Grantee Research Project Results
Final Report: Development of Molecular Biomarkers to Measure Environmentally Induced Immune Responses
EPA Grant Number: R832947Title: Development of Molecular Biomarkers to Measure Environmentally Induced Immune Responses
Investigators: Miller, Lisa A. , Gilliland, Frank D. , Abel, Kristina , Margolis, Helene , Gern, James , Joad, Jesse
Institution: University of California - Davis
EPA Project Officer: Callan, Richard
Project Period: June 15, 2006 through June 14, 2009 (Extended to December 14, 2011)
Project Amount: $712,423
RFA: Early Indicators of Environmentally Induced Disease (2004) RFA Text | Recipients Lists
Research Category: Human Health
Objective:
The primary objective of this project was to develop a panel of immune biomarkers that could be used to detect environmentally induced disease in humans, focusing on parameters of allergy and asthma. Phenotypic evaluation of immune cells is effectively done by flow cytometry; however, this approach often requires immediate processing and can be problematic with small sample quantities. We hypothesized that molecular biology techniques could accurately provide quantitative measurements of immune cell phenotype and function within small biological samples. To test this hypothesis, we designed a panel of oligonucleotide primers and probes for real-time quantitative RT-PCR analysis of relevant markers of allergy and asthma, then utilized cloned plasmid DNA standards to establish absolute quantitation of immune cell phenotype and associated cytokines. The significance of each immune biomarker in the context of environmentally induced airway disease was subsequently determined by characterization of peripheral blood and airway cells obtained from an infant rhesus monkey model of environmentally induced childhood asthma.
To address the question of relevant immune biomarkers for environmentally induced airway disease, we completed the following two research aims within the context of this project:
- Develop a high-throughput quantitative RT-PCR assay for leukocyte cluster of differentiation markers to measure key immune cell populations associated with allergy and asthma in small biological samples.
- Develop a high-throughput quantitative RT-PCR assay for key cytokine markers associated with allergy and asthma to measure immune cell function in small biological samples.
The equipment and reagents required to conduct this analysis are readily available and not limited to this campus. As such, the molecular reagents and methods characterized within this proposal can be directly implemented into a large population study of human environmental exposures. Importantly, values obtained by this method are absolute and can be compared among different individuals and at different time points within a population study of any magnitude. In addition, biological samples can be collected in a stabilizing reagent without substantial technical training and stored indefinitely before analysis is completed. This approach is particularly applicable for longitudinal population studies that require birth cohorts, due to the transient nature of immune system development during infancy and early childhood.
Summary/Accomplishments (Outputs/Outcomes):
METHODS
For the first specific aim, we proposed to develop a molecular method to measure numbers of T lymphocyte populations in small biological specimens. We initially focused our efforts on testing and optimizing RT-PCR primer/probe reagents that can detect both human and rhesus macaque targets, with the intent that this strategy could be immediately incorporated into an epidemiology study with human subjects. Purified T lymphocyte cell populations were subsequently generated from adult rhesus monkey peripheral blood and confirmed for purity. Finally, a standard curve was generated using optimized RT-PCR reagents and serial dilutions of purified T lymphocytes. Peripheral blood samples from a cohort of 12 adult rhesus monkeys were tested for correlation of T lymphocyte numbers with copy numbers of mRNA.
Animals
Animals from the California National Primate Research Center (CNPRC) were selected based on age (> 4 years of age), and evenly distributed by sex and country of origin.
Table I. Adult Rhesus Monkey Cohort for Peripheral Blood Analysis
| Animal Number | Country of Origin | Sex | Age (Months) |
| 1 | Full Chinese | F | 105 |
| 2 | Full Chinese | F | 119 |
| 3 | Full Chinese | M | 56 |
| 4 | Full Chinese | M | 57 |
| 5 | Chinese/Indian Hybrid | F | 107 |
| 6 | Chinese/Indian Hybrid | F | 69 |
| 7 | Chinese/Indian Hybrid | M | 72 |
| 8 | Chinese/Indian Hybrid | M | 59 |
| 9 | Full Indian | M | 108 |
| 10 | Full Indian | M | 84 |
| 11 | Full Indian | F | 95 |
| 12 | Full Indian | F | 83 |
Reagents
We obtained commercially available purified plasmids containing complete cDNA inserts for human CD3 epsilon, human CD4 p55, and human CD8 alpha chain genes. RT-PCR primer/probe reagents that target both human and rhesus macaque monkey gene sequences were tested. Cross-reactivity for human and rhesus was confirmed by amplification of human cDNA targets as well as monkey peripheral blood samples. Standard curves for CD3 epsilon, CD4 p55 and CD8 alpha chain mRNA copy number were generated by RT-PCR amplification of serially diluted plasmids.
