Grantee Research Project Results
2009 Progress Report: Rapid Concentration, Detection, and Quantification of Pathogens in Drinking Water
EPA Grant Number: R833840Title: Rapid Concentration, Detection, and Quantification of Pathogens in Drinking Water
Investigators: Hu, Zhiqiang , Riley, Lela K. , Lin, Mengshi
Institution: University of Missouri - Columbia
EPA Project Officer: Aja, Hayley
Project Period: May 1, 2008 through April 30, 2011 (Extended to April 30, 2013)
Project Period Covered by this Report: May 1, 2009 through April 30,2010
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens and Cyanobacteria and Their Toxins in Drinking Water (2007) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The objectives of this research are to: 1) evaluate a lanthanum-based colloidal destabilization method to rapidly concentrate pathogens in water; 2) determine the efficiency of fluorescence-based oxygen microrespirometry in differentiating live/dead pathogens; 3) develop and validate a new SERS-based method for pathogen detection and quantification; and 4) improve pathogen detection using modified concentration and molecular detection methods and compare the detection efficiencies to the Environmental Protection Agency’s (EPA) existing methods with seeded or unseeded drinking water samples.Progress Summary:
In this report, we summarize the efforts and results achieved in the following three areas: 1) concentration of microbes in water using a lanthanum-based colloidal destabilization method; 2) enumeration of microbes using standard quantitative real-time PCR (qPCR); and 3) development of a SERS-based method for bacterial detection and quantification.
- Concentration of microbes in water using a lanthanum-based colloidal destabilization method. Rapid concentration and detection of microorganisms, particularly pathogens, are important but remain a challenge. In this research, lanthanum chloride (LaCl3) was used to concentrate E. coli in water and the results were compared with those obtained using traditional flocculants, such as ferric sulfate and aluminum sulfate. A turbidimetric assay and a microrespirometric assay were employed to enumerate the bacteria in water samples by monitoring the absorbance of bacteria and the oxygen-based fluorescence intensity, respectively. The microrespirometric method requires less time than the turbidimetric assay. Both assays could linearly enumerate the bacteria at the concentration range from 101 to 109 cells/mL. Based on the turbidimetric assay, the relative concentration efficiencies of the three flocculants were 75% (LaCl3), 40% (FeCl3) and 33% (Al2(SO4)3), while for the microrespirometric assay, the concentration efficiencies were 85% (LaCl3), 34% (FeCl3) and 32% (Al2(SO4)3). The microbial recovery rates, defined as the ratio of cell number in the sediment after coagulation/flocculation to that of the controls, were 94% (LaCl3), 69% (FeCl3) and 51% (Al2(SO4)3) from the turbidimetric assay. The results demonstrate that compared with traditional flocculants, LaCl3 has higher relative concentration and recovery efficiencies and thus possesses the potential for microbial concentration in water samples. Our microbial concentration results have been published in Water Research (Water Research, 2010, 44, 3385-3392).
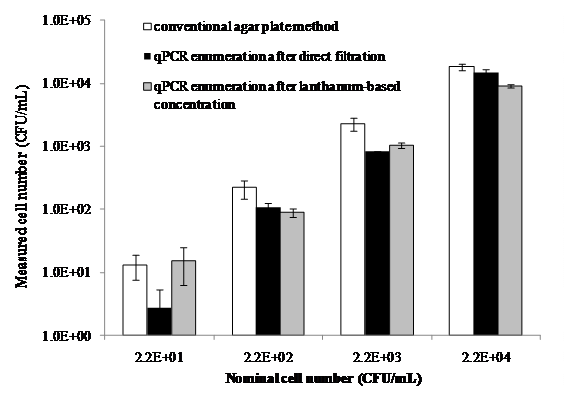
- Enumeration of microbes using standard quantitative real-time PCR (qPCR) Rapid detection of low numbers of bacteria, especially pathogenic strains, in water samples is important but challenging. We have made some breakthroughs about bacterial detection using our combined concentration, DNA extraction, and qPCR detection. In a recent study (Lanthanum-based concentration and real-time PCR detection of microbes in water, submitted), we developed the overal method. Compared with traditional flocculants such as FeCl3 and Al2(SO4)3, chemical flocculation by lanthanum chloride (LaCl3) resulted in higher bacterial concentration and recovery efficiencies. The objective of this study was to develop and evaluate a lanthanum-based bacterial concentration method coupled with quantitative real-time PCR (qPCR) to enumerate bacteria in water. The flocs containing enmeshed bacteria after chemical coagulation/flocculation must be dissolved before PCR detection. Two types of chemicals, hydrochloric acid and ethylenediaminetetraacetic acid (EDTA), were evaluated for their effectiveness in floc dissolution. The efficiency of additional bacterial concentration through filtration and centrifugation was also determined. EDTA completely dissolved the flocs after lanthanum-based coagulation/flocculation, but it did not fully dissolve the flocs after traditional coagulation/flocculation using FeCl3 or Al2(SO4)3. When compared with HCl to dissolve the lanthanum-based flocs, EDTA appeared to have higher efficacy for bacterial recovery. However, additional filtration and rinsing steps were required to remove residual EDTA that inhibited DNA extraction and PCR amplification. Lanthanum-based concentration coupled with real-time PCR was able to detect bacteria at very low concentrations (e.g., 10 CFU/mL in raw water, Figure 1). With a reduction of water volume by a factor of hundreds to possibly millions, the lanthanum-based concentration coupled with qPCR technique is therefore a promising method for detection of low numbers of specific bacteria or pathogens in water. Future study will include detection of the selected pathogens (i.e., Helicobacter pylori) in water using our developed method.
