for exposures associated with increased risk. This formed the rationale for specifically targeting education of Latinx families with messages relevant to these time periods.
Non-traditional outreach includes an ongoing e-newsletter (English and Spanish) for Promotoras de Salud, early childhood workers, and other health workers. We have published 7 of these so far for a regular audience of 165+. In our 2019/2020 year we have established a collaboration with the PEHSU program in region 6 which will expand the outreach to that border region. Other non-traditional outreach has included presentations on many webinars targeted to both professional and lay populations throughout the cycle. Traditional media coverage has included Dr. Metayer being featured in a PBS documentary “On the Front Lines of Cancer”.
The materials described in aim 3 above were released and are available in Spanish and specifically address the needs of the Latino community. They have been used in numerous workshops presented to parents, promotores de salud (lay health workers), child care workers, and at health fairs. This is part of our program to “Improve the Environmental Health Literacy of Young Adults in which CIRCLE, Commonweal, and the Western States Pediatric Environmental Health Specialty Unit (PESHU) collaborate.
For 3 years we were able to recruit Jose Camacho, a health educator with 20 years of experience working with the Latino community, to conduct a number of workshops as well as to include dissemination of our materials at health fairs and other events that he attends regularly in other capacities. He hass done a superb job and is extremely well liked by the participants. Twice CIRCLE had booths at the annual conference of Vision y Compromiso and once were able to present a session at this major conference for promotoras. We also presented a session at the XVII Binational Promotoras Conference at the UC Berkeley School of Public Health
Examples of additional outreach events included (1) Members of the COTC group met with leaders in the Latino community working with adolescents and young adults in Richmond to solicit advice on outreach; and (2) Workshops presented at the Tiburcio Vasquez Health Center (serving primarily and immigrant Latinx community), Planned Parenthood (for Spanish speaking staff), county based First Five Centers, etc.
Aim 5. Collaborate with California EPA, Office of Environmental Health Hazard Assessment and the UCSF Pediatric Environmental Health Specialty Unit to conduct symposia that will update state, federal, and other scientists and the public on current research findings of the Children’s Environmental Health Research Centers. CIRCLE joined with California, EPA, OEHHA and the WSPEHSU to co-sponsor our annual children’s environmental health symposium series. In person and online attendance has averaged about 300+ state and federal scientists and policy makers, stakeholders, and others each year. The meetings are also viewed in a non-synchronous fashion. In 2017 we had a first ever meeting specifically on “Environmental Justice and Children” (https://oehha.ca.gov/risk-assessment/presentation/2017-childrens-environmental-health-symposium-presentations). This was the first ever symposium to focus specifically on this issue and is relevant to our CIRCLE COTC aims as many of the EJ community exposures of concern are also linked to childhood leukemia risk. Symposia are archived and include:
- Air Pollution and Life Course Neurological Impacts (https://oehha.ca.gov/risk-assessment/presentation/2018-childrens-environmental-health-symposium-presentations)
- New Findings from California (https://oehha.ca.gov/environmental-justice/agenda-background/2016-childrens-environmental-health-symposium)
In summary, the COTC has accomplished the range of goals set out at the beginning of the cycle and far more than we had anticipated. An overarching goal that arose from our initial plan was to advance the idea that prevention of childhood leukemia needs to be given additional importance beside the current emphasis on treatment. When CIRCLE started to advance these ideas there were no other organizations or governments agencies with programs discussing this. Over the last several years in addition to increased interest at NCI, advocacy groups such as the Cancer Free Economy Network, The Reason Why, Children’s Environmental Health Network and others have developed programs aimed at raising awareness of this issue and developing effective responses.
Core C – Mouse Model Core Facility:
Aim 1: Design means, doses, and timing of in utero exposures for mouse model experiments.
Aim 2: Obtain plasma samples from exposed and control animals for analysis of immunomodulatory cytokines (samples obtained by the Core for analyses performed in Project 1).
Aim 3: Generate DNA samples from blood cells and from immature B-cells in bone marrow of exposed and control animals for analyses of DNA methylation (samples obtained by the Core for analyses performed in Project 3).
Aim 4: Analyze the impact of exposures on immune cell numbers, including pre-leukemic immature B-cells (analyses performed by the Core for Projects 1 and 3).
Aim 5: Assess the impact of exposures on leukemia development (analyses performed by the Core for Projects 1 and 3).
What was accomplished under these goals:
Third Hand Smoke (THS): Third hand smoke has been recognized as an important mechanism through which children are exposed to tobacco carcinogens. We hypothesized that exposure to THS during pregnancy and following birth would contribute to ALL in our mouse models. Exposure regimen consisted of exposing pregnant dams from time of mating until weaning of pups to THS exposed or control cotton terrycloth, one exposed swatch of cloth each week for 6 weeks. In order to limit the development of sarcomas as a competing risk, given our use of Cdkn2a null animals (susceptible to leukemia and to sarcoma), we transplanted bone marrow from THS exposed animals into histocompatible irradiated recipient mice, and followed these animals for disease. In order to understand how THS might impact immune development we also examined plasma cytokine levels at 2 days of age, as well as blood counts and immune cell numbers at 5 weeks of age. These early endpoints could provide insight into mechanisms through which such pre-natal chemical exposures may influence the health of offspring.
We found that in utero and early life THS exposure caused significant alterations in blood parameters including broad reductions in plasma cytokine concentrations and alterations in immune cell populations in blood, spleen and bone marrow, which appeared more pronounced in male mice. Spleen and bone marrow B-cell populations were significantly lower in THS exposed mice. Although we did not observed accelerated leukemia following THS exposure, the increased pace of development of solid tumors in our irradiated recipient animals suggests that the immunosuppressive effects of in utero and early life THS exposure may contribute to carcinogenesis by lowering the host defense to other exposures. These data extend our knowledge of an important mechanism through which pre-natal chemical exposures can influence child health: by impairing the development of the immune system. Although our experiments did not demonstrate the ability of these exposures to accelerate leukemia development, this negative finding is limited to the specific conditions of our studies which reflect a particular mouse genetic background, a specific genetic lesion, and a particular exposure.
Two additional lines of investigation we pursued in this model were in the areas of methylation and mutation spectrum. In the first of these lines of investigation, whole genome DNA methylation analysis of blood cells from THS exposed mice at 5 weeks of age were compared to the DNA methylation patterns of control exposed animals. The methylation patterns did not appear to be substantially different under these experimental conditions: in non‑supervised hierarchical clustering of the methylation data the animals did not separate out by exposure group. In the second additional investigation we postulated that even though THS exposure did not accelerate leukemia development, there might be differences in the pathway to leukemic transformation as a result of THS exposure. We selected leukemias that arose following THS exposure and performed low-depth DNA sequencing of the tumor tissue, in order to identify regions of chromosomal gain and loss. We compared the gains and losses to those observed in leukemias following control exposure. Although we detected chromosomal gains and losses, similar to those previously observed in this model, and the study involved small number of leukemias (n=4 for each group) we did not observe significant differences in the extent of copy number changes between the exposed and non-exposed groups.
Polyaromatic Hydrocarbons (PAHs): For the PAH Dibenz[a,h]anthracene our exposure protocol included 1.5 mg/kg exposures every other day during pregnancy for a total of 5 doses. Study design otherwise paralleled what is described above for THS. This PAH study was largely a negative study, with only minor impacts on cytokine levels at 2 days of age, animal weights at weaning, blood counts and immune cell numbers at 5 weeks of age, and DNA methylation at 5 weeks of age. No changes in leukemia development were observed.
