Grantee Research Project Results
Final Report: Pathway Analysis Core
EPA Grant Number: R835737C005Subproject: this is subproject number 005 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: Human Models for Analysis of Pathways (H MAPs) Center
Center Director: Murphy, William L
Title: Pathway Analysis Core
Investigators: Roy, Sushmita
Institution: University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
The overarching goal of the Pathway Analysis Core is to provide an integrated understanding of cellular response to environmental perturbations by developing novel network biology tools that are applicable to variety of cell types and tissue models. We will apply these methods to dissect regulatory networks of tissue and cell-type specific responses to small molecules and toxins from the central nervous system and liver hepatocyte development. Specifically, the core is developing methods to analyze data measured by the Brain MAPS and Liver MAPS projects.
Summary/Accomplishments (Outputs/Outcomes):
1. Inference of regulatory networks of early neural stem cell differentiation from BRAIN MAPs RNA-seq data.
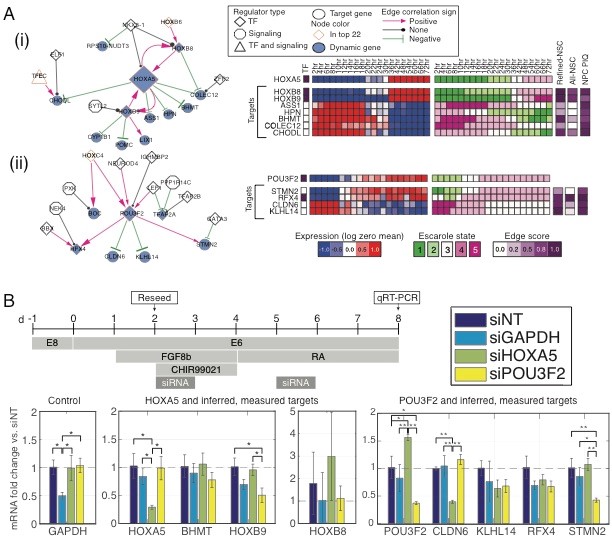
We have developed a novel computational approach to identify regulatory networks and their dynamics from RNA-seq time course data. Our approach includes a novel method, Escarole, to analyze time course gene expression to identify dynamic genes and to infer a context-specific transcriptional regulatory network. We applied this approach to the RNA-seq dataset collected by BRAIN MAPs to infer regulatory network dynamics during early hindbrain differentiation (Fig 1). We used the inferred network to prioritize regulators and validated the targets of two candidate prioritized regulators, HOXA5 and POU3F2, using siRNA followed by qPCR of the targets. This work is currently under revision in Cell Systems, "Inferring dynamic regulatory programs in non-stationary expression time courses with applications to early human neural development".
Fig 1. Inference and validation of regulatory networks in hind brain neural stem cells. A. Subnetworks showing neighborhoods of top-ranked regulators HOXA5 (i, #1), POU3F2 (ii, #5). Heatmaps on the right show the gene expression (mean-centered), Escarole states of the regulators and their dynamic target genes, and the score of the edge from different inferred or sequence-specific networks. B. qRTPCR results showing mRNA levels for candidate regulatory network target genes or control genes following treatment with non-targeting siRNA (siNT, dark blue) or RNAi-mediated knockdown of GAPDH (siGAPDH, aqua), HOXA (siHOXA5, green), or POU3F2 (siPOU3F2, yellow). The x-axis labels indicate the gene being measured with qRT-PCR. Error bars denote standard deviation.
2. Integrative analysis of RNA-seq and ATAC-seq data from LIVER MAPs.
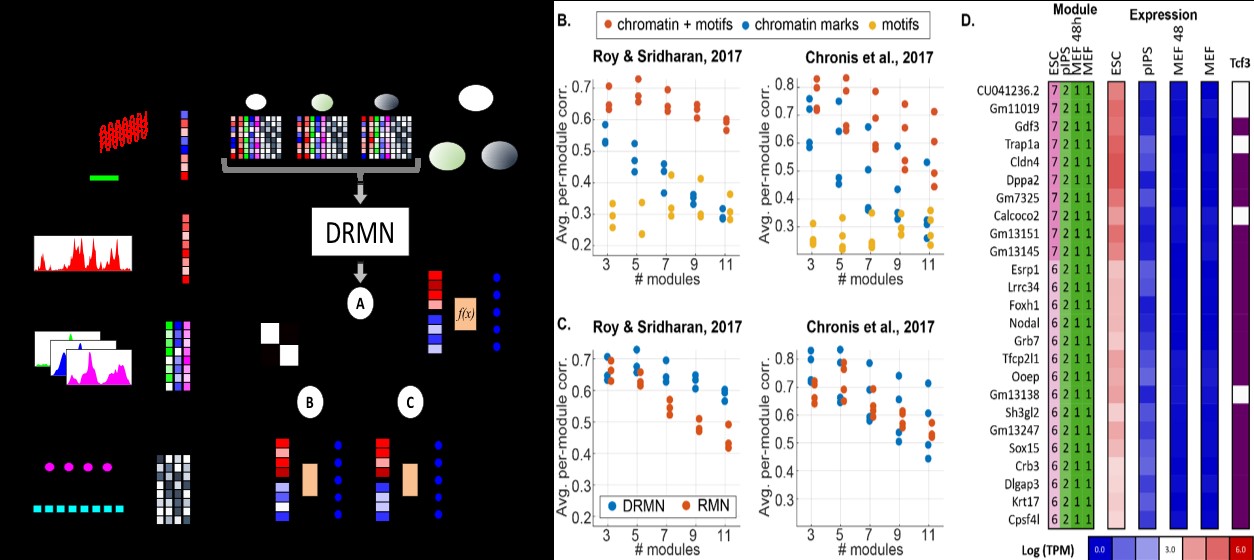
We are developing a novel approach, Dynamic Regulatory Module Networks (DRMNs) to analyze parallel transcriptomic and epigenomic time courses (Fig 2A), which would be applicable to the datasets being generated by the LIVER MAPs project to decipher the regulatory network governing the hepatocyte state. DRMNs are based on a non-stationary probabilistic graphical model, where the regulatory network at each time point or cell type is represented by a concise set of gene expression modules and their associated regulatory programs. Inputs to DRMNs are cell type specific gene expression levels, a relationship among the cell types (e.g., time or lineage), sequence-based motif network and cell type specific epigenetic data (e.g. chromatin marks and accessibility). As preliminary work, we applied DRMN to two different data sets (Roy & Sridharan 2017, Chronis et al 2017) to study the regulatory network dynamics during cellular reprogramming. Each data set included multiple activating and repressive chromatin marks and gene expression for the starting differentiated cell type (mouse embryonic fibroblasts), the pluripotent cell type, and one or more partially reprogrammed intermediate cell types, which we integrated with sequence specific motifs of 342 transcription factors identified within +/-2kbp of the gene's transcription start site (TSS). For both data sets, integrating sequence motifs with chromatin marks was important for accurately predicting expression, suggesting that sequence motifs capture additional variation not explained by chromatin mark data (Fig. 2B). DRMN outperformed or matched the performance of a baseline model, that learned predictive programs for each cell type independently (Fig. 2C). Inspection of the inferred regulatory modules revealed cell type-specific enrichment of motifs of known regulators of pluripotency, including c-Myc, Klf4, Tcf3 (Fig. 2D), and Esrrb. In conclusion, DRMN is able to predict cell type-specific expression from cis-regulatory signals, and multi-task learning improves performance. We are currently applying the DRMN approach to datasets generated by the Thomson lab. In particular, the Thomson has generated ATAC-seq and RNA-seq data in mouse hepatocytes over 16 time points over a period of 36 hours. Application of DRMNs to this time course data is expected to identify important regulators associated with expression state changes during dedifferentiation, that can be tested functionally for their impact on maintenance of the hepatocyte state.
Fig 2 A. Dynamic Regulatory Module Networks (DRMN) schematic. DRMNs take as input cell type gene expression, a cell lineage relationship, and additional cell type specific epigenetic data (ATAC-seq, ChIP-seq for chromatin marks), and sequence specific motifs. DRMNs produce cell type specific expression modules and their regulatory programs (f(x)) and transition matrices. B. Performance of different regulatory signal data sets using DRMNs. Each panel shows results from one of the two reprogramming studies. Expression prediction in cross-validation is scored by the correlation between true and predicted expression for each module, averaged across all modules. Each dot is one cell type. C. Performance of DRMN versus a base line model (RMN) using motif and chromatin regulatory features. D. A set of genes with ESC-specific expression (Chronis et al, 2017), overrepresented for motif instances of Tcf3. Modules are numbered from 1 (low expression) to 9 (high expression). The Tcf3 column indicates presence of the motif within 2kbp of the gene's TSS.
Journal Articles on this Report : 1 Displayed | Download in RIS Format
| Other subproject views: | All 17 publications | 9 publications in selected types | All 8 journal articles |
|---|---|---|---|
| Other center views: | All 215 publications | 82 publications in selected types | All 81 journal articles |
| Type | Citation | ||
|---|---|---|---|
|
|
Wheeler HE, Ploch SA, Barberia AN, Bonazzola R, Andaleon A, Fotuhi S, Saha A, Battle A, Roy S, Im HK. Imputed gene associations identify replicable trans-acting genes enriched in transcription pathways and complex traits. Genetic Epidemiology 2019. |
R835737C005 (Final) |
|
Progress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 Human Models for Analysis of Pathways (H MAPs) Center Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
8 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center