Grantee Research Project Results
Final Report: Cancer MAPs: A 3D Organotypic Microfluidic Culture System to Identify Chemicals that Impact Progression and Development of Breast Cancer
EPA Grant Number: R835737C003Subproject: this is subproject number 003 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: Human Models for Analysis of Pathways (H MAPs) Center
Center Director: Murphy, William L
Title: Cancer MAPs: A 3D Organotypic Microfluidic Culture System to Identify Chemicals that Impact Progression and Development of Breast Cancer
Investigators: Beebe, David , Murphy, William L
Institution: University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
To produce a 3D organotypic culture model of ductal breast cancer that is compatible with higher throughput screening and high-content screening approaches to discern effects of chemicals on breast cancer development and progression.
Aim 1. Optimize and automate our microfluidic 3Din vitroductal breast cancer model to be used for chemical library screening.
Aim 2. Develop an AOP based model of estrogen-receptor (ER) mediated invasive ductal carcinoma (IDC).
Aim 3. Conduct high throughput screens using chemicals from the ToxCast library to identify chemicals that promote ER-mediated and non ER-mediated IDC.
Summary/Accomplishments (Outputs/Outcomes):
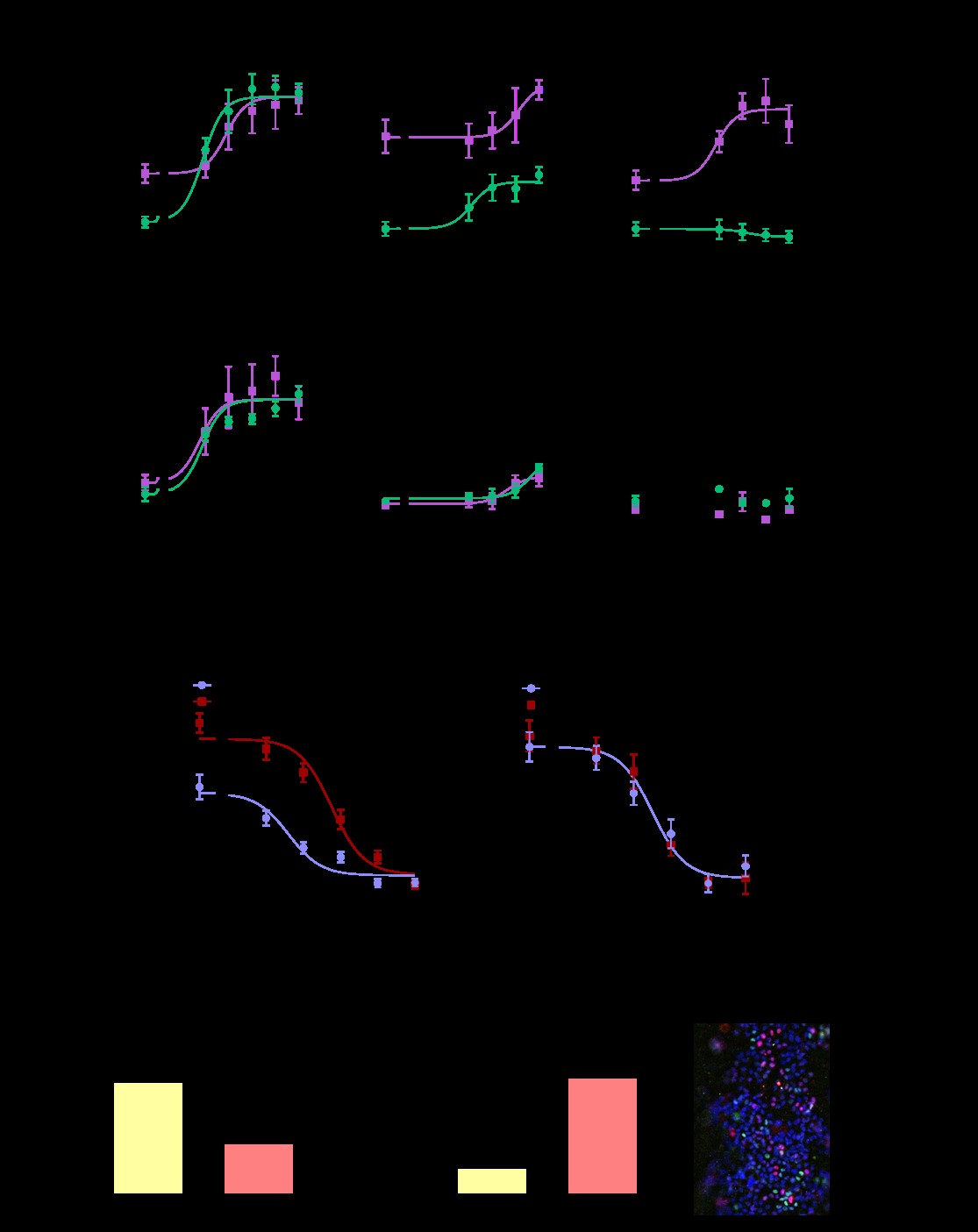
We first tested the hypothesis that responses to environmental chemicals would differ depending on if MCF7s were grown as a 2D monoculture (e.g. a traditional screening model), compared to when grown as biomimetic ducts co-cultured with adipose stromal cells. We tested chemicals that had been identified by the Toxcast database to disrupt the estrogen receptor and/or the estrogen-producing enzyme aromatase. Each test chemical was evaluated at five different concentrations alone and in combination with 1 nM testosterone. 1 nM of testosterone was chosen because the primary source of estrogen in post-menopausal women occurs locally through aromatase metabolizing T to E2, and 1 nM is a physiologically relevant concentration for post-menopausal woman. Some chemicals, such as fenarimol (Fig. 1A) and diethylstilbestrol (Fig. 1B), had similar effects in the traditional model compared to the organotypic