Insecticides: Cypermethrin and permethrin dosing regimen included a combination of 35 mg/kg cypermethrin + 200 mg/kg permethrin each day for a total of 10 days during pregnancy, and again paralleled what is described above for THS. Pre-natal insecticide exposure caused profound suppression of cytokine levels at 2 days of age, and modest differences in immune cell populations at 5 weeks of age. No changes in leukemia development were observed.
Quality Assurance: 1) Animal experiments for a given exposure were performed over a period of months with multiple different breeders and offspring. 2) Both male and female offspring were studied. And 3) Each experimental animal was independent. Only a single male and a single female animal was studies at a particular time point from each exposed dam. Only a single male and a single female animal were followed for disease from each exposed dam. Only a single transplant recipient was generated from each of these donor mice.
Summary: Our results extend previous observations regarding the negative impacts on the immune system (immune suppression, altered immune cell numbers) of pre-natal exposures, including exposures to tobacco and particular pesticides. In fact, in our studies we showed that these two pre-natal exposures caused profound depression of immune regulatory cytokines at birth, with potential for long-term impact on child health. Although our experiments did not demonstrate the ability of these exposures to accelerate leukemia development, this negative finding is limited to the specific conditions of our studies which reflect a particular mouse genetic background, a specific genetic lesion, and a particular exposure. The observed negative immune impacts of pre-natal exposure to environmental toxins raise questions of how such exposures influence human health in ways other than leukemogenesis, including response to infections and risk of allergies and autoimmunity
Project 1:
Aim 1: Evaluate the impact of in utero chemical exposures and neonatal immune status at birth on childhood ALL risk, in the California Childhood Leukemia Study (CCLS). Quality Assurance: In order to have the most sensitive assays for neonatal blood spot extracts, which are difficult to work with, we performed additional pilot studies using a variety of immunoassay platforms to increase sensitivity of the assays. These pilot assays used Meso Scale Discovery, Simoa, and Luminex platforms. Our work in this area led to modifications in our protocol for Aims 1 and 2 as follows: we extract archived blood spots in half the volume as prior work to get more concentrated proteins. We used the Curiox plate washer system to allow for smaller volume incubations and cleaner washes. We now use an ultra-sensitive Luminex assay (R&D Systems) instead of Millipore or Bio-rad assays, as the R&D is more sensitive. With these changes, we were able to assay all of our blood spots.
In line with Aim 1 we evaluated the interplay between levels of immunomodulatory cytokines at birth, prenatal environmental exposures (including small-molecule features and human serum albumin adducts measured in Project 2), and the risk of childhood acute lymphoblastic leukemia (ALL). Nine cytokines – interleukin 1 (IL1), IL4, IL6, IL8, IL10, IL12p70, granulocyte-macrophage colony-stimulating factor, tumor necrosis factor alpha (TNFα), and vascular endothelial growth factor (VEGF) – were measured in blood spots collected at birth from 1,020 childhood ALL cases and 1,003 controls participating in the California Childhood Leukemia Study.
Cytokines were related to several demographic and birth characteristics. Cytokine levels were negatively correlated with the timing of blood collection, suggesting that concentrations decreased in the hours and days after birth. Cesarean delivery, premature gestational age, and low birth weight were also linked to lower cytokine levels. Concentrations of IL1β, IL4, IL8, and IL10 differed by mother’s race/ethnicity – with newborns of non-Latina white mothers having the lowest levels. Newborns from high-income households tended to have lower levels of IL1β, IL6, and IL8 in their blood at birth than newborns from low-income households.
Levels of all cytokines were correlated with each other at birth, with Spearman rank correlation coefficients of 0.10 or more and all p-values < 0.05. To accommodate the correlation between cytokines in risk analyses, we used principal component analysis to reduce the dimensionality of the data from nine individual cytokines to three principal components (PCs). These three PCs explained 69% of the variance in cytokine levels. PC1 was a linear combination of the nine individual cytokines, loaded most heavily by IL8, IL1β, VEGF, and TNFα.
Odds ratios associated with an interquartile range increment in cytokine levels were calculated using logistic regression adjusting for sociodemographic and birth characteristics. Higher levels of PC1 were consistently associated with increased odds of childhood ALL using several different modeling approaches. We also found that ALL patients were born with higher levels of a group of correlated cytokines – IL1β, IL8, IL10, IL12p70, TNFα, and VEGF – than controls. Stratified regression analyses were conducted by race/ethnicity, ALL subtype, diagnosis age, birth year, and sex. These associations were observed among children of Latina mothers, but not among children of non-Latina white mothers, and were seen most dramatically for ALL with high hyperdiploidy.
We used a mediation analysis (mediate in R) to evaluate whether cytokines were mediating the relationship between prenatal environmental exposures and ALL risk. We found that neonatal cytokine levels were correlated with neonatal levels of small molecule biomarkers which had been previously associated with ALL risk (in Project 2). There were 16 positive links between levels of one of the 9 cytokines at birth and levels of one of the prenatal environmental exposures, with Spearman correlation coefficients from +0.15 to +0.30. Of these 16 strongest positive relationships, 11 involved a small-molecule biomarker measured at birth in the same blood spots (most commonly an unknown metabolite with putative molecular formula of C26H44O2). There were 36 negative links between a cytokine and a prenatal environmental exposure, with correlation coefficients ranging from -0.15 to -0.33. Of these 36 strongest negative relationships, 34 involved a small-molecule biomarker (most commonly a glycerophospholipid, a sphingolipid, and two unknown molecules measured as part of Project 2). However, there was no evidence that the cytokines were mediating the relationship between these small molecules and ALL risk.
We posit that children who are born with altered cytokine levels are set on a trajectory towards an increased risk for subsequent aberrant immune reactions that can initiate ALL (manuscript submitted to Cancer Epidemiology Biomarkers and Prevention).
In complement to the in utero immune-related hypothesis that is the focus of Project 1, we have completed analyses related to immune function (--at birth and after birth--) and childhood ALL: We investigated the association between self-reported child’s allergies and the risk of childhood ALL, using CCLS subjects. Overall, no associations were observed between childhood ALL risk and specific allergy phenotypes or any allergy, as a group. However, having any allergy was associated with a younger age at diagnosis (-0.66 years, 95% CI: -0.02, -1.31). We also conducted meta-analyses with data from the CCLS and 14 previously published studies. In random-effect models, a reduced odds of ALL was associated with hay fever (meta OR=0.66, 95% CI: 0.48, 0.91); however, restricting the analysis to studies that used medical records for assessment of allergy or recently published studies, led to null or attenuated results. Overall, our analyses suggest a lack of association between allergy diagnosed after birth and childhood ALL. The limitation of self-reports may explain this negative finding, and further support the need of biomarker studies to better characterize immune profile. A manuscript was published in a peer-reviewed journal (Wallace AD, et al., Cancer Epidemiol Biomarkers Prev. 2018 Oct;27(10):1142-1150).
Aim 2: Evaluate the impact of in utero chemical exposures, maternal immune status during pregnancy, and neonatal immune status at birth on childhood ALL risk, in the CA Birth Cohort. We have pursued Aim 2 as originally proposed and made steady progress. First, we have collaborated with investigators from the Berkeley and San Francisco campuses of University of California to ensure that appropriate blood samples of subjects (cases, controls, and their biological mothers) are processed and measured for immunomodulatory cytokines.
