Grantee Research Project Results
Final Report: Liver MAPs
EPA Grant Number: R835737C001Subproject: this is subproject number 001 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: The Morgridge Institute for Research
Center Director: Thomson, James
Title: Liver MAPs
Investigators: Thomson, James
Institution: The Morgridge Institute for Research , University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
- OBJECTIVE 1. Identify regulatory networks that control liver maturation.
- OBJECTIVE 2. Screen for factors that promote human ES/iPS cell-derived hepatocyte maturation.
- OBJECTIVE 3. Optimize 3D organotypic cultures of human ES/iPS cell-derived hepatocytes with mature metabolic function.
Summary/Accomplishments (Outputs/Outcomes):
OBJECTIVE 1
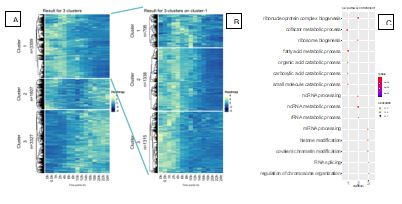
Figure 1. A. K-mean clustering of gene expression across the time course. B. K-mean clustering of genes in cluster 1 of A. C. Gene Ontology analysis of clusters from A, only the Biological Porcesses ontology in shown.
| A) | Cluster | B) | Cluster |
| Alb | 1 | Alb | 1.1 |
| Cyp1a2 | 1 | Cyp1a2 | 1.2 |
| Cyp2e1 | 1 | Cyp2e1 | 1.2 |
| Cyp2f2 | 1 | Cyp2f2 | 1.2 |
| Cup3a11 | 1 | Cyp3a11 | 1.2 |
| G6pc | 1 | G6pc | 1.3 |
| Gjb1 | 1 | Gjb1 | 1.2 |
| Tat | 1 | Tat | 1.2 |
| Tdo2 | 1 | Tdo2 | 1.2 |
| Ugt1a6a | 1 | Ugt1a6a | 1.2 |
| Ugt1a6b | 1 | Ugt1a6b | 1.1 |
| Ug2b1 | 1 | Ug2b1 | 1.2 |
| Ugt2b36 | 1 | Ugt2b36 | 1.2 |
| Ugt2b5 | 1 | Ugt2b5 | 1.2 |
Table1. A. Cluster from 1A that hepatocyte marker genes are found in. B. Cluster from 1B that hepatocyte marker genes are found in.
Due to requirement of high cell numbers for DNase-seq, we focused on ATAC-seq, which is robust and requires only 5 X 104 input cells compared to the 2 X 107cells for DNase-seq. Concurrently we also performed matched RNA-Seq to obtain the transcriptomic landscape of de-differentiating hepatocytes in our time course experiment. We collected nucleic acids every 2 hours for the first 24 hours from primary hepatocytes in tissue culture and performed both ATAC-Seq and RNA-Seq. We have completed a detailed examination of the RNA-Seq experiment and are working to analyze the ATAC-Seq data. Analysis of the RNA-seq results revealed 3 main clusters of genes that capture the majority of variation in the dataset (Figure 1). Figure 1A shows that the 3 clusters consists of 1) genes that are high at 0 hours and subsequently lose their expression 2) genes that are relatively stable across the time course with a drop in expression at around 10 hours and 3) genes that have low expression early in the time course but increase towards the end. When we look at several marker genes of mature hepatocytes it is apparent that they all cluster together in cluster 1 (Table 1A). Gene ontology analysis (Figure 1C) determined that genes in cluster 1 are enriched in metabolic processes related to hepatocyte function; cluster 2 consisted of genes related to ribosome and transcription and genes in cluster 3 are related to major changes in nuclear organization such as chromosome and histone modifications.
Cluster 1 contained genes of interest and further analysis was performed on this cluster which showed 3 major sub-clusters. Sub-cluster 3 was comprised of genes that are down regulated the fastest followed by cluster 2 where genes were down regulated slower and cluster 1 included genes that were downregulated the slowest (Figure 1B). We confirmed the clustering by looking at individual genes. We found slower downregulated genes such as albumin in cluster 1, cytochrome P450 genes, which are downregulated faster are found in cluster 2 and very fast downregulated genes such as glucose-6-phosphatase catalytic subunit are found in cluster 3 (Table 1B).
We are in the process of developing a pipeline for the analysis of the ATAC-Seq data. We plan to cluster peaks across the time course and perform motif analysis of clusters of peaks to identify important transcription factors. We are also going to use the clustering done on RNA-Seq data to identify groups of ATAC-Seq peaks that are associated with the clusters identified in figure 1.
OBJECTIVE 2
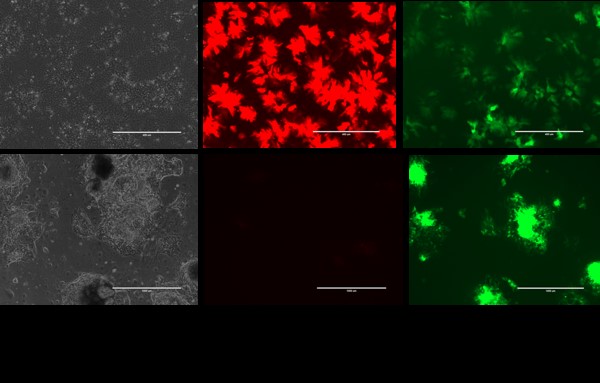
Figure 1. Top: H9 ALB-GFP/AFP-td tomato hepatocytes. Bottom: SHPCs derived from above.