Cell Isolation
We isolated CD3+, CD4+, and CD8+ rhesus macaque lymphocytes using a commercially available antibody-conjugated magnetic bead system. Purity of lymphocyte preparations was confirmed by flow cytometry; we were able to obtain 98-99% purity of CD3+, CD4+ and CD8+ cell populations. Using defined numbers of CD3+, CD4+, and CD8+ cells, we isolated RNA and generated cDNA for RT-PCR analysis. CD3 epsilon, CD4 p55, and CD8 alpha chain mRNA were amplified from defined numbers of cells.
RESULTS
Using this strategy, we used purified cell preparations from a pool of six randomly selected adult monkeys to generate a standard curve for cell numbers, such that quantitative estimates of T lymphocyte populations could be determined from unknown blood samples. We subsequently found that mRNA levels in purified cell preparations were significantly reduced due to the isolation procedures, such that the use of molecular values from a standard curve obtained by this approach were not comparable to whole blood preparations. To circumvent this technical problem, we focused our efforts on whole blood preparations to avoid artifact imposed by isolation procedures. Instead of using purified cell preparations, we were able to accurately quantify T lymphocyte subset numbers by incorporation of counting beads during flow cytometry.
As shown in Figure 1, values obtained from a cohort of 12 adult rhesus macaque monkeys showed good correlation of CD3 and CD8 mRNA copy numbers with cell numbers obtained via flow cytometry analysis. CD4 mRNA did not correlate as well with CD4 cell numbers by flow cytometry; it is possible that this was due to low level expression of CD4p55 on monocyte populations in peripheral blood.
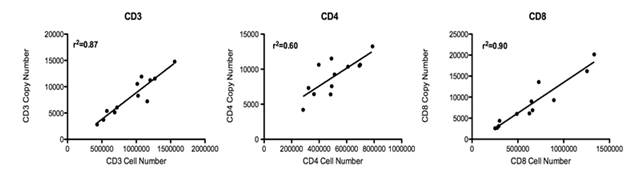
Figure 1. Correlation of T lumphocyte Cell Numbers With Corresponding mRNA Copy Numbers
METHODS
Reagents
For the second specific aim, we focused on the testing and optimization of RT-PCR reagents that detect both rhesus and human immune biomarkers associated with allergy and asthma. In our original application, we proposed to evaluate 23 different immune targets in conjunction with the establishment of standard curves using plasmids containing human cDNA inserts. At completion of this study, we have generated standard curves for 32 different RT-PCR reagents that can quantitatively measure both rhesus monkey and human mRNA targets for immune cells and cytokines (Table II).
| Target | Function |
|---|---|
| CD3 epsilon | Cell Type Specific |
| CD4 p55 | Cell Type Specific |
| CD8 alpha chain | Cell Type Specific |
| CCR3 | Cell Type Specific |
| Eosinophil Cationic Protein | Cell Type Specific |
| FoxP3 | Cell Type Specific |
| IL-4 | Adaptive Immunity |
| IL-5 | Adaptive Immunity |
| IL-17 | Adaptive Immunity |
| IL-2 | Adaptive Immunity |
| IL-13 | Adaptive Immunity |
| IFN gamma | Adaptive Immunity |
| IL-10 | Adaptive Immunity |
| CD25 | Adaptive Immunity |
| TGF beta 2 | Adaptive Immunity |
| CCL11 | Innate Immunity |
| CCL24 | Innate Immunity |
| CCL26 | Innate Immunity |
| CCL20 | Innate Immunity |
| IL-12p35 | Innate Immunity |
| IL-6 | Innate Immunity |
| IL-8 | Innate Immunity |
| IL-3 | Innate Immunity |
| IL-9 | Innate Immunity |
| TNF alpha | Innate Immunity |
| GMCSF | Innate Immunity |
| TLSP | Innate Immunity |
| TLR3 | Innate Immunity |
| TLR4 | Innate Immunity |
| TLR5 | Innate Immunity |
| GAPDH | Housekeeping Gene |
| Beta actin | Housekeeping Gene |
Animals
Peripheral blood and lavage samples were obtained from infant rhesus monkeys exposed to house dust mite (Dermatophagoides pteronyssinus) aerosol, ozone, or combined house dust mite aerosol and ozone exposure (n = 6 per group). In brief, newborn monkeys were sensitized to house dust mite by intradermal injection with purified Der p1 and Der p2. From 1 month to 6 months of age, animals were exposed to house dust mite aerosols and/or ozone at 0.5 ppm in a cyclical fashion.
RESULTS
To determine which immune markers would best identify biomarkers of allergy and asthma in peripheral blood, we completed a longitudinal analysis of 10 different mRNA targets in peripheral blood samples collected from infant rhesus monkeys exposed to house dust mite (D. pteronyssinus), ozone, or combined house dust mite and ozone. Targets were selected based on literature surveys of both rodent models as well as human studies, with the intent of initially screening for those cytokines that have been experimentally shown to promote allergy and/or asthma. To determine the effect of developmental time point and exposure on cytokine expression levels, we collected peripheral blood every 4 weeks for the first 6 months of life, starting at 2 weeks of age. Whole blood samples were stored in an RNA storage solution immediately after collection and RNA was collected at a later time point.