Second, we have analyzed the cytokine profiles of 137 cases of childhood acute lymphoblastic leukemia (ALL) and 499 control subjects matched to the cases on month and year of birth, sex, and race/ethnicity. The cytokines included IL-1β, IL-2, IL-4, IL-5, IL-6, IL-8, IL-10, IL-12p70, GM-CSF, INF-γ, TNF- α, and VEGF. After an initial evaluation, three of the cytokines (IL-2, IL-5, and INF-γ) were removed from further analyses, because more than 50% of study subjects had levels that were non-detectable. For the remaining nine cytokines, we assigned half of the detection limit to subjects whose levels were not detectable. We then evaluated whether the level of these cytokines was associated with a number of factors, including year of birth, race (white, Hispanic, other), sex, age at the collection of blood spots (in hours, continuous variable), birth weight (<2500, 2500-2999, 3000-3499, 3500-3999, ≥4000 grams), gestational age (37, 37-41, >41 weeks, or missing), plurality (singleton/multiple birth), birth order (1, 2, ≥3), mode of delivery (vaginal, C-section), mother’s age at delivery (<25, 25-34, ≥35 years), and mother’s birth place (US, other). These evaluations informed our choice of covariates for inclusion in the multivariable model. As we observed high correlations across the cytokines and were concerned about collinearity, we utilized the principle component approach to derive six components that are independent from each other.
Since the first three components explained more than 72% of the variance, we decided to focus on these components. Interestingly, each of the components was dominated by unique cytokines. For example, Component 1 was heavily influenced by IL-6 and IL-8, while IL-4 and IL-12p70 appeared to play an important role in Component 2. When we compared the components between cases and controls, both Components 1 and 3 were significantly associated with the risk of childhood ALL (see table below). While it is premature to draw conclusions at this point, it appears that our hypothesis regarding the role of immunomodulatory cytokines in leukemogenesis is supported by the current data. In addition, it may be important to assess subsets of cytokines that operate through different mechanisms in immune modulation.
| Case (%) | Control (%) | OR (95% CI) | P value | |
| Component 1 | 61(44.5) | 250(50.1) | 1.00 | |
| 76(55.5) | 249(49.9) | 1.52 (1.01-2.30) | 0.04 | |
| Component 2 | 63(46.0) | 250(50.1) | 1.00 | |
| 74(54.0) | 249(49.9) | 1.19 (0.80-1.77) | 0.38 | |
| Component 3 | 55(40.1) | 250(50.1) | 1.00 | |
| 82(59.9) | 249(49.9) | 1.66 (1.11-2.50) | 0.01 |
Table 1 Cytokines and Risk of Childhood ALL
Arginase 2 (ARG2) is a key regulator in perinatal immunity, suppressing T cells through an anti-inflammatory cascade resulting from arginine depletion. Emerging evidence suggests that susceptibility to neonatal infections is related to active immune suppression within the neonatal environment. Given the function of ARG2 in neonatal immune function and response to early-life infections, we also investigated whether variation in ARG2 levels at birth may be associated with risk of developing ALL in childhood. We utilized the neonatal blood spots from the 137 cases and 500 cancer-free control children from Project 1, Aim 2. The multivariable analysis suggested that the risk of childhood ALL increased by more than two-fold in subjects whose level of ARG2 at birth was in the third or fourth quartile, compared to those whose ARG2 level was in the lowest quartile. This finding suggests that immune changes related to ARG2 levels are evident long before the clinical manifestation of childhood leukemia. This novel finding increases evidence for a role of immune dysregulation at birth in the development of childhood ALL. The results of the analysis were published in a peer-reviewed journal (Nielsen AB, et al., Haematologica 2019 104(11):e514-e516)
In complement to the in utero immune-related hypothesis that is the focus of Project 1, we have completed analyses related to immune function (--at birth --) and childhood ALL: We have analyzed data from the California Childhood Cancer Record Linkage Project (study population from which the California Mother-Child Birth Cohort is derived) and found that cesarean section (C-section), especially elective C-section, increases the risk of childhood ALL (manuscript published in the American Journal of Epidemiology; see Publications section below for detail). This study underscored the contrast between emergency and elective C-section in terms of their impact on risk of childhood ALL, which is likely related to the different patterns of exposure to microbiome in the birth canal. As the prevalence of C-section in the United States increased from 5.8% in 1970 to 32.9% in 2009 and most of the increase was due to elective C-section, this finding has important public health implications. It may be possible to target elective C-section as a modifiable risk factor for childhood ALL, the incidence of which has continuously risen in the United States over the last few decades. For example, there are ongoing clinical trials evaluating the possibility of deliberately exposing C-section-delivered newborns to maternal microbiome. An original manuscript was published in a peer-reviewed journal. Results were consistent with a pooled analysis conducted by the Childhood Leukemia International Consortium, including CIRCLE investigators (Wang R, et al. Am J Epidemiol. 2017 Jan 15;185(2):96-105; Marcotte EL, et al., Lancet Haematol. 2016 Apr;3(4):e176-85). The full list of publications is available in the executive summary.
Aim 3: Elucidate the mechanism(s) by which in utero chemical exposures and maternal/neonatal immune status interact to initiate ALL in an engineered mouse model with a propensity to develop a mouse-analog of childhood ALL. During the period of this grant we have aimed to utilize a mouse model of childhood ALL to elucidate mechanisms by which in utero chemical exposures and maternal/neonatal immune status interact to influence ALL development. To this end we have evaluated the impact on this animal model of pre-natal exposure to polychlorinated biphenyls (Aroclor 1260), tobacco (tobacco smoke residue/third hand smoke), pesticides (permethrin/cypermethrin), and a polyaromatic hydrocarbon (dibenz [a,h] anthracene). In regard to the mechanisms through which such pre-natal chemical exposures may influence the health of offspring, we observed that tobacco smoke residue and pesticides cause substantial effects on the cytokine profiles of offspring assessed during the second day of life, and that persistent changes in immune cell populations were present in animals at 5 weeks of age. Hence, these data extend our knowledge of an important mechanism through which pre-natal chemical exposures can influence child health: by impairing the development of the immune system. Although our experiments did not demonstrate the ability of these exposures to accelerate leukemia development, this negative finding is limited to the specific conditions of our studies which reflect a particular mouse genetic background, a specific genetic lesion, and a particular exposure. Three manuscripts describing cytokine levels in 2-day pups following prenatal exposure to tobacco, permethrin/cypermethrin, and dibenz [a,h] anthracene are in preparation and will be submitted for publication in peer-reviewed academic journals.
Project 2:
Aim 1: Finalize methods for profiling small molecules and Cys34 adducts in ANBS. Quality Assurance: We developed untargeted methods for extracting small molecules and Cys34 adducts from ANBS and then measuring these chemicals via metabolomics (Petrick et al. 2017) and adductomics (Yano et al. 2019) in control children from the California Childhood Leukemia Study (CCLS). Each set of untargeted analyses was performed with 4.7-mm punches from ANBS and included measurement of potassium and/or hemoglobin (Hb) to facilitate normalization of analyte levels for hematocrit content. Of the myriad small molecules that were tested across 106 control children, significant associations were detected with ethnicity (three metabolites) and birth weight (15 metabolites) after adjusting for multiple testing (Petrick et al. 2017). The adductomics method was validated with 49 archived DBS collected from newborns whose mothers either actively smoked during pregnancy or were nonsmokers. Using an ensemble of linear and nonlinear models, the Cys34 adduct of cyanide was found to be consistently elevated in ANBS from children of smoking mothers (Yano et al. 2019).
