Grantee Research Project Results
2007 Progress Report: Quantitative Assessment of Pathogens in Drinking Water
EPA Grant Number: R833002Title: Quantitative Assessment of Pathogens in Drinking Water
Investigators: Schwab, Kellogg J. , Halden, Rolf U. , Graczyk, Thaddeus
Institution: The Johns Hopkins University
EPA Project Officer: Aja, Hayley
Project Period: August 25, 2006 through August 24, 2009 (Extended to September 30, 2010)
Project Period Covered by this Report: August 25, 2006 through August 24,2007
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
A major limiting factor in assessing the human health risk of microbial pathogens in raw and finished drinking water is the lack of robust, efficient methods for concentrating, identifying, and quantifying low levels of bacteria, viruses, and protozoa simultaneously, effectively, and rapidly. We will develop a microbial isolation and detection protocol capable of qualitative and quantitative identification of waterborne microbial pathogens by combining the latest high-efficiency filtration technology with rapid and sensitive molecular detection techniques, including quantitative PCR (qPCR), quantitative reverse transcription-PCR (qRT-PCR), fluorescent in situ hybridization (FISH), and matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS). The sensitivity and specificity of the proposed pathogen recovery and detection approach will be directly compared to current USEPA methods via spiking and analysis of raw and finished drinking water samples collected from various water resources and distribution systems. Following method validation, a series of unspiked raw or finished waters (including waters from distribution systems) will be monitored for pathogenic microorganisms to demonstrate the utility of the approach in real world situations.
Progress Summary:
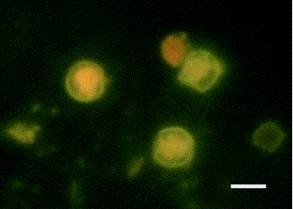
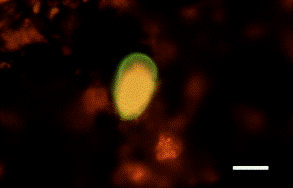
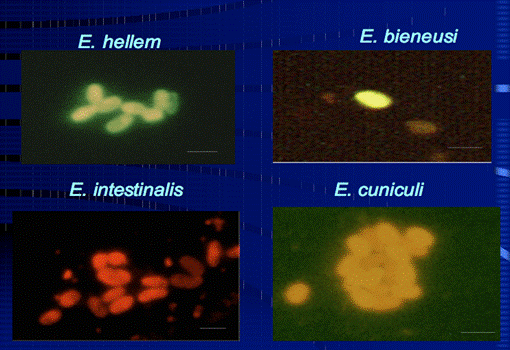
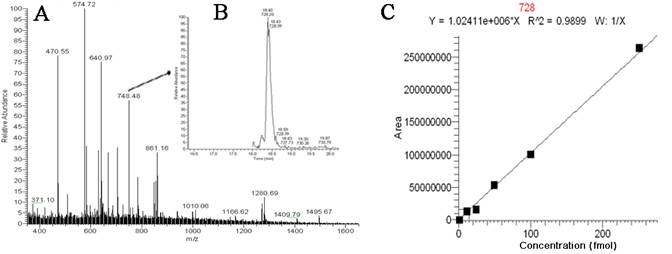
Research during the first 12 months of this project include the optimization and streamlining of several fluorescence in situ hybridization (FISH) methods into two primary assays for the simultaneous detection of 1) Cryptosporidium parvum oocysts, and Giardia lamblia cysts and 2) human-virulent microsporidia spores such as Enterocytozoon bieneusi, Encephalitozoon intestinalis, Encephalitozoon hellem, and Encephalitozoon cuniculi. Efforts have also been made towards the further development of mass spectrometry methods for the detection and quantification of virus capsid proteins, which could allow for estimation of virus concentrations within a sample. Additionally, a bench-scale tangential flow ultrafiltration system has been developed to evaluate the ability of commercially available hollow fiber membranes to simultaneously recover viruses and bacteria from 3 different water types. Research in the development of methods for downstream concentration and detection of these recovered microorganisms is also ongoing. A summary of the research that has either been initiated and/or completed during the current reporting period (August 2006-August 2007) is presented below.