We have developed culture methods to induce proliferation of primary human hepatocytes (PHHs) and pluripotent stem cell-derived hepatocytes via generation of small hepatocyte proliferating cells (SHPCs). Our culture conditions not only maintain mature phenotype of PHHs over multiple passages, but also induce maturity in embryonic stem cell-derived hepatocytes which is our goal and is reported here.
| GENE | Uncultured PHH | Dual hepatocyte | Dual hepatocyte SHPC |
|---|---|---|---|
| ALB | 50870.7 | 39376.26 | 35351.04 |
| AFP | 17.79 | 40549.13 | 273.38 |
| CYP3A4 | 14.77 | 0 | 67.19 |
| CYP3A7 | 5.27 | 811.33 | 16.02 |
| CYP2C9 | 652.99 | 57.35 | 311.98 |
| CYP2E1 | 2556.93 | 0.35 | 586.67 |
| UGT1A9 | 43.08 | 9.43 | 153.45 |
| UGT1A1 | 91.38 | 25.93 | 783.37 |
| SERPINA1 | 17172.69 | 4882.91 | 27901.25 |
Hepatocytes differentiated from a dual reporter ES cell line (H9-Albumin-GFP/Alpha-fetoprotein-TdTomato) expressed high levels of the fetal marker alpha-fetoprotein (AFP) and low levels of the mature hepatocyte marker, Albumin (Alb). SHPCs generated from the above hepatocytes displayed higher Alb and no detectable AFP marker (Fig. 1). RNA-Seq showed up-regulation of mature hepatic genes including those important for xenobiotic metabolism along with down-regulation of fetal markers such as AFP (Table 1).
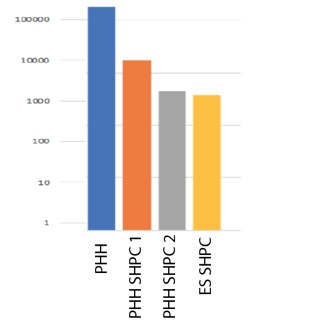
Figure 2. Humand Alb (ng/ml) in mouse blood
SHPCs derived from both ES cells and PHHs were transplanted into Tk-NOG mice via injection into the spleen. ELISA for human specific albumin in transplanted mouse blood confirmed their engraftment (Figure 2). We plan to test xenobiotic metabolism by ES-derived hepatocytes by LCMSMS for common drugs that are metabolized by phase I and II enzymes.
OBJECTIVE 3
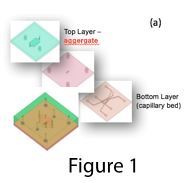
We had previously reported that three-dimensional culture of human iPS-derived hepatocytes improved their mature gene expression and function. We had also shown that coculture of iPS-hepatocytes (iHEPs) with human sinusoidal endothelial cells (ECs) significantly improved their functionality. We had observed that ECs also formed organized vascular network in the iHEP aggregates. These vascularized iHEP aggregates have the potential to be incorporated to the capillary bed in microfluidic platform for generation of scalable multiplexed drug screening chips where the drugs could be administered through the vasculature and their effects on the hepatic compartment could be monitored via imaging.
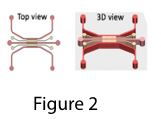
To achieve this goal, we used a three-layer microfluidic platform in the earliest attempt where vascular network were formed in the bottom layer and iHEP aggregates were placed in the top layer and vascular network were encouraged to migrate to the top layer by VEGF gradient induction (Figure 1). Preliminary results showed successful engagement of vascularized aggregates with the capillary bed and increased life span of aggregates. However, due to stacking nature of the device, it was hard to image the top layer of the device where the iHEP aggregates are.
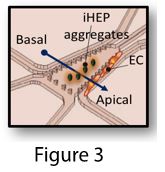
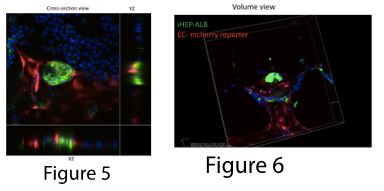
To address the imaging issue, we modified the first design. In the new design, vascular networks and aggregates are on the same layer (Figure 2), where iHEP aggregates are in the middle channels and endothelial cells are in the side channels (Figure 3). We encouraged vascular sprouting toward iHEP aggregates (the middle channel) by creating an apical-basal flow (Figure 4). Our results showed successful integration of endothelial sprouts with the iHEP aggregates in the middle channel (Figure 5 and 6). However, the natural extracellular matrix (Matrigel) was not stable for carrying the experiment over 10 days. Matrigel disintegration results in collapse of 3D iHEP aggregates and spreading in a 2D layer format. Therefore, we are explored ssynthetic polyethylene glycol (PEG) based hydrogels as a substitute for Matrigel. Previously, we showed that PEG-norbornene hydrogel modified with RGD peptide is permissive for capillary network formation (Ardalani et al., Acta Biomater. 35 (2016) 32-41), so we are screening for the optimal formulation of PEG-NB with different pendant groups that would be suitable for both hepatocytes and endothelial cells
Journal Articles:
No journal articles submitted with this report: View all 8 publications for this subprojectProgress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 The Morgridge Institute for Research Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
4 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center