Aim 2: Measure and annotate “omic” features in ANBS from childhood ALL cases and matched controls. We characterized two sets of omic features in ANBS, i.e. small molecules (metabolomics) and HSA adducts (adductomics). The small-molecule and adduct features were measured and annotated in ANBS from 400 CL cases and matched controls from CCLS (Petrick et al. submitted; Petrick et al. 2019; Yano et al. 2020) and with both ANBS and maternal blood from 200 cases and matched controls from the mother-child cohort.
Aim 3: Process and compare data from cases and controls to find discriminating “omic” features. We developed processing pipelines to increase throughput of omics workflows (Edmands et al. 2017; Edmands et al. 2018; Schiffman et al. 2019) and a bioinformatic pipeline with which we compared omic features between 200 pairs of CCLS childhood leukemia cases and controls (Petrick et al. submitted; Petrick et al. 2019; Yano et al. 2020). Regarding comparisons of small molecules between ALL cases and matched controls, feature selection pointed to potential roles of several lipids in the etiology of ALL, including linolenic and linoleic acids and phospholipids with 18:2 and eicsoatetraenoic acid moieties. Interestingly, many of the selected metabolites were highly associated with breastfeeding practice in children diagnosed with ALL after age 6, suggesting that particular nutrients in breast milk are protective for ALL. When the bioinformatics pipeline was extended to 48 AML cases and 46 controls from the CCLS, there was no overlap between the 16 predictors of AML in females and 15 predictors in males, suggesting that neonatal metabolomic profiles of pediatric AML risk are sex-specific (Petrick et al. submitted). In females, four predictors of AML were putatively annotated as ceramides, a class of metabolites that has been linked with cancer cell proliferation, and two metabolite predictors of AML were strongly correlated with breastfeeding practice, indicating that breastfeeding may also be protective of pediatric AML. Regarding the 28 Cys34 adducts measured the CCLS subjects, we found no differences in adduct abundances between childhood leukemia cases and controls overall. However, cases of T-cell ALL had higher abundances of adducts of reactive carbonyl species and a Cys34 disulfide of homocysteine was present at lower levels in AML cases. These results suggest that oxidative stress and lipid peroxidation may be etiologic factors of T-cell ALL, and alterations in one-carbon metabolism and epigenetic changes may be predictors of AML.
Aim 4. Perform targeted analyses of small molecules chosen a priori as possible biomarkers of childhood ALL. In parallel with Aim 2, we performed quantitative assays to measure targeted small-molecule biomarkers of putative causes of childhood leukemia. The particular biomarkers included metabolites of benzene (free and conjugated phenol, hydroquinone and catechol), maternal alcohol consumption (ethyl glucuronide), maternal coffee consumption (free and conjugated caffeine, trigonelline, and atractyligenin) and maternal smoking (cotinine) as well as several Cys34 adducts (oxidation products and disulfides) that are biomarkers of reactive oxygen species. We detected several of these biomarkers in ANBS from infants of smoking and nonsmoking mothers, namely caffeine, phenol, hydroquinone, and atractyligenin plus all targeted Cys34 adducts. However, there was no evidence that any of these biomarkers differed in abundance between cases and controls.
Aim 5. Measure and compare “omic” features between pairs of ANBS and maternal blood samples collected during pregnancy from 200 mothers of childhood ALL cases and 400 mothers of control children. As mentioned under Aim 2, we characterized metabolomes and adductomes in 200 child-mother pairs of ALL cases and controls. Although preprocessing workflows have been completed for all samples, quantitation of metabolomic and adductomic features have not yet been performed due to depletion of funds at the end of the project. Unanticipated expenses were incurred in the last year of Project 2 due to the need to perform metabolomics under a subcontract to Dr. L. Petrick who had left UC Berkeley to join the faculty at the Mount Sanai School of Medicine. This change of venue for metabolomics was necessitated by the untimely death of Dr. W. Edmands who had been scheduled to perform the metabolomics during Year 07 in Prof. Rappaport’s laboratory at UC Berkeley. All of the raw and processed metabolomics and adductomics data have been transferred to CIRCLE’s secure server in anticipation that renewed funding under various avenues will permit completion of this aim.
In summary: We developed untargeted methods for extracting small molecules and Cys34 adducts of HSA from ANBS and then measuring these chemicals via metabolomics and adductomics in childhood leukemia cases and controls. These methods employed innovative sample processing pipelines to facilitate rapid and efficient collection of data. Using data from untargeted metabolomics and adductomics with ANBS from childhood leukemia and controls, we employed a novel bioinformatic pipeline to find discriminating omics features. Comparisons of small molecules between ALL cases and matched controls point to potential roles of several lipids in the etiology of ALL, including some that were highly associated with breastfeeding practice. Our results suggest that breastfeeding may reduce ALL risks because breast milk has lower abundances than infant formula for particular lipids derived from linoleic and linolenic acids. Additional analyses of AML cases and controls indicated that small molecules associated with AML differ between males and females and that breastfeeding may reduce risks of AML in females. Although no overall differences were detected between abundances of Cys34 adducts in childhood leukemia cases and controls, cases of T-cell ALL had higher abundances of adducts of reactive carbonyl species and cases of AML had lower abundance of a Cys34 disulfide of homocysteine. These results suggest that oxidative stress and lipid peroxidation may be etiologic factors of T-cell ALL, and alterations in one-carbon metabolism and epigenetic changes may be predictors of AML. The list of publications and presentation is available in the executive summary.
Project 3:
Aim 1: Construct a constitutive genetic framework for DNA methylation profiles at birth.
- Conduct genome-wide assessment of the impact of genetic polymorphisms (Affymetrix LAT array) on DNA methylation (Illumina HM450K methylation array) in neonatal dried blood spots from 200 childhood ALL cases and 400 controls in the CA Birth Cohort.
- Replicate associations found in Aim 1a in another set of 200 childhood ALL cases and 400 controls from the CCLS that have complete genome-wide association study (GWAS) and DNA methylation data.
Aim 2: Determine the perturbation of DNA methylation by in utero chemical, dietary, and immune factors, incorporating constitutive genetics, and the subsequent effect on childhood ALL risk.
- Assess the role of newly identified chemical risk factors (Project 2, Aims 3 and 4), previously identified chemical risk factors (first cycle of funding), immune factors (Project 1), and dietary factors (exploratory, folic acid pathway nutrients), on DNA methylation events and childhood ALL risk.
- Assess the effects of environmental factors (described in Aim 2a) on DNA methylation and childhood ALL risk, accounting for the role of constitutive genetic factors refined from Aim 1.
Aim 3: Assess the impact of in utero chemical exposures on pre-B cell development, pre-B cell DNA methylation, and leukemogenesis in the offspring of exposed mice.
- Using an engineered mouse model of the most common subtype of childhood ALL, test the effect of in utero chemical exposures on pre-B cell development, pre-B cell DNA methylation changes in murine genetic regions that are homologous to chemically-sensitive human genetic regions, and leukemogenesis.
What was accomplished under these goals:
To complete Aims 1 and 2, we received from the California State’s biobank the archived blood spots for neonatal DNA methylation studies and genotyping. DNA methylation and genotyping were assessed on all subjects, resulting in four sets of data. Our first two sets include DNA methylation with the Illumina HM450K panel, and later two sets with the EPIC850K panel. Genotyping was performed with Illumina Omni Express (3 sets) and Atffymetrix LAT array (one set) and all subjects imputed for missing genoptypes using HRC v1.1 and more recently with TOPMed reference panels, which is more appropriate for our majority Latino sample set.