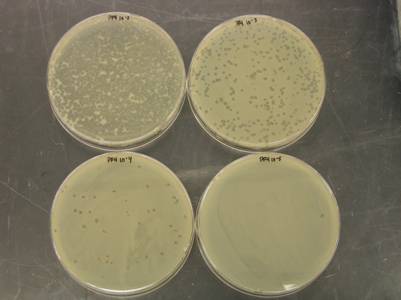
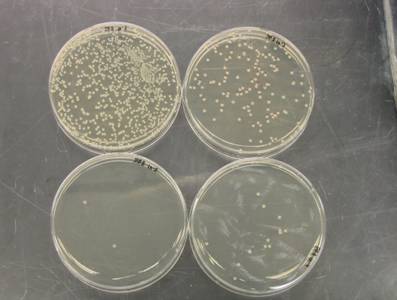
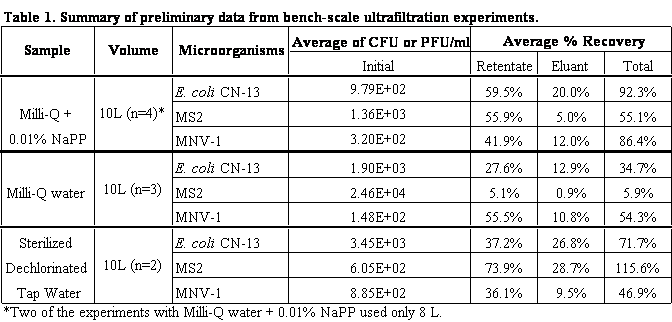
previously in this report.
Future Activities:
The following research will be conducted during the next project period (August 2007 to August 2008):
- Further evaluations of the multiplexed FISH assays will be conducted by spiking 10 L water samples with low concentrations of cysts, oocysts, and spores (1-10) for the detection of G. lamblia, C. parvum, and human-virulent microsporidia, respectively.
- Our mass spectrometry protocol will be tested further for its ability to detect enteric viruses, like norovirus, within a complex matrix such as a human stool extract. Once optimized, MS will be utilized in the detection of microorganisms from the ultrafiltration process in combination with the established culture methods of detection as well as quantitative molecular analysis.
- Evaluation and optimization of the bench-scale ultrafiltration system will continue. The following parameters will be involved:
- The system will be challenged by spiking progressively lower concentrations of E. coli, MNV-1, and MS2 into 10 L and 100 L samples of sterile dechlorinated tap water.
- The retentate, eluant, and secondary concentrate will be analyzed for the microorganisms by both culture and quantitative molecular techniques which have been established in our lab.
- Additional microorganisms will be added to the water samples including poliovirus, norovirus, Hepatitis A virus, adenovirus, enterococci, Clostridium perfringens, Pseudomonas diminuta, Cryptosporidium and Microsporidia. This will help to evaluate the ability of the ultrafilter to recovery a wide variety of microorganisms ranging in size, shape, and type.
- Large volume surface, ground, and distribution system water samples will be collected and spiked with previously mentioned microorganisms. These samples will then be processed using the ultrafiltration setup and detection methods described within this report.
References:
1. Colquhoun, D. R., K. J. Schwab, R. N. Cole, and R. U. Halden. 2006. Detection of norovirus capsid protein in authentic standards and in stool extracts by matrix-assisted laser desorption ionization and nanospray mass spectrometry. Appl Environ Microbiol 72:2749-55
2. DaSilva, A. J., S. B. Slemenda, G. S. Visvesvara, D. A. Schwartz, C. M. Wilcox, S. Wallace, and N. J. Pieniazek. 1997. Detection of Septata intestinalis (Microsporidia) Cali et al. 1993 using polymerase chain reaction primers targeting the small subunit ribosomal RNA coding region. Mol. Diagn. 2:47-52.
3. DaSilva, A. J., F. J. Bornay-Llinares, L. N. S. Moura, S. B. Slemenda, J. L. Tuttle, and N. J. Pieniazek.1999. Fast and reliable extraction of protozoan parasite DNA from fecal specimens. Mol. Diagn. 4:57-64.
4. Deere D, Vesey G, Milner M, Williams K, Ashbolt N, Veal DA. 1998. Rapid method for fluorescent in situ ribosomal RNA labeling of Cryptosporidium parvum. J Appl Microbiol 85: 807-818.
5. deGroote, M. A., G. S. Visvesvara, M. L. Wilson, N. J. Pieniazek, S. B. Slemenda, A. J. DaSilva, G. J. Leitch, R. T. Bryan, and R. Reeves. 1995. Polymerase chain reaction and culture confirmation of disseminated Encephalitozoon cuniculi in patient with AIDS: successful therapy with albendazole. J. Infect. Dis. 171:1375-1378.
6. Dorsch MR, Veal DA, 2001. Oligonucleotide probes for specific detection of Giardia lamblia cysts by fluorescent in situ hybridization. J Appl Microbiol 90: 836-842.
7. Gerber, S. A., J. Rush, O. Stemman, M. W. Kirschner, and S. P. Gygi. 2003. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc Natl Acad Sci U S A 100:6940-5.