model. Other chemicals, such as the organotin compound tributyltin chloride (TBT) had different effects in the traditional 2D platform compared to the organotypic platform; TBT had no effect on ER transactivation when MCF7s were grown as a 2D monoculture, in the presence or absence of testosterone. However, when grown in the organotypic co-culture model in the absence of testosterone, TBT acted as an ER antagonist (Fig. 1C). We concluded via qPCR that TBT induced the mRNA expression of CYP19A1, the gene encoding aromatase, in adipose stromal cells. We also found that TBT inhibited ER-driven gene induction in breast cancer cells. As breast cancer cells express ER but not aromatase, these findings highlight the value of incorporating co-culture into screening platforms.
While the above studies showed that our organotypic platform identified different hits than a 2D monoculture platform, we next wanted to investigate if the organotypic platform was more predictive of human responses than a 2D co-culture model. We chose to study the prototypic aromatase inhibitor anastrazole, which is commonly used to treat ER+ breast cancer. Clinical evidence suggests that obese women are less sensitive to aromatase inhibitors than lean women. To compare how stromal cells derived from lean and obese women influence response to the aromatase inhibitor, anastrazole, we incorporated human-derived adipose breast stroma in the MCF7-derived duct model; the adipose stromal cells were isolated from lean or obese women undergoing reduction mammoplasties. We found that the obese adipose stromal cells exhibited increased aromatase mRNA compared to the lean controls. In addition, MCF7-derived ducts co-cultured with obese stromal cells exhibited higher maximal aromatization-induced ER transactivation and reduced anastrazole sensitivity (Fig. 1D). In contrast, the 2D co-culture model revealed no differences between the lean and obese samples (Fig. 1E). These findings suggest that some chemical interactions and human sensitivities are missed when evaluated in 2D, and that platforms that include structure and/or matrix may be needed to better understand population level effects. A manuscript detailing these findings has been accepted for publication by FASEB.