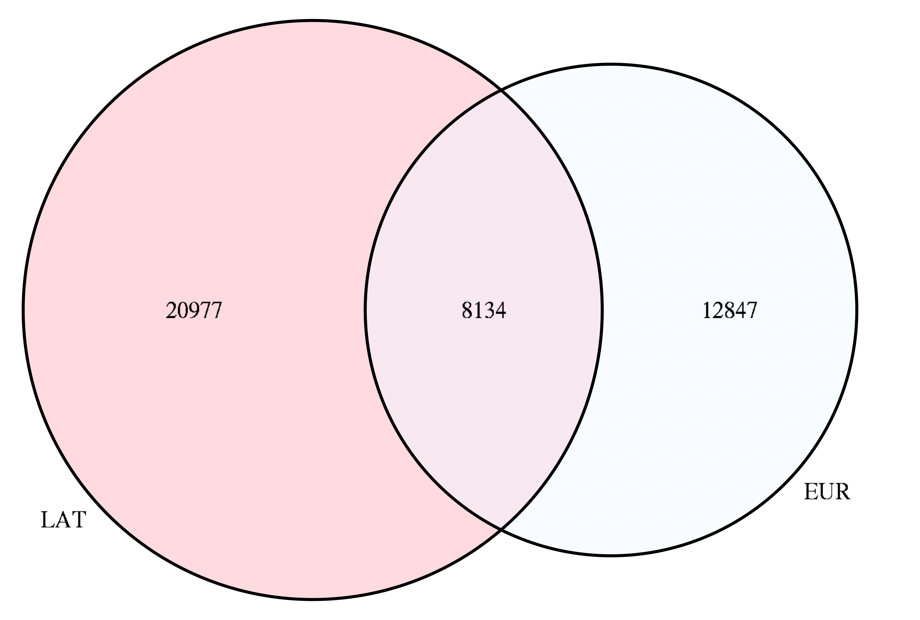
Quality Assurance: For Aim 1, our goal was to create a matrix of data on methylation-quantitative trait loci (mQTL) which can be used to account for genetic effects on DNA methylation when examining environmental risk factors. QTLTools was used with 100 permutations to determine cis-mQTLs in all datasets. This resulted in the identification of over 5 million mQTLs. Filtering of minor QTLs was performed – resulting in a stringent subset of 29,813 mQTLs for Latinos and 21,945 in non-Latino (NL) Whites. There was a significant overlap of mQTLs (Figure 1) as should be expected, given the European admixture in Latinos, but importantly the large number of non-overlapping mQTLs is another reason these groups need to be stratified in additional analyses. This mQTL matrix is now being utilized to adjust for genetic effects in environmental analyses which are ongoing to complete Aim 2. Our current work is using this mQTL matrix to analyze some covariates well known to affect DNA methylation, including birthweight and smoking which themselves will be novel analyses, and then to all the additional environmental risk variables (Aim 2).
Figure 1. top mQTL loci in HM450K loci (also present on HM850K EPIC array) in Latinos (LAT) and non-Latino Whites (EUR), demonstrating a significant but limited overlap between the two groups
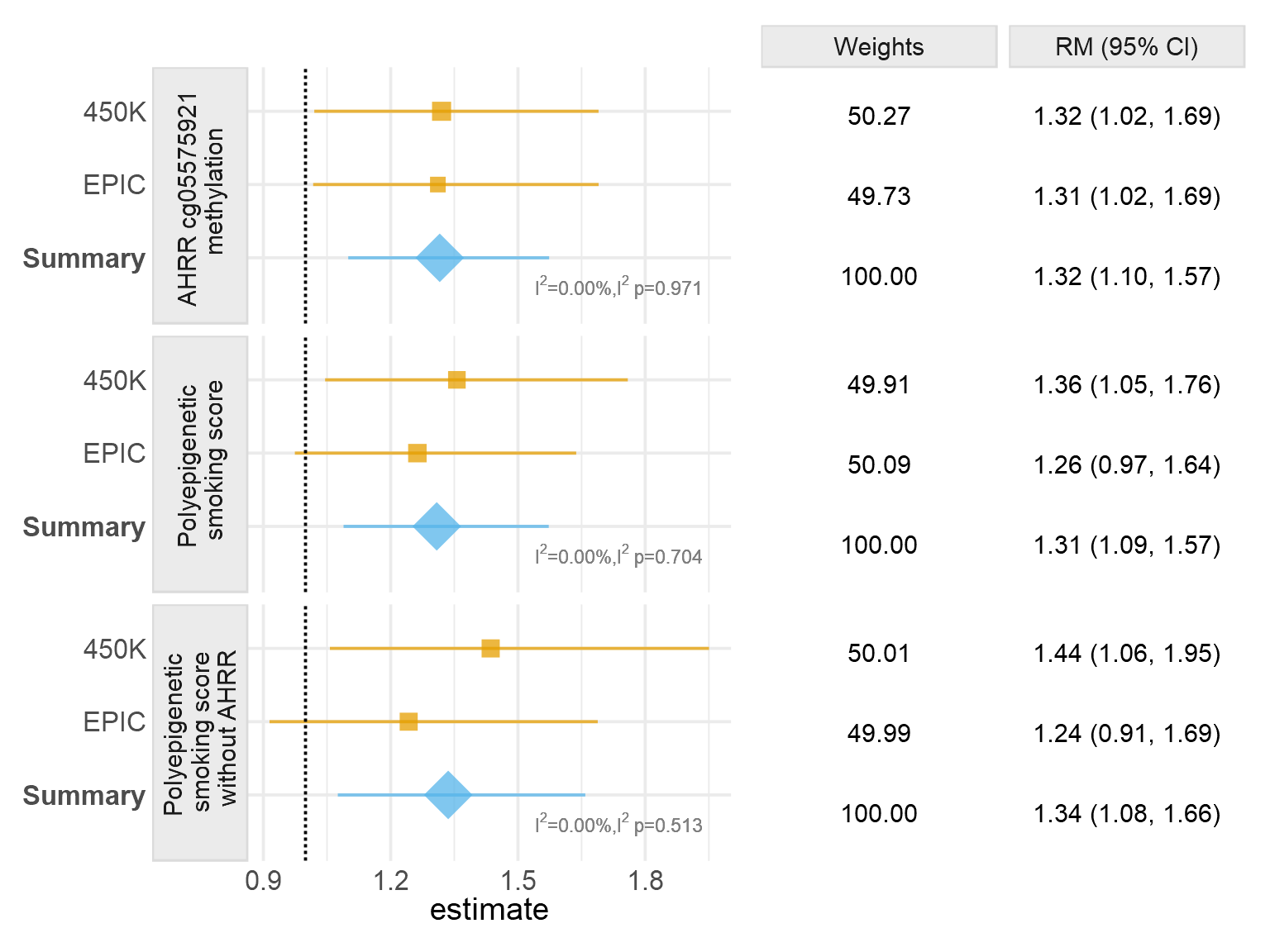
“Epigenetic biomarker” of maternal smoking: We noted that the top CpG loci recently replicated in PACE consortium studies were also the top hits in our data. This allowed us the potential to use CpG methylation at specific sites as an “epigenetic biomarker” of maternal smoking. Interestingly epidemiologic case-control studies have long shown a consistent risk estimate for paternal smoking and childhood leukemia, but no relationship with maternal smoking. There are potential biases which could affect this – maternal smoking may be underreported or the risk may occur via sidestream smoke from other housing members for instance. We therefore used DNA methylation at the most sensitive locus for maternal smoking, cg05575921, and found that DNA methylation at this locus was lower (which is the direction indicating increased smoking) at birth in our leukemia cases compared to controls (P = 0.009, n = 242 cases and 238 controls). We also assessed maternal smoking and tumor genetic features of leukemia cells, finding that self-reported smoking was associated with numbers of deletions in leukemia cells in a dose-dependent fashion (de Smith et al., 2017). Recently we completed a replication of this analysis using additional data created from our supplemental funding for this project on the 850K EPIC array, and found the same result in additional case subjects (See Figure 2 below). This result was also corroborated by using DNA methylation of cg05575921 as a biomarker of maternal smoking. In addition, we used a DNA methylation “smoking score” to further specify the association, correcting for genetic effects on the same CpG sites (as pertaining to Aim 1 above). This work is under preparation for publication.
