Grantee Research Project Results
Final Report: Assessment of Microbial Pathogens in Drinking Water using Molecular Methods Coupled with Solid Phase Cytometry
EPA Grant Number: R833830Title: Assessment of Microbial Pathogens in Drinking Water using Molecular Methods Coupled with Solid Phase Cytometry
Investigators: Pyle, Barry H , Camper, Anne
Institution: Montana State University - Bozeman , Little Big Horn College
EPA Project Officer: Aja, Hayley
Project Period: March 1, 2008 through February 28, 2011 (Extended to February 28, 2013)
Project Amount: $599,996
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens and Cyanobacteria and Their Toxins in Drinking Water (2007) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The determination of microbial safety of drinking water is still done by assessing the presence of fecal contamination through culturing indicator bacteria, such as E. coli, rather than by direct detection of pathogens from the water. Ideally, the absence of the indicators should always signal the absence of pathogens; however, this is not always the case. Because pathogens and indicator bacteria react differently to environmental stresses, pathogens can be present in drinking water even when no indicator bacteria have been detected (1, 2). Additionally, because the methods for detection of indicator organisms rely on culture techniques, it can take days before indicator bacteria can be identified. The goal of this study was to find a rapid method to detect multiple pathogens directly from water. Although this is a global issue, the project was structured toward identifying microbial contamination and its significance within Montana, particularly on the Crow Indian Reservation east of Billings. The objectives of this study were to: (1) develop a sensitive method for rapid/real-time detection and enumeration of target microbial pathogens, including assessment of viability and infectivity using fluorescent in situ hybridization (FISH) coupled with Solid Phase Laser Cytometry, (SPLC), which facilitates very rapid, sensitive and reliable enumeration and immediate confirmation by epifluorescent microscopy (EFM); (2) evaluate this method for identification and detection of Escherichia coli O157:H7, Helicobacter pylori, Legionella pneumophila, Mycobacterium avium, Aeromonas hydrophila, Giardia lamblia, and Cryptosporidium parvum; (3) examine the possibility of detection of multiple pathogens at the same time; and (4) determine if the method could be used in environmental conditions in collaboration with the Crow community.
Summary/Accomplishments (Outputs/Outcomes):
| Frequency Table Thresholding | |
|---|---|
| Ko | 0 |
| K1 | 0.05 |
| Pretrigger | 0 |
| Trig Delay | 5 |
| FISH Hybridization |
| R2A broth with nalidixic acid 2 4 hours ↓ 4% paraformaldehyde 1 hour ↓ 50 % 80% 94% ETOH baths 4 min each ↓ 60 µl TE rinse 5 min ↓ Lysozyme 10 min ↓ 60 µl 0.02 N HCL 10 min ↓ 60 µl TE rinse 5 min ↓ Hybridize 90 min at 48 °C ↓ Wash buffer 100 µl 30 min 48°C ↓ Rinse TNT buffer 5 min ↓ FITC tyramide signal amplification 75 µl 10 min ↓ Rinse TNT buffer 5 min ↓ Store dry on cellulose pad at 4˚C until analyzed |
| Date | EPA | ScanRDI | ||
| | # | Log | # | Log |
| 9/9/2011 | 196 | 2.29 | 315 | 2.50 |
| 9/21/2011 | 109 | 2.04 | 235 | 2.37 |
| 2/21/2012 | 236 | 2.37 | 62 | 1.79 |
| 4/13/2012 | 973 | 2.99 | 204 | 2.31 |
| 5/22/2012 | 45 | 1.65 | 26 | 1.41 |
| 6/28/2012 | 158 | 2.20 | 3 | 0.52 |
| Mean | 286 | 141 |
| |
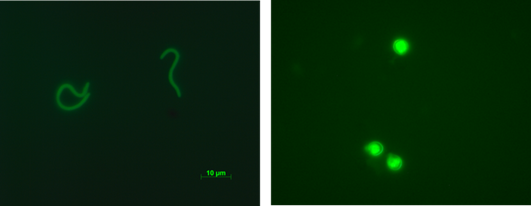
Objective 4 Use of method in environmental conditions in collaboration with the Crow community
As noted above, the Crow Environmental Health Steering Committee specifically requested that we look for Cryptosporidium parvum in the Little Big Horn River. This river is the source water for the water treatment facility at Crow Agency, Montana and may be contaminated by both cattle operations and rural houses upstream from the treatment plant. Nine water samples were taken from the source water and the treated drinking water over the course of 1 year, from June 2011 through June 2012. The samples were collected using filtration, concentrated by centrifugation and purified with immunomagnetic separation as described in EPA Method 1623. The resulting sample was split evenly and labeled with either the FAb/DAPI/DIC and examined microscopically or labeled with the newly developed FISH method and examined with the ScanRDI. At each sampling point, both river water and drinking water were also tested for coliforms and E. coli using Hach mColiBlue.
