Grantee Research Project Results
Final Report: Prediction of Allergenicity by Linear and Conformational Epitopes
EPA Grant Number: R834823Title: Prediction of Allergenicity by Linear and Conformational Epitopes
Investigators: Braun, Werner , Schein, Catherine H. , Ivanciuc, Ovidiu I
Institution: The University of Texas Medical Branch - Galveston
EPA Project Officer: Aja, Hayley
Project Period: September 15, 2010 through September 30, 2014
Project Amount: $425,000
RFA: Approaches to Assessing Potential Food Allergy from Genetically Engineered Plants (2009) RFA Text | Recipients Lists
Research Category: Human Health
Objective:
Recombinant genetic tools are increasingly used by the biotechnology industry to introduce novel proteins into crops for protection against pesticides, as plant-incorporated protectants (PIP) or to increase the nutritional value of the foods. Appropriate safeguards must thus be in place to ensure that these techniques will not introduce new proteins that are potentially allergenic. As up to 8% of infants and young children and 2% of adults are allergic to one or more food allergens, the potential individuals who might be harmed by those GMO products represent a substantial fraction of the U.S. population. In cases such as allergies to peanuts, the physiological effect can be quite severe, as exposure of sensitive individuals to certain proteins in the peanut plant can lead to anaphylactic shocks with a deadly outcome.
The goal of this project was to provide quantitative bioinformatics criteria to determine the potential allergenicity of transgenic proteins in food products and PIPs using allergen specific motifs. A computational approach, using sequence and structural similarities of the new gene product to known allergenic proteins, can quantitatively address the potential allergenicity of a novel gene product. In this project, we contributed to the safety assessment by providing software tools and archiving sequence and structural details on allergenic proteins and make them publicly available on our “Structural Database of Allergenic Proteins” (SDAP) website (http://fermi.utmb.edu/SDAP/).
The specific aims of the projects were three-fold:
Aim1: To maintain and expand the currently available information in our Structural Data Base of Allergenic Proteins (SDAP) on the sequences, 3D structures and IgE epitopes of allergens.
Aim2: To establish a data base of allergen specific motifs that can be quantitatively screened to estimate the potential allergenicity of a query protein sequence. A major effort in pursuing aim 2 is the experimental validation of the computational tools in SDAP.
Aim3: Develop novel computational methods to detect similar conformational epitopes.
Summary/Accomplishments (Outputs/Outcomes):
Results for Aim 1.
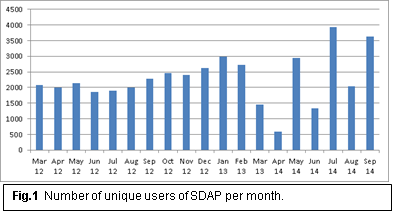
(a) We maintained and expanded our Structural Data Base of Allergenic Proteins (SDAP) as a user friendly Web server to the allergenic community. The website is still maintained and available through two ports, an http port (http://fermi.utmb.edu/SDAP/) and a special secure https port (https://fermi.utmb.edu/SDAP/) with a widely accepted digital certificate. SDAP was heavily used by the allergenic community with approximately 1,500 unique users. Currently, the following information on allergenic protein for SDAP users can be downloaded from our website: 1,526 allergens and isoallergens, 1,312 protein sequences for allergens and isoallergens, 92 allergens with experimentally determined three-dimensional structures, 458 three-dimensional models, specific IgE epitope information on 29 allergens, a list of protein families grouped according to the PFAM (Protein families) database, maintained by the European Bioinformatics Institute. An SDAP user can perform keyword and similarity sequence searches in the database, and execute the FAO/WHO bioinformatics guidelines for a query sequences or calculate peptide similarity searches with our property distance (PD) tool. The PD tool is also available as a downloadable script from the website.
(b) We proposed a new Markup Language, AllerML, as a standard computer readable format for allergens. The information exchange between major allergen services, databases and users is currently hindered by the absence of a common standard language. AllerML can streamline the data exchange between different databases, bioinformatics servers, and users. AllerML has been implemented for all entries in SDAP, thus providing a computer-readable access to our allergen database. The flexibility of the markup language allows defining new tags to accommodate information from other allergen databases. Details for the AllerML tags have been published in the journal, Regulatory Toxicology and Pharmacology (Ivanciuc et al., Regul. Toxicol. Pharmacol. 2011;60(1):151-160).