In Figure 2, longitudinal mRNA expression profiles are shown for IL-4, IL-12p35, IL-6, and IFN gamma in peripheral blood; these cytokines were selected based on their known essential roles in directing T helper 1, T helper 2, and T regulatory differentiation. Other targets evaluated included IL-17, IL-2, IL-10, and IL-13. An important finding from this study is that cytokines do not necessarily follow a linear trajectory with regards to expression over time during postnatal development. For example, IFN gamma expression in peripheral blood cells is increased with age; however, IL-12 had multiple peaks of expression during the evaluation period. It should also be noted that mRNA levels for different cytokines were variable. IFN gamma mRNA was readily detectable, but IL-12p35 mRNA was expressed at very low levels.
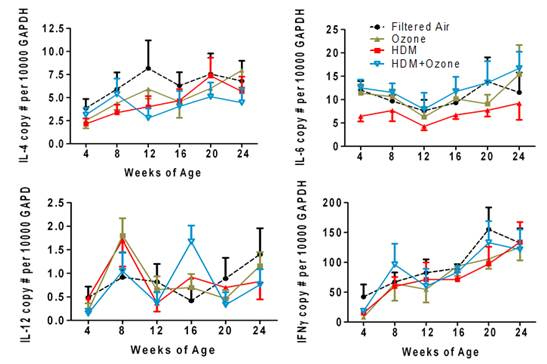
Figure 2. Longitudinal analysis of IL-4, IL-6, IL-12 and IFN gamma mRNA in infant comkey
peripheral blood: effect of house dist mite and/or ozone exposure during the first six months
of life.
In addition to surveying cytokine mRNA expression in a longitudinal fashion, we also quantified mRNA expression of CD3 epsilon and Foxp3, which are measures of the total T lymphocyte population as well as the T regulatory cell population. As shown in Figure 3, CD3 mRNA appears to follow a linear trajectory over time, whereas Foxp3 expression is modulated with either ozone or house dust mite exposure. An important question that was addressed in this study is whether peripheral blood responses correspond with immune events within the airways. As shown in Figure 4, the combination of house dust mite and ozone results in a striking increase in CD3 mRNA in lavage cells. Foxp3 mRNA expression is also increased in lavage cells from animals exposed to ozone or house dust mite and ozone combined. Although we do observe modest changes in response in peripheral blood due to experimental exposures, these findings would suggest that immunopathology within the lung is distinct from the circulation.
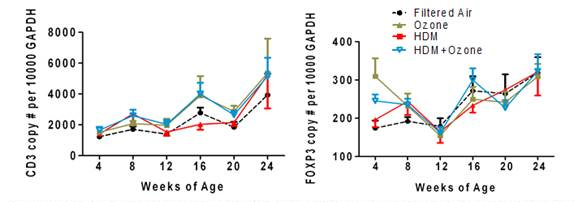
Figure 3. Longitudinal analysis of CD3 epsilon and Foxp3 in infanty monkey peripheral blood: effect
of house dust mites and/or ozone exposure.
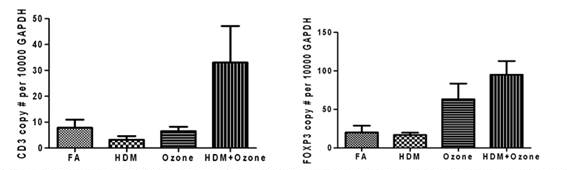
Figure 4. Expression of CD3 episilon and Foxp3 in infanty monkey bronchoalyeolar lavage cells:
effect of house dust mites and/or ozone expsoure
Conclusions:
- In the development of T lymphocyte specific markers, we found that mRNA copy numbers were not always indicative of cell number. Depending upon the mRNA target, expression levels may be variable with age and possibly gender. If utilized in a large population study, it will be important to conduct a preliminary study with combined RT-PCR and flow cytometry on multiple age groups (infant, juvenile, adult) to determine if expression levels are associated with chronological age.
- Longitudinal peripheral blood studies in very young animals have demonstrated that immune cell number and cytokine profile does not always follow a linear trajectory. This finding has important implications for peripheral blood studies in young children, as even small incremental differences in age may contribute to variability in data outcome and will not be reflective of exposure.
- Immunological profiles within the peripheral blood may not be reflective of mucosal sites such as the lung.
Journal Articles on this Report : 4 Displayed | Download in RIS Format
| Other project views: | All 6 publications | 4 publications in selected types | All 4 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Chou DL, Gerriets JE, Schelegle ