Grantee Research Project Results
2016 Progress Report: Pathway Analysis Core
EPA Grant Number: R835737C005Subproject: this is subproject number 005 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: Human Models for Analysis of Pathways (H MAPs) Center
Center Director: Murphy, William L
Title: Pathway Analysis Core
Investigators: Roy, Sushmita
Institution: University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
Project Period Covered by this Report: December 1, 2014 through February 28,2016
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
The overarching goal of our Center's Pathway Analysis Core is to provide an integrated understanding of cellular response to environmental perturbations by developing novel network biology tools that are applicable to a variety of cell types and tissue models. Our specific aims are to develop methods that (1) integrate temporal dynamics in regulatory network model reconstruction and (2) identify subnetworks that are perturbed under exposure to changing environmental stimuli. We will apply these methods to dissect regulatory networks of tissue and cell-type specific responses to small molecules and toxins from the central nervous system and liver hepatocyte development. Specifically, the core is developing methods to analyze data measured by the Brain MAPS and Liver MAPS projects.
Progress Summary:
Aim 1. Application of dynamic network and module inference methods to Brain MAPs RNA sequenced (RNA-seq) data. The Brain MAPs RNA-seq data profiles region-specific neural stem cell (NSC) populations at discrete time points corresponding to consecutive spatial locations in the hindbrain. Our analyses were motivated by two goals: (A) to characterize gene expression dynamics across the NSC time course, and (B) to identify genes and subnetworks that establish and drive transitions between populations.
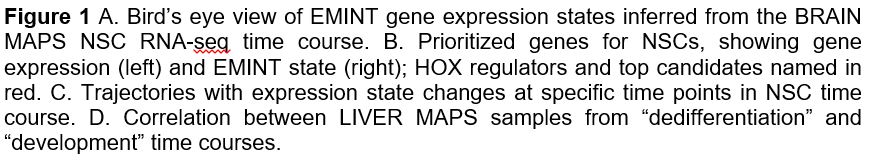
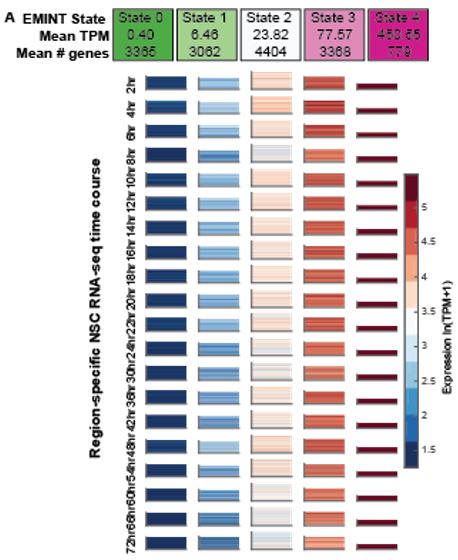
Characterizing gene expression dynamics in NSC time course. We have applied EMINT (Expression Module Inference on Trees), a new approach to examine expression time courses. EMINT takes as input a genome-wide expression profile from each time point and produces a clustering for genes within each time point, where each cluster is interpreted as a discrete expression state, and the states are matched one-to-one across time points. We used EMINT to discretize gene expression values into five distinct states at each time point (Figure 1A). By inspecting the state assignments, we identified which genes are in which state (high or low) at a particular time and assessed how genes change their state over time. Gene ontology enrichment of these states shows that the induced state is associated with translation and cell cycle, and the repressed state is associated with later neural differentiation pathways, which is consistent with the overall state of early NSCs.
Prediction of novel genes and subnetworks associated with the NSC state specification. To prioritize novel genes for association with specific time points or subsets of time points we focused on genes that change their EMINT state. About two-thirds of the genes did not change their state, reflecting the high level of similarity of the time points. We identified 29 dynamic genes with a state range of at least 3, including one phosphatase (SGGP2) and six transcription factors, which may be important for distinguishing the cell types.
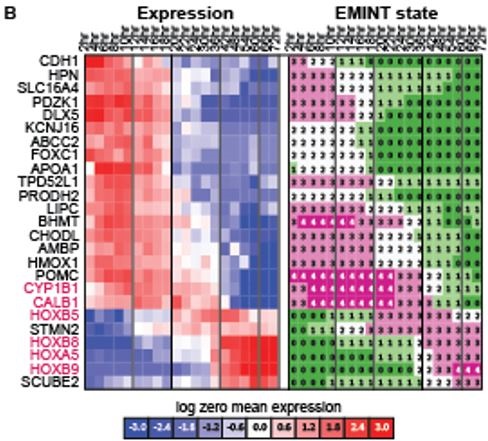
In parallel, we identified genes with associated variation in expression (ANOVA p < 0.001) and ranked genes by their standard deviation in EMINT state assignment (Figure 1B). In collaboration with the Ashton lab, we have examined these gene sets and identified several important genes to test using in situ hybridization in chicken embryo, including four HOX genes and two candidates for novel function in the developing hindbrain (CALB1, CYP1B1).
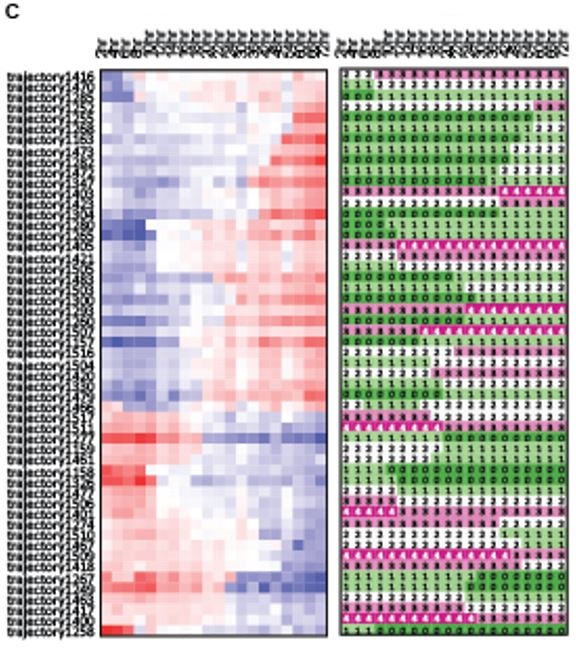
We also identified trajectories that are shared by groups of genes that change their state coordinately at specific times (Figure 1C). These trajectories are hypothesized to be effector genes of region-specific regulators.
Identification of gene regulatory networks associated with region-specific NSC expression. We are using tools developed in the Roy lab to infer an NSC-specific network from publicly available RNA-seq data in neural lineage cell types. With input from Dr. Ashton, we have created an initial RNA-seq dataset including 84 samples, 28 RNA-seq samples from the Gene Expression Omnibus, 36 samples from the Thomson laboratory and 20 samples from the Brain MAPs project. By applying a network inference tool developed in our lab (MERLIN), we have obtained an initial neural lineage regulatory network with 30,377 connections between 502 transcription factors to 9,058 target genes.
2. Adapt and apply network information flow methods to examine tissue-specific response to different toxins. We have not had a concrete research question to which to apply these approaches in collaboration with Brain MAPs or Liver MAPs thus far.
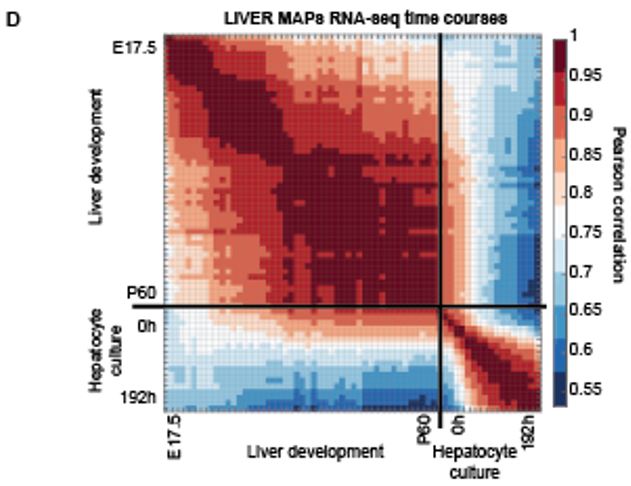
3. Initial transcriptomic and epigenomic data analysis from LIVER MAPs project. The Thomson lab gave us two RNA-seq datasets: a “dedifferentiation” time course of primary mouse hepatocytes in culture, and a “development” time course of the developing mouse liver from before birth to adulthood. We performed a principal components analysis (PCA) to examine the major sources of variation in the data. The first PC captures 60.5% of the variance in the dedifferentiation time course and 58% in the development time course indicating the presence of a strong temporal component. Fibroblast genes were positively strongly correlated with the first PC in the differentiation time course indicating the loss of mature phenotype. The early time points in the differentiation time course and late samples in the development time course appear highly correlated suggesting these time courses represent similar end and begin points (Figure 1D).
We did not begin epigenomic analysis as the data are not yet available. We performed an extensive literature review of recent approaches to map regulatory networks in cell lineages by integrating expression, epigenomic and sequence data (Chasman & Roy, under review).
Future Activities:
Our future activities include:
- Finalize EMINT analysis, network inference, interpretation, and gene prioritization for NSCs for validation by the Ashton lab.
- Draft manuscript with the Ashton lab to describe the EMINT approach and application.
- Continue analysis of RNA-seq and epigenomic data as available from LIVER MAPs.
Journal Articles on this Report : 2 Displayed | Download in RIS Format
| Other subproject views: | All 17 publications | 9 publications in selected types | All 8 journal articles |
|---|---|---|---|
| Other center views: | All 215 publications | 82 publications in selected types | All 81 journal articles |
| Type | Citation | ||
|---|---|---|---|
|
|
Chasman D, Roy S. Inference of cell type specific regulatory networks on mammalian lineages. Current Opinion in Systems Biology 2017;2:130-139. |
R835737 (2017) R835737C004 (2017) R835737C005 (2016) R835737C005 (2017) |
Exit |
|
|
Roy S, Sridharan R. Chromatin module inference on cellular trajectories identifies key transition points and poised epigenetic states in diverse developmental processes. Genome Research 2017;27(7):1250-1262. |
R835737 (2017) R835737C005 (2016) |
Exit Exit Exit |
Supplemental Keywords:
regulatory networks, transcriptional modules, chromatin state, dynamic networks, temporal networksProgress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 Human Models for Analysis of Pathways (H MAPs) Center Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
8 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center