Figure 2. Forest plot from fixed effect meta-analyses of Poisson regression results in two sets of samples (450K and EPIC). The upper panel includes Poisson regression results for the association between DNA methylation at the AHRR CpG cg05575921 and deletion numbers. The middle panel includes Poisson regression results for the association between the poly-epigenetic smoking score and deletion numbers. The bottom panel includes Poisson regression results for the association between the polyepigenetic smoking score excluding AHRR cg05575921 and deletion numbers. Ratio of means (RMs) were calculated for every 0.1 beta value decrease of AHRR CpG cg05575921 and every 4-unit increase of polyepigenetic smoking score. All the Poisson regression models were adjusted for cell type heterogeneity and genetic ancestry. Models with exposure variable DNA methylation at the AHRR cg05575921 were additionally adjusted for genotype at SNP rs148405299 in the 450K dataset and genotype at SNP rs77111113 in the EPIC dataset. Centers of squares and horizontal bars through each indicate point estimates and 95% confidence intervals (CI) of individual set RM. Area of squares indicate relative weights of individual set. Vertical apices of diamonds and horizontal bars through each indicate summary RM and 95% CI. Relative weights (%) of two sets, RM, sRM, and 95% CI were summarized in the right panel.
At this date of writing, we have initiated the analysis with Project 2 “exposomic” data, in which case we are analyzing the top small molecules which are associated with leukemia case status, and Project 1 cytokine data where we are examining neonatal immune phenotypes.
“Novel epigenetic markers predicting childhood leukemia”: We have worked with our colleagues at the International Agency for Research on Cancer (Zdenko Herceg and Akram Ghantous) and the University of Melbourne (Richard Saffery) to discover and validate new DNA methylation marks at birth that predict future risk of leukemia. We have discovered two new loci which seem to validate in all three datasets, and we will report on these in the coming year as well as analyze the role of environmental and genetic determinants that affect their DNA methylation status. This work is now a complete manuscript under editing by coauthors.
Our research group is also heavily involved in a new field called “immunomethylomics,” which is the analysis of blood cell types by their DNA methylation patterns and how this influences biology and disease incidence and progression. Our goal for neonates is to identify the presence of myeloid-derived suppressor cells and determine their role in response to environmental chemicals and leukemogenesis. This represents a connection between Project 3 (Epigenetics) and Project 1 (Immune function). As new blood cell reference sets have become available, we also found that certain CpG loci are capable of identifying fetal versus postnatal developed stem cells. This stem cell profile is highly variable among our leukemia case population and we are now working to determine the relationship of the fetal stem cell profile with demographic and exposure variables. We have found that cell subsets predicted by DNA methylation are highly correlated to certain cytokines and arginase-2, indicating that DNA methylation is a supplemental source of information on immune cell development which will be used to enhance Project 1.
For Aim 3, we have continued work on dosing strategies for the mouse model and testing DNA methylation in the mouse pups. We have completed this work for PCBs, third-hand smoke, benzo(a)pyrene, and pyrethroid pesticide exposures, and performed enhanced reduced representation bisulfite sequencing of the mouse DNA specimens as a service provided by Weill College epigenetic profiling services (Cornell University) to perform this work. Blood from the mouse dams was taken at week 5 after pregnancy exposures to the chemicals involved. The methylation patterns did not appear to be substantially different under these experimental conditions: in non‑supervised hierarchical clustering of the methylation data the animals did not separate out by exposure group (see Core C Progress Report for more details).
Conclusions:
Journal Articles: 35 Displayed | Download in RIS Format
| Other center views: | All 37 publications | 35 publications in selected types | All 35 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Breton CV, Marsit CJ, Faustman E, Nadeau K, Goodrich JM, Dolinoy DC, Herbstman J, Holland N, LaSalle JM, Schmidt R, Yousefi P, Perera F, Joubert BR, Wiemels J, Taylor M, Yang IV, Chen R, Hew KM, Freeland DM, Miller R, Murphy SK. Small-magnitude effect sizes in epigenetic end points are important in children's environmental health studies:the Children's Environmental Health and Disease Prevention Research Center's Epigenetics Working Group. Environmental Health Perspectives 2017;125(4):511-526. |
R836159 (2018) R834515 (Final) R835436 (2017) |
|
|
|
de Smith AJ, Walsh KM, Morimoto LM, Francis SS, Gonseth S, Jeon S, Chen M, Sun H, Luna-Fineman S, Antillon F, Hansen HM, Kang AY, Smirnov I, Xiao X, Whitehead TP, Barcellos LF, Sender L, Healy J, Laverdière C, Sinnett D, Taub JW, Birch JM, Thompson P, Pombo-de-Oliveira MS, Spector LG , DeWan AT, Mueller BA, Chiang C, Metayer C, Ma X, Wiemels JL (In Press) Heritable variation at the chromosome 21 gene ERG is associated with acute lymphoblastic leukemia risk in children with and without Down syndrome. Leukemia. |
R836159 (2019) |
Exit |
|
|
de Smith AJ, Kaur M, Gonseth S, Endicott A, Selvin S, Zhang L, Roy R, Shao X, Hansen HM, Kang AY, Walsh KM, Dahl GV, McKean-Cowdin R, Metayer C, Wiemels JL. Correlates of prenatal and early-life tobacco smoke exposure and frequency of common gene deletions in childhood acute lymphoblastic leukemia. Cancer Research 2017;77(7):1674-1683. |
R836159 (2017) R836159 (2018) R836159C003 (2017) |
Exit Exit Exit |
|
|
Edmands WMB, Petrick LM, Barupal DK, Scalbert A, Wilson MJ, Wickliffe JK, Rappaport SM. compMS2Miner:an automatable metabolite identification, visualization, and data-sharing R package for high-resolution LC-MS data sets. Analytical Chemistry 2017;89(7):3919-3928. |