8. Graczyk, T. K., J. Bosco-Nizeyi, A. J. DaSilva, L. N. S. Moura, N. J. Pieniazek, M. R. Cranfield, and H. D. A. Lindguist. 2002. A single genotype of Encephalitozoon intestinalis infects free-ranging gorillas and people sharing their habitats, Uganda. Parasitol. Res. 88:926-931
9. Graczyk, T. K., D. B. Conn, F. Lucy, D. Minchin, L. Tamang, L. N. S. Moura, and A. J. DaSilva. 2004. Human waterborne parasites in zebra mussels (Dreissena polymorpha) from the Shannon River drainage, Ireland. Parasitol. Res. 93:389-391
10. Halden, R. U., and D. H. Paull. 2004. Analysis of triclocarban in aquatic samples by liquid chromatography electrospray ionization mass spectrometry. Environmental Science & Technology 38:4849-4855.
11. Hester, F. D., H. D. A. Linquist, A. M. Bobst, and F. W. Schaffer. 2000. Fluorescent in situ detection of Encephalitozoon hellem spores with a 6carboxyfluorescein-labeled ribosomal RNA-targeted oligonucleotide probe. J. Eukaryot. Microbiol. 47:299-308.
12. Slodkowicz-Kowalska, A., T. K. Graczyk, L. Tamang, S. Jedrzejewski, A. Nowosad, P. Zduniak, P. Solarczyk, A. S. Girouard, and A. C. Majewska. 2006. Microsporidia species known to infect humans are present in aquatic birds; implications for transmission via water? Appl. Environ. Microbiol. 72:4540-4544.
13. Vesey G, Ashbolt N, Fricker EJ, Deere D, William KL, Veal DA, Dorsch M, 1998. The use of a ribosomal RNA targeted oligonucleotide probe for fluorescent labelling of viable Cryptosporidium parvum oocysts. J Appl Microbiol 85: 429-440.
14. Visvesvara, G. S., A. J. DaSilva, G. P. Croppo, N. J. Pieniazek, G. J. Leitch, D. Ferguson, H. Moura, S. Wallace, S. B. Slemenda, and I. Tyrrell. 1995. In vitro culture and serologic and molecular identification of Septata intestinalis isolated from urine of a patient with AIDS. J. Clin. Microbiol. 33:930-936.
15. Wobus, C. E., S. M. Karst, L. B. Thackray, K. O. Chang, S. V. Sosnovtsev, G. Belliot, A. Krug, J. M. Mackenzie, K. Y. Green, and H. W. Virgin. 2004. Replication of a Norovirus in cell culture reveals a tropism for dendritic cells and macrophages. PLoS Biol. 2:e432.
Journal Articles on this Report : 2 Displayed | Download in RIS Format
| Other project views: | All 8 publications | 4 publications in selected types | All 4 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Graczyk TK, Sunderland D, Rule AM, da Silva AJ, Moura INS, Tamang L, Girouard AS, Schwab KJ, Breysse PN. Urban feral pigeons (Columba livia) as a source for air- and waterborne contamination with Enterocytozoon bieneusi spores. Applied and Environmental Microbiology 2007;73(13):4357-4358. |
R833002 (2007) R833002 (2008) R833002 (Final) |
Exit Exit Exit |
|
|
Graczyk TK, Majewska AC, Schwab KJ. The role of birds in dissemination of human waterborne enteropathogens. Trends in Parasitology 2008;24(2):55-59. |
R833002 (2007) R833002 (2008) R833002 (Final) |
Exit Exit |
Supplemental Keywords:
drinking water, human health, molecular detection, monitoring, quantitative PCR, Fish, proteomics, mass spectrometry, pathogens, viruses, protozoa, bacteria, exposure, risk assessment, environmental microbiology, Maryland (MD), California (CA), Pollutants/Toxics, Water, International Cooperation, Scientific Discipline, RFA, Physical Aspects, Drinking Water, Physical Processes, Environmental Engineering, Environmental Chemistry, Environmental Monitoring, Microorganisms, bacteria, drinking water contaminants, mass spectrometry, pathogenic protozoa, molecular detection, viruses, pathogens, ultrafiltration, exposure, human health,, RFA, Scientific Discipline, PHYSICAL ASPECTS, INTERNATIONAL COOPERATION, Water, POLLUTANTS/TOXICS, Physical Processes, Environmental Chemistry, Microorganisms, Drinking Water, Environmental Engineering, Environmental Monitoring, human health, mass spectrometry, viruses, ultrafiltration, water quality, pathogenic protozoa, pathogens, drinking water contaminants, exposure, drinking water monitoring, molecular detectionProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.