A criticism of our organotypic breast model is that it utilizes MCF7 cells as the epithelial cell type, which is an immortalized breast cancer cell line. Towards increasing the physiological relevance of our organotypic system, we enriched for primary mammary epithelial cells derived from women undergoing normal reduction mammoplasties (referred to as HMECs) and evaluated their behavior and estrogen responsiveness when grown in the organotypic mammary duct model. To evaluate if the ductal structure induced differences in HMEC gene expression, HMECs were grown in 2D or in ductal structures for 48 hours then evaluated for the expression of ESR1, which encodes ER protein, as well as SPCA1 and SPCA2, which are two calcium pump genes associated with increased calcification found in DCIS. In both patients evaluated, expression of ESR1, SPCA1, and SPCA2 were increased when HMECs were grown as luminal structures. To evaluate estrogen responsiveness, HMEC-derived ducts were exposed to a vehicle control or the potent estrogen, 17B-estradiol then evaluated for the expression of ER-driven genes ESR1 and PGR. Estradiol decreased ESR1 mRNA (Fig. 1F) and increased PGR mRNA (Fig. 1G). To further validate that estradiol was inducing effects through ER, HMEC-derived ducts were exposed to a vehicle control, estradiol, or estradiol and 4-hydroxytamoxifen, which is the active form of the ER antagonist tamoxifen. After a 48-hour exposure, cultures were fixed then stained for ER protein and the proliferation marker, Ki67. Similar to normal human breast tissue, cells rarely co-expressed Ki67 and ER, and approximately 10-20% of epithelial cells expressed ER (representative picture shown in (Fig. 1H). While estrogen decreased ER protein in HMEC-derived ducts, co-treatment with tamoxifen and estrogen increased ER protein. The percentage of Ki67+ cells decreased with estrogen treatment, while co-treatment with tamoxifen had no effect. Altogether, these findings demonstrate that we can culture HMECs in the biomimetic duct model, and that in this model the HMECs are estrogen responsive and exhibit different gene expression profiles versus 2D.
Journal Articles on this Report : 5 Displayed | Download in RIS Format
| Other subproject views: | All 6 publications | 6 publications in selected types | All 6 journal articles |
|---|---|---|---|
| Other center views: | All 215 publications | 82 publications in selected types | All 81 journal articles |
| Type | Citation | ||
|---|---|---|---|
|
|
LIvingston M, Morgan M, Daly W, Murphy W, Johnson B, Virumbrales-Munoz M. Evaluation of PEG-Based Hydrogel Influence on Estrogen-Receptor-Driven Responses in MCF7 Breast Cancer Cells. ACS Publications 2019;5(11):6089-6098. |
R835737 (2018) R835737C003 (Final) |
Exit |
|
|
Morgan MM, Livingston MK, Warrick JW, Stanek EM, Alarid ET, Beebe DJ, Johnson BP. Mammary fibroblasts reduce apoptosis and speed estrogen-induced hyperplasia in an organotypic MCF7-derived duct model. Scientific Reports 2018;8(1):7139. |
R835737 (2017) R835737C003 (2017) R835737C003 (Final) |
Exit |
|
|
Morgan M, Arendt L, Alarid E, Beebe D, Johnson B. Mammary adipose stromal cells derived from obese women reduce sensitivity to the aromatase inhibitor anastrazole in an organotypic breast model. FASEB Journal 2019;33(7):8623-8633. |
R835737 (2018) R835737C003 (Final) |
Exit |
|
|
Stallcop LE, Alvarez-Garcia YR, Reyes-Ramos AM, Ramos-Cruz KP, Morgan MM, Shi Y, Li L, Beebe DJ, Domenech M, Warrick JW. Razor-printed sticker microdevices for cell-based applications. Lab on a Chip 2018;18(3):451-462. |
R835737 (2017) R835737C003 (2017) R835737C003 (Final) |
|
|
|
Ayuso JM, Gillette A, Lugo-Cintrón K, Acevedo-Acevedo S, Gomez I, Morgan M, Heaster T, Wisinski KB, Palecek SP, Skala MC, Beebe DJ. Organotypic microfluidic breast cancer model reveals starvation-induced spatial-temporal metabolic adaptations. EBioMedicine 2018;37:144-157. |
R835737C003 (Final) |
|
Progress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 Human Models for Analysis of Pathways (H MAPs) Center Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
6 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center