R836159 (2017) R836159C002 (2017) |
Exit Exit Exit |
|
|
Felix JF, Joubert BR, Baccarelli AA, Sharp GC, Almqvist C, Annesi-Maesano I, Arshad H, Baiz N, Bakermans-Kranenburg MJ, Bakulski KM, Binder EB, Bouchard L, Breton CV, Brunekreef B, Brunst KJ, Burchard EG, Bustamante M, Chatzi L, Cheng Munthe-Kaas M, Corpeleijn E, Czamara D, Dabelea D, Davey Smith G, De Boever P, Duijts L, Dwyer T, Eng C, Eskenazi B, Everson TM, Falahi F, Fallin MD, Farchi S, Fernandez MF, Gao L, Gaunt TR, Ghantous A, Gillman MW, Gonseth S, Grote V, Gruzieva O, Haberg SE. Cohort profile: Pregnancy And Childhood Epigenetics (PACE) Consortium. International Journal of Epidemiology 2018;47(1):22-23u. |
R836159 (2018) R836159 (Final) R835442 (2018) |
Exit Exit Exit |
|
|
Giddings BM, Whitehead TP, Metayer C, Miller MD. Childhood leukemia incidence in California: high and rising in the Hispanic population. Cancer 2016;122(18):2867-2875. |
R836159 (2017) R836159 (2018) |
Exit Exit Exit |
|
|
Gonseth S, Roy R, Houseman EA, de Smith AJ, Zhou M, Lee ST, Nussle S, Singer AW, Wrensch MR, Metayer C, Wiemels JL. Periconceptional folate consumption is associated with neonatal DNA methylation modifications in neural crest regulatory and cancer development genes. Epigenetics 2015;10(12):1166-1176. |
R836159 (2017) R836159 (2018) R836159 (2019) R836159C003 (2016) R834511 (2014) |
Exit Exit Exit |
|
|
Gonseth S, de Smith AJ, Roy R, Zhou M, Lee S-T, Shao X, Ohja J, Wrensch MR, Walsh KM, Metayer C, Wiemels JL. Genetic contribution to variation in DNA methylation at maternal smoking-sensitive loci in exposed neonates. Epigenetics 2016;11(9):664-673. |
R836159 (2017) R836159 (2018) R836159C003 (2017) |
Exit Exit |
|
|
Kaur M, de Smith AJ, Selvin S, Zhang L, Cunningham M, Kang MW, Hansen HM, Cooper RM, McKean-Cowdin R, Wiemels JL, Metayer C. Tobacco smoke and Ras mutations among Latino and non-Latino children with acute lymphoblastic leukemia. Archives of Medical Research 2016;47(8):677-683. |
R836159 (2018) |
Exit Exit |
|
|
McCanlies EC, Fekedulegn D, Mnatsakanova A, Burchfiel CM, Sanderson WT, Charles LE, Hertz-Picciotto I. Parental occupational exposures and autism spectrum disorder. Journal of Autism and Developmental Disorders 2012;42(11):2323-2334. |
R836159 (2019) R833292 (2012) R833292 (Final) |
Exit |
|
|
Metayer C, Dahl G, Wiemels J, Miller M. Childhood leukemia: a preventable disease. Pediatrics 2016;138 (Suppl 1):S45-S55. |
R836159 (2017) R836159 (2018) |
Exit Exit Exit |
|
|
Miller MD, Valenti M, Schettler T, Tencza B. A multimedia e-book — A story of health: filling a gap in environmental health literacy for health professionals. Environmental Health Perspectives 2016;124(8):A133-A136. |
R836159 (2017) R836159 (2018) |
|
|
|
Miller MD, Valenti M, Schettler T, Tencza B. A story of health: filling a gap in environmental health literacy for health professionals. San Francisco Medicine, Journal of San Francisco Medical Society 2016;89(10):20-24 (Reprinted and edited with permission from Environmental Health Perspective). |
R836159 (2018) |
Exit |
|
|
Milne E, Greenop KR, Petridou E, Bailey HD, Orsi L, Kang AY, Baka M, Bonaventure A, Kourti M, Metayer C, Clavel J. Maternal consumption of coffee and tea during pregnancy and risk of childhood ALL: a pooled analysis from the Childhood Leukemia International Consortium. Cancer Causes & Control 2018;29(6):539-550. |
R836159 (2018) R836159 (2019) |
Exit Exit |
|
|
Orsi L, Magnani C, Petridou ET, Dockerty JD, Metayer C, Milne E, Bailey HD, Dessypris N, Kang AY, Wesseling C, Infante-Rivard C, Wunsch-Filho V, Mora AM, Spector LG, Clavel J. Living on a farm, contact with farm animals and pets, and childhood acute lymphoblastic leukemia: pooled and meta-analyses from the Childhood Leukemia International Consortium. Cancer Medicine 2018;7(6):2665-2681. |
R836159 (2018) R836159 (2019) |
Exit Exit Exit |
|
|
Petrick L, Edmands W, Schiffman C, Grigoryan H, Perttula K, Yano Y, Dudoit S, Whitehead T, Metayer C, Rappaport S. An untargeted metabolomics method for archived newborn dried blood spots in epidemiologic studies. Metabolomics 2017;13(3):27 (19 pp.). |
R836159 (2017) R836159 (2018) R836159C002 (2017) |
Exit Exit |
|
|
Petridou ET, Georgakis MK, Erdmann F, Ma X, Heck JE, Auvinen A, Mueller BA, Spector LG, Roman E, Metayer C, Magnani C, Pombo-de-Oliveira MS, Ezzat S, Scheurer ME, Mora AM, Dockerty JD, Hansen J, Kang AY, Wang R, Doody DR, Kane E, Rashed WM, Dessypris N, Schüz J, Infante-Rivard C, Skalkidou A. Advanced parental age as risk factor for childhood acute lymphoblastic leukemia: results from studies of the Childhood Leukemia International Consortium. European Journal of Epidemiology 2018;33(10):965-976. |
R836159 (2018) R836159 (2019) |
Exit Exit |
|
|
Rappaport, S.M. Redefining environmental exposure for disease etiology. NPJ Systems Biology and Application. 2018(4):30. |
R836159 (2019) |
Exit |
|
|
Sharp GC, Salas LA, Monnereau C, Allard C, Yousefi P, Everson TM, Bohlin J, XuZ, Huang RC, Reese SE, Xu CJ, Baiz N, Hoyo C, Agha G, Roy R, Holloway JW, Ghantous A, Merid SK, Bakulski KM, Kupers LK, Zhang H, Richmond RC, Page CM, Duijts L, Lie RT, Melton PE, Vonk JM, Nohr EA, Williams-DeVane C, Huen K, Rifas-Shiman SL, Ruiz-Arenas C, Gonseth S, Rezwan FI, Herceg Z, Ekstrom S, Croen L, Falahi F, Perron P, Karagas MR, Quraishi BM, Suderman M, Magnus MC, Jaddoe VWV, Taylor JA, Anderson D, Zhao S, Smit HA, Josey MJ, Bradman A, Baccarelli AA, Bustamante M, Haberg SE, Pershagen G, Hertz-Picciotto I, Newschaffer C, Corpeleijn E, Bouchard L, Lawlor DA, Maguire RL, Barcellos LF, Davey Smith G, Eskenazi B, Karmaus W, Marsit CJ, Hivert MF, Snieder H, Fallin MD, Melen E, Munthe-Kaas MC, Arshad H, Wiemels JL, Annesi-Maesano I, Vrijheid M, Oken E, Holland N, Murphy SK, Sorensen TIA, Koppelman GH, Newnham JP, Wilcox AJ, Nystad W, London SJ, Felix JF, Relton CL. Maternal BMI at the start of pregnancy and offspring epigenome-wide DNA methylation: findings from the pregnancy and childhood epigenetics (PACE) consortium. Human Molecular Genetics 2017;26(20):4067-4085. |