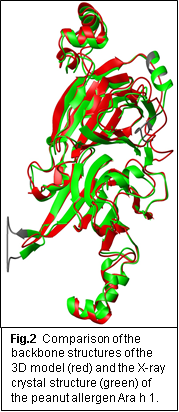
(c) We generated three-dimensional models for those allergen proteins that had no 3D structures at the time of the modelling and made them available on the website. As new three-dimensional protein structures became available, we used it as an opportunity to assess the quality of those 3D models. The comparison of the 3D models in SDAP to their experimental structures, determined after our modeling efforts, showed the high quality for backbone fold and mapping of IgE epitopes on the surface of the protein structures if the identity between the allergen sequence and the sequence of the template structure is higher than 25%. As an example of the quality of the 3D models, we show in Figure 2 the comparison of the backbone structure of our 3D model for the peanut allergen Ara h 1 and the experimentally determined 3D structure for that allergen. We published a detailed analysis of this comparison in the journal, Proteins. (Power et al., Proteins: Struct. Funct. and Bioinf. 2013;81(4):545-554).
(d) A new feature of the SDAP website is the installation of a secure https port (https://fermi.utmb.edu/SDAP). This port allows users to access the SDAP website over a secure connection that prevents potential interception of the information by third parties. Specifically it guarantees that the user of SDAP will communicate with a trusted website as certified by a well-established authority by a digital certificate.
Results for Aim 2.
(a) We demonstrated that a score function based on allergen-specific motifs discriminates between non-allergens and allergens. We used our PCPMer software tool to generate allergen specific sequence motifs. These sequence motifs are short linear sequences of typically 8 to 12 amino acid residues with conserved physical chemical properties in a multiple sequence alignment of all allergenic sequences in a protein family. The quantitative definition of a motif is a profile type characterization of the multiple sequence alignment with the average values and standard deviations of our quantitative descriptors E1 to E5. The relative entropy, a measure for conservation, is also calculated and used as criterion for selection. We had previously shown that only 130 different proteins domains occur in allergenic proteins, among more than 10,000 different domains in the Pfam classification (Ivanciuc et al., Mol. Immun. 46(4), 559-568, 2009). Thus, only a small subset of those domains contains most allergens. We have now generated sequence motifs for highly populated allergen families. These families include all major allergens from common allergenic foods, such as milk, eggs, fish, crustacean shellfish, tree nuts, peanuts, wheat and soybeans which account for 90% of food allergic reactions. In a new development for allergen research, we defined a score function for a given user query sequence that determines is degree of similarity to any of the highly populated allergen family. The score function was tested against a control set of 84 939 non-allergenic protein sequences generated from a set of non-redundant protein sequences in UniProt. The distribution of the score values shows a clear separation of the values between non-allergenic sequences and allergenic sequences. We also have data showing the discriminating power of the score function between allergens and non-allergens within the same family (manuscript in preparation).
(b) Predicted short linear peptides by the PD tool in SDAP for peanut and nut allergens were experimentally tested by mono-clonal antibodies and verified as cross-reactive.
Dr. Schein, in collaboration with Dr. Soheila Maleki, Ph.D., of the USDA, New Orleans and Dr. Suzanne Teuber, M.D., of the UC-Davis, experimentally tested whether short linear peptides common to peanut and nut allergens could be the basis of clinically observed cross-reactivity. That project helped us in validating the computational tools in SDAP for clinical and biological research use, and in bringing these tools to the attention of experimental allergen researchers. We have previously shown that the PD scale in SDAP can identify similar IgE binding areas that may be important for cross-reactivity between allergens (Ivanciuc et al., Mol. Immun. 2009;46(5):873-883). Searching the SDAP using the PD tool revealed many potential IgE epitopes in other nut allergens, including several 7S, 2S and 11S albumins, with similar physicochemical properties (low PD value) to known Ara h 2 epitopes. Cross-reactivity of the predicted allergens were tested with anti-Ara h 2 monoclonal antibodies. Those monoclonal antibodies recognized many of the predicted vicilin, conglutinin, and glycinin nut allergens. All of the monoclonal antibodies recognized the Jug r 1 allergen. Four antibodies were highly reactive with the Jug r 2 leader sequence, confirming the presence of similar antigenic regions. Thus, repeated sequences similar to known IgE epitopes are common to many different allergenic proteins from nuts and seeds. The importance of these regions for clinically relevant IgE cross-reactivity is indicated by these results. In addition, we synthesized short peptides that correspond to motifs of the peanut allergens and tested their IgE binding affinity with serum from peanut sensitive patient by micro array experiments. Some of the motifs overlap with previously reported epitopes. Results are published in Nesbit et al., Mol. Nutr. Food Res. 2012;56,1, and Schein et al., BMC Bioinformatics, 2012;13(Suppl 13):S9.
Results for Aim 3.
Prediction of conformational IgE epitopes from phage display data. In the past, computational methods to predict clinically observed cross-reactivity were based entirely on amino acid sequence analysis of the proteins involved. We developed a new computational method that also includes 3D structural information on predicting conformational epitopes, and tested the new 3D prediction method based on phage display experiments for the dust mite allergen Bla g 2 (Tiwari et al., Int. Arch. Allergy Immunol., 2012;157(4):323-330).
