Grantee Research Project Results
2015 Progress Report: Liver MAPs
EPA Grant Number: R835737C001Subproject: this is subproject number 001 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: The Morgridge Institute for Research
Center Director: Thomson, James
Title: Liver MAPs
Investigators: Thomson, James
Institution: The Morgridge Institute for Research
Current Institution: The Morgridge Institute for Research , University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
Project Period Covered by this Report: December 1, 2014 through November 30,2015
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Progress Summary:
OBJECTIVE 1. Identify regulatory networks that control liver maturation. We are performing detailed time series RNA-seq during mouse liver development to select time points during hepatocyte maturation for use in chromatin immunoprecipitation sequencing (ChIP-seq) against six histone modifications shown to be sufficient to identify promoters and enhancers, as well for DNase-seq to identify transcription factors binding to regulatory elements. We are collecting livers daily from embryonic day 17.5 to postnatal day 42 and then every 5 days from day 45 to day 60. We will exclude livers from female mice to avoid variation caused by the estrus cycle. For time points when sex cannot be determined, we will perform sex identification using PCR. At this time, we have collected 35 of 48 time points. Samples will be sequenced and analyzed upon completion of the timeline.
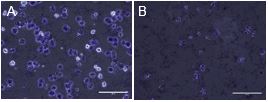
We are in the process of optimizing the next step of DNase-seq and have standardized purification of nuclei from hepatocytes of adult mice with optimum NP40 concentration (Fig. 1). We have further standardized DNase digestion critical for optimal fragmentation of hypersensitive sits (Fig. 2.).
Fig. 1. Hepatocytes before (A) nuclei are isolated (0% NP40) and after optimal nuclei isolation (0.25% NP40) (B) where the cell membrane is dissolved and only the nuclei remain.
Fig. 2. Ratio of two hypersensitive sites (albumin and RASSf6) against two inactive sights by qPCR to determine optimal fragmentation.
OBJECTIVE 2. Screen for factors that promote human embryonic stem/induced pluripotent stem (ES/iPS) cell-derived hepatocyte maturation. We have taken a three-pronged approach to screen for small molecules to promote maturation: (1) We have generated a dual reporter human ES cell clone that expresses tdTomato fluorescent-protein driven by the fetal hepatic gene alpha-fetoprotein (AFP) locus along with green fluorescent-protein driven by the mature hepatic gene albumin (ALB) locus. We have differentiated this clone to hepatocytes and performed a pilot screen of a library of 160 small molecules of known function that interact with key cellular pathways, designed in house on a customized robotic workstation in a sterile controlled atmosphere. We have treated the dual reporter clone-derived hepatocytes with the library for 7 days and have quantified green fluorescent protein (GFP) and tdTomato in the cells using an automated imaging system (Biostation CT, Nikon) designed specifically for live cell high-throughput fluorescence time-lapse imaging. Images are processed with CL-Quant software (DRVision Technologies) that automates fluorescence intensities in a high-throughput workflow. Visual inspection post-screen has identified several candidates. We are in the process of completing image analysis and shall follow up on the identified candidates by RNA-seq confirmation of a larger cohort of fetal and adult hepatic genes. (2) With the help of independent contract research organization InGenious Targeting Laboratory, we have generated a targeted double knock-in mouse ES cell line with the P. pyralis luciferase gene regulated by the AFP locus and the R. reniformis luciferase gene regulated by the ALB locus. We will use the new mouse ES cell dual reporter lines to develop defined hepatocyte differentiation conditions that we will use to screen for molecules that increase maturation using our robotic workstation and automated imaging set-up. (3) In contrast with InGenious Targeting Laboratory, we have generated chimeric mice using the double knock-in ES cell line described above. Once we receive the founder mice and establish a colony, we will isolate hepatocytes from adult livers and use our robotic screen to identify small molecules that will help retain the maturity of adult hepatocytes in culture, where they de-differentiate and lose adult phenotype rapidly.
Fig. 3. A simple 3D aggregate culture of iCell hepatocytes (CDI, Madision, WI). Improved mature hepatic markers (albumin, HNf4a, alpha 1 antitrypsin) concomitantly down-regulating fetal marker alpha fetoprotein (B) and maintaining phase I and II enzymes in 10 days (C).
Fig. 4. Confocal image of a representative aggregate of hepatocytes and iCell cells cultured for 10 days. staining shows endothelial cells assembling to form vascular cords.
OBJECTIVE 3. Optimize 3D organotypic cultures of human ES cell-derived hepatocytes with mature metabolic function. We followed up on our recent observation that suspension culture of human ES cell-derived hepatocytes in 3D aggregates significantly improved metabolic activities and have optimized their co-culture with non-parenchymal liver cells to further improve metabolic functions. A simple 3D aggregate culture of iCell hepatocytes (CDI, Madison, WI) (Fig. 3 A) improved mature hepatic markers (albumin, HNf4α, alpha 1 antitrypsin) while down-regulating fetal marker AFP and maintaining phase I and II enzymes in 10 days (Fig. 3 B).
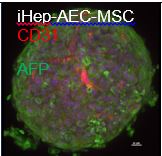
We also optimized the co-culture media for survival of all liver cell types and formed aggregates of iCell hepatocytes (CDI, Madison, WI), iCell endothelial cells (CDI, Madison, WI), or iPSC-derived arterial endothelial cells and iPSC-derived mesenchymal stem cells (Fig. 4) that further increased mature hepatic markers and phase I and phase II enzymes compared to iHep 3D monolayer culture (Fig. 5).
Fig. 5. Optimized co-culture media further increased mature hepatic markers and phase I and phase II enzymes compared with iHep 3D monolayer culture.
Journal Articles:
No journal articles submitted with this report: View all 8 publications for this subprojectSupplemental Keywords:
Hepatoxicity, liver metabolism, toxicology, drug development, high throughput screening, combinatorial screening, phenotypic screening, microphysiological systemProgress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 The Morgridge Institute for Research Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
4 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center