Figure 1. A comparison of the measured bacterial numbers in raw water samples at different initial bacterial concentrations among the three methods: conventional agar plate enumeration, qPCR coupled with direct filtration, and qPCR coupled with lanthanum-based concentration. Error bars represent one standard error of the mean (n=3).
- Rapid detection of water- and foodborne bacteria by surface enhanced Raman spectroscopy Rapid methods with high sensitivity and specificity for bacterial detection are of critical importance to food safety and human health. Rapid and early detection can prevent and control pathogen related clinical diagnostic diseases and bioterroristic attacks. SERS is a novel and sensitive analytical tool that can be used in testing chemical and biochemical samples using SERS-active substrates. Our previous studies show that SERS coupled with gold SERS-active substrates is reliable for identifying and differentiating different virus strains. In this study, a protocol of SERS coupled with internal-coated silver nanoparticles was established for detection of bacterial samples.
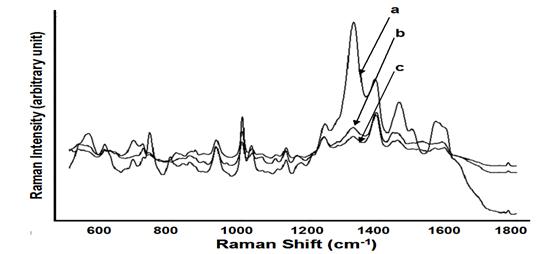
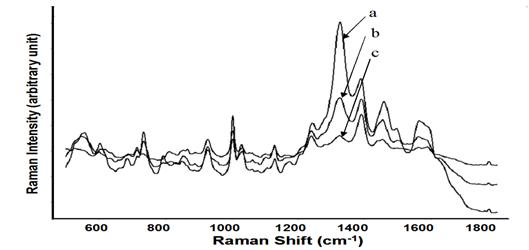
Development of rapid and sensitive methods to detect foodborne pathogens is an important aspect of improving food safety. This study aimed to develop a new method to detect and discriminate Escherichia coli O157:H7 and Staphylococcus epidermidis by surface enhanced Raman spectroscopy (SERS) coupled with silver nanomaterials. An internal coating method using silver nanostructures for bacterial samples was established and assessed to achieve satisfactory SERS performance. SERS spectra were obtained from different bacterial cells (Figure 2). Distinctive differences in SERS spectral data between bacterial species were observed, specifically in the Raman shift region between 700-1500 cm-1 (Figure 2). The detection limit of SERS coupled with silver nanosubstrates could reach the level of single cells. Significant differences were observed between the spectra of E. coli O157:H7 and S. epidermidis. A mixture test for these two bacterial strains was also conducted (Figure 3). The results of this study indicate that coupled with internal coated silver nanosubstrates, SERS could be a rapid and sensitive method for testing foodborne pathogens.
The results demonstrate that internal coating with silver nanostructures for bacterial cells is a feasible and effective approach to conduct SERS measurement for bacterial samples. Second, coupled with internal-coated silver nanostructures, SERS was able to rapidly detect different water- and foodborne bacteria. Figure 4 shows the SEM pictures of S. epidermidis and E. coli O157:H7 cells internally coated with silver nanostructures. Third, SERS could be used to test mixed bacterial samples for water and food safety control.
Figure 2. Average SERS spectra (n=7) acquired from 3 bacterial species with internal coating treatment in a concentration of 109: S. epidermidis (a), E. coli K-12 (b), E. coli O157:H7 (c)
Figure 3. Average SERS spectra (n=7) acquired from S. epidermidis, E. coli O157:H7 and the 1:1 ratio mixture with a concentration of 109: S. epidermidis (a), the 1:1 ratio mixture of S. epidermidis and E. coli O157:H7 (b), E. coli O157:H7 (c).
Figure 4 Scanning electron microscopy images of bacterial cells internally coated with silver nanostructures: S. epidermidis (a), E. coli O157:H7 (b)
Journal Articles on this Report : 2 Displayed | Download in RIS Format
| Other project views: | All 6 publications | 5 publications in selected types | All 5 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Fan C, Hu Z, Riley LK, Purdy GA, Mustapha A, Lin M. Detecting food-and waterborne viruses by surface-enhanced Raman spectroscopy. Journal of Food Science 2010;75(5):M302-M307. |
R833840 (2009) R833840 (Final) |
Exit |
|
|
Zhang Y, Riley LK, Lin M, Hu Z. Lanthanum-based concentration and microrespirometric detection of microbes in water. Water Research 2010;44(11):3385-3392. |
R833840 (2009) R833840 (Final) |
Exit |
Supplemental Keywords:
drinking water, exposure, waterborne pathogens, microbiology, monitoring, measurement methods, physical processes, health effects, field-based detection, lanthanum, respirometry, Surface Enhanced Raman Spectroscopy, nanotechnology, biotechnology,, RFA, Health, Ecosystem Protection/Environmental Exposure & Risk, Scientific Discipline, PHYSICAL ASPECTS, Water, Health Risk Assessment, Physical Processes, Risk Assessments, Environmental Chemistry, Monitoring/Modeling, Drinking Water, Environmental Monitoring, virulence factor activity relationships, microbial contamination, E. Coli, microbiological organisms, microorganism, measurement, measurement , microbial risk assessment, human exposure, microbial risk management, bacteria monitoring, monitoring, virulence factor biochip, assessment technology, exposure, drinking water contaminants, other - risk assessment, human health risk, detection, exposure and effectsProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.