In silico screening of cross-reactive epitopes in a 3D database of allergens. Our new method to predict clinically observed cross-reactivities from 3D structures has achieved first results. Currently, we have tested the method for recognizing similar surface exposed patches in all allergens of SDAP with a known 3D structure. We used the known conformational epitopes of the birch pollen allergen Bet v1, the hyaluronidase, a major bee venom allergen, the beta-lactoglobulin allergen, the cockroach allergen Bla g 2 and the grass pollen allergen Phl p 2 as a test case and searched in SDAP for allergens with similar epitopes. The new method successfully predicted the cross-reactivity of allergens consistent with known clinical observations. A manuscript describing our new method is in preparation.
Conclusions:
- SDAP is an essential resource for allergy researchers and regulators worldwide. The user-friendly website can perform several standard bioinformatics searches as well as novel similarity searches that will inform about the potential risk of an allergenic protein.
- Allergen-specific motifs are defined as short linear sequences that are common to many related known allergens. They provide an alternative way to predict IgE cross-reactivity. We defined those motifs in 17 major families of allergenic proteins and showed that these motifs can distinguish allergens in a protein family from non-allergenic members in the same family.
- We showed the value of 3D structures of proteins in the risk assessment, and considerably expanded the knowledge on 3D structures of allergens by a large scale 3D modeling effort. We could demonstrate the high quality of the 3D models for allergen research but also pointed out what quality assessment criteria need to be applied. Thus, we suggest that high-quality 3D models can be used for finding cross-reactive regions of allergens.
- Computational methods to predict clinically observed cross-reactivity were based in the past entirely on amino acid sequence analysis of the proteins involved. We developed a new computational method that also includes 3D structural information on conformational epitopes in the prediction of cross-reactivity.
- The PD tool was experimentally validated for the identification of cross-reactivity among peanut and nut allergens.
Journal Articles on this Report : 6 Displayed | Download in RIS Format
| Other project views: | All 19 publications | 6 publications in selected types | All 6 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Ivanciuc O, Gendel SM, Power TD, Schein CH, Braun W. AllerML: markup language for allergens. Regulatory Toxicology and Pharmacology 2011;60(1):151-160. |
R834823 (2011) R834823 (2013) R834823 (Final) R834066 (Final) |
Exit Exit Exit |
|
|
Negi SS, Braun W. Cross-React: a new structural bioinformatics method for predicting allergen cross-reactivity. Bioinformatics 2017;33(7):1014-1020. |
R834823 (Final) |
Exit Exit |
|
|
Nesbit JB, Hurlburt BK, Schein CH, Cheng H, Wei H, Maleki SJ. Ara h 1 structure is retained after roasting and is important for enhanced binding to IgE. Molecular Nutrition and Food Research 2012;56(11):1739-1747. |
R834823 (2012) R834823 (2013) R834823 (Final) R834066 (Final) |
Exit |
|
|
Power TD, Ivanciuc O, Schein CH, Braun W. Assessment of 3D models for allergen research. Proteins 2013;81(4):545-554. |
R834823 (2012) R834823 (2013) R834823 (Final) |
Exit |
|
|
Schein CH, Bowen DM, Lewis JA, Choi K, Paul A, van der Heden van Noort GJ, Lu W, Filippov DV. Physicochemical property consensus sequences for functional analysis, design of multivalent antigens and targeted antivirals. BMC Bioinformatics 2012;13(Suppl 13):S9. |
R834823 (2012) R834823 (2013) R834823 (Final) R834066 (Final) |
Exit Exit |
|
|
Tiwari R, Negi SS, Braun B, Braun W, Pomes A, Chapman MD, Goldblum RM, Midoro-Horiuti T. Validation of a phage display and computational algorithm by mapping a conformational epitope of Bla g 2. International Archives of Allergy and Immunology 2012;157(4):323-330. |
R834823 (2012) R834823 (2013) R834823 (Final) R833137 (Final) |
Exit Exit |
Supplemental Keywords:
Relational database, large scale 3D modeling of proteins, WHO/EFSA recommendations for risk assessment of proteinsRelevant Websites:
(1) Home page of the Structural Data base of Allergens (SDAP): https://fermi.utmb.edu/SDAP/ Exit
(2) Presentation of Dr. Braun in the HESI PATC and IFBiC Biotechnology Symposium 2013, Arlington, VA. (video Exit )
(3) Presentation of Dr. Schein in the HESI PATC and IFBiC Biotechnology Symposium 2013, Arlington, VA. (video Exit )
Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.