R836159 (2018) R835442 (2018) |
Exit Exit Exit |
|
|
Wallace AD, Francis SS, Shao X, de Smith AJ, Walsh KM, Mckean-Cowdin R, Ma X, Dahl G, Barcellos LF, Wiemels JL, Metayer C. A germ-line deletion of APOBEC3B does not contribute to subtype-specific childhood acute lymphoblastic leukemia etiology. Haematologica 2018;103(1):e29-e31. |
R836159 (2018) |
Exit Exit Exit |
|
|
Wallace AD, Francis SS, Ma X, McKean-Cowdin R, Selvin S, Whitehead TP, Barcellos LF, Kang AY, Morimoto L, Moore TB, Wiemels JL, Metayer C. Allergies and childhood acute lymphoblastic leukemia: a case-control study and meta-analysis. Cancer Epidemiology, Biomarkers and Prevention 2018;27(10):1142-1150. |
R836159 (2018) R836159 (2019) |
Exit |
|
|
Walsh KM, Whitehead TP, de Smith AJ, Smirnov IV, Park M, Endicott AA, Francis SS, Codd V, ENGAGE Consortium Telomere Group, Samani NJ, Metayer C, Wiemels JL. Common genetic variants associated with telomere length confer risk for neuroblastoma and other childhood cancers. Carcinogenesis 2016;37(6):576-582. |
R836159 (2018) |
Exit Exit Exit |
|
|
Wang R, Wiemels JL, Metayer C, Morimoto L, Francis SS, Kadan-Lottick N, DeWan AT, Zhang Y, Ma X. Cesarean section and risk of childhood acute lymphoblastic leukemia in a population-based, record-linkage study in California. American Journal of Epidemiology 2017;185(2):96-105. |
R836159 (2017) R836159 (2018) R836159C001 (2017) |
Exit Exit Exit |
|
|
Wang R, Metayer C, Morimoto L, Wiemels JL, Yang J, DeWan AT, Kang A, Ma X. Parental age and risk of pediatric cancer in the offspring: a population-based record-linkage study in California. American Journal of Epidemiology 2017;186(7):843-856. |
R836159 (2018) |
Exit Exit Exit |
|
|
Whitehead TP, Metayer C, Wiemels JL, Singer AW, Miller MD. Childhood leukemia and primary prevention. Current Problems in Pediatric and Adolescent Health Care 2016;46(10):317-352. |
R836159 (2017) R836159 (2018) |
Exit Exit Exit |
|
|
Wiemels JL, Walsh KM, de Smith AJ, Metayer C, Gonseth S, Hansen HM, Francis SS, Ojha J, Smirnov I, Barcellos L, Xiao X, Morimoto L, McKean-Cowdin R, Wang R, Yu H, Hoh J, DeWan AT, Ma X. GWAS in childhood acute lymphoblastic leukemia reveals novel genetic associations at chromosomes 17q12 and 8q24.21. Nature Communications 2018;9(1):286 (8 pp.). |
R836159 (2018) |
Exit Exit Exit |
|
|
Yano Y, Grigoryan H, Schiffman C, Edmands W, Petrick L, Hall K, Whitehead TP, Metayer C, Dudoit S, Rappaport SM. Untargeted adductomics of cys34 modifications to human serum albumin in newborn dried Blood Spots. Analytical and Bioanalytical Chemistry 2019. |
R836159 (2019) |
Exit |
|
|
Schiffman C, Petrick L, Perttula K, Yano Y, Carlsson H, Whitehead TP, Metayer C, Hayes J, Edmands W, Rappaport SM, Dudoit S. Data-adaptive pipeline for filtering and normalizing metabolomics data. Biorxiv. 2018. |
R836159 (2019) |
Exit Exit Exit |
|
|
de Smith AJ, Walsh KM, Francis SS, Zhang C, Hansen HM, Smirnov I, Morimoto L, Whitehead TP, Kang A, Shao X, Barcellos LF, McKean-Cowdin R, Zhang L, Fu C, Wang R, Yu H, Hoh J, Dewan AT, Metayer C, Ma X, Wiemels JL. BMI1 enhancer polymorphism underlies chromosome 10p12.31 association with childhood acute lymphoblastic leukemia. Int J Cancer 2018;143(11):2647-2658. |
R836159 (2019) |
Exit |
|
|
Panagopoulou P, Skalkidou A, Marcotte E, Erdmann F, Ma X, Heck JE, Auvinen A, Mueller BA, Spector LG, Roman E, Metayer C, Magnani C, Pombo-de-Oliveira MS, Scheurer ME, Mora AM, Dockerty JD, Hansen J, Kang AY, Wang R, Doody DR, Kane E, Schüz J, Christodoulakis C, Ntzani E, Petridou ET. Parental age and the risk of childhood acute myeloid leukemia:results from the Childhood Leukemia International Consortium. Journal of Cancer Epidemiology. 2019(59):158-165. |
R836159 (2019) |
Exit |
|
|
Schiffman C, Petrick L, Perttula K, Yano Y, Carlsson H, Whitehead T, Metayer C, Hayes J, Rappaport S, Dudoit S. Filtering procedures for untargeted LC-MS metabolomics data. BMC Bioinformatics. 2019(1):20:334. |
R836159 (2019) |
Exit |
|
|
Petrick LM, Schiffman C, Edmands WMB, Yano Y, Perttula K, Whitehead T, Metayer C, Wheelock CE, Arora M, Grigoryan H, Carlsson H, Dudoit S, Rappaport SM. Metabolomics of neonatal blood spots reveal distinct phenotypes of pediatric acute lymphoblastic leukemia and potential effects of early-life nutrition. Cancer Letters 2019;452:71-78. |
R836159 (2019) |
Exit Exit |
|
|
Yano Y, Schiffman C, Grigoryan H, Hayes J, Edmands W, Petrick L, Whitehead T, Metayer C, Dudoit S, Rappaport S. Untargeted adductomics of newborn dried blood spots identifies modifications to juman serum albumin associated with childhood leukemia. Leukemia Research. 2020(8):106268 |
R836159 (2019) |
Exit |
|
|
Karalexi MA, Dessypris N, Clavel J, Metayer C, Erdmann F, Orsi L, Kang AY, Schüz J, Bonaventure A, Greenop KR, Milne E, Petridou ET. Coffee and tea consumption during pregnancy and risk of childhood acute myeloid leukemia:A Childhood Leukemia International Consortium (CLIC) study. Cancer Epidemiology. 2019(62):101581 |
R836159 (2019) |
Exit |
|
|
Nielsen AB, Zhou M, de Smith AJ, Want R, McCoy L, Hansen H, Morimoto L, Gronbaech K, Johansen C, Kogan SC, Metayer C, Bracci PM, Ma X, Wiemels JL. Increased neonatal level of arginase 2 in cases of childhood acute lymphoblastic leukemia implicates immunosuppression in etiology. Haematologica. 2019. |
R836159 (2019) |
|
Relevant Websites:
Center for Integrative Research on Childhood Leukemia and the Environment Exit
GithubsimExTargId, Exit GithubMetabolomics-Filtering Exit
Sound Cloud Maternal Smoking Exit
Sound Cloud Males Smoking Exit
Progress and Final Reports:
Original Abstract Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R836159C001 In Utero Chemical Exposures, Immune Status, and Childhood Leukemia
R836159C002 Identifying In Utero Exposures that are Risk Factors for Childhood Leukemia
R836159C003 Prenatal Exposures, Constitutive Genetics, DNA Methylation & Childhood Leukemia
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
- 2019 Progress Report
- 2018 Progress Report
- 2017 Progress Report
- 2016 Progress Report
- Original Abstract
35 journal articles for this center