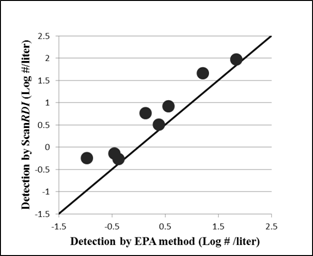
Cryptosporidium parvum oocysts were found in all river water samples tested by both EPA and ScanRDI methods, with highest numbers found in June and July, 2012 (Table 4). Figure 3 shows that the number of oocysts labeled with FISH detected by the SPLC was higher than the number of oocysts labeled with FAb/DAPI enumerated by the manual microscopy for every sample taken from the river.
There was no correlation between the number of E. coli or coliforms and the number of oocysts found in a sample.
| Date | Coliform (CFU/100 ml) | E. coli (CFU/100ml) | Oocysts EPA #/liter | Oocysts ScanRDI#/liter | Total oocysts in Sample (Calculated) | Total oocysts/liter (Calculated) |
| 6/17/2011 | 6130 | 267 | 16.6 | 44.9 | 461.1 | 30.7 |
| 7/8/2011 | 6700 | 130 | 70.0 | 91.7 | 1212.8 | 80.9 |
| 8/10/2011 | 5230 | 333 | 2.4 | ND* | 24.0 | 1.2 |
| 9/8/2011 | 7667 | 23 | 0.1 | 0.6 | 6.8 | 0.3 |
| 10/27/2011 | 630 | 13 | 0.4 | 0.5 | 9.56 | 0.5 |
| 2/22/2012 | 1313 | 10 | 2.5 | 3.1 | 27.9 | 2.8 |
| 4/13/2012 | 597 | 50 | 1.4 | 5.7 | 30.0 | 3.0 |
| 5/22/2012 | 2800 | 370 | 3.7 | 8.2 | 51.3 | 6.0 |
| 6/28/2012 | 1537 | 63 | 0.4 | 0.7 | 8.0 | 0.5 |
Conclusions:
| Species | Probe | Sequence | Ref. |
| Legionella pneumophila | LEGPNE1 | 5'-ATCTGACCGTCCCAGGTT-3' | (16) |
| Helper for L. pneumophila | LEG H343 | 5-TTATGACCAACTGAGCTC-3 | This lab |
| Helper for L. pneumophila | LEG H307 | 5`- ACACTTTAAGGACCCGAA-3 | This lab |
| Aeromonas hydrophila | AER66 | 5'-CTACTTTCCCGCTGCCGC-3' | (17) |
| Helper for A. hydrophila | AER66H21 | 5`- GATTGTGTACGTTCAGCT -3` | This lab |
| Helper for A. hydrophila | AER66H57 | 5`- AACGATGAAAACGGCCGC -3` | This lab |
| Cryptosporidium parvum | CRY-1 | 5′-CGGTTATCCATGTAAGTAAAG-3′ | (18) |
| E. coli non specific | ECO541 | 5'-CCGATTAACGCTTGCACC-3' | (19) |
| E. coli | Colinsitu | 5 GAGACTCAAGATTGCCAGTATCAG -3 | (9) |
| Helper for E. coli | HColin_L | 5-ATGCAGTTCCCAGGTTGAG | (5) |
| Helper for E. coli | HColin_R | ACCTGGAATTCTACCCCCCTCTAC | (5) |
| Negative control | NONEUB | 5'-ACTCCTACGGGAGGCAGC-3' | (20) |
| Positive control (Bacteria) | EUB 338 | 5-GCTGCCTCCCGTAGGAGT-3 | (21) |
References:
Journal Articles:
No journal articles submitted with this report: View all 6 publications for this projectSupplemental Keywords:
Water, drinking water, groundwater, wastewater, effluent, feedlot, health effects, organisms, pathogens, bacteria, protozoa, biology, microbiology, monitoring, measurement methods, Northwest, Montana, MT, EPA Region 8, Native American, Crow Reservation, Cryptosporidium, E. coli, Legionella,Aeromonas hydrophila, fluorescent in situ hybridization, solid phase laser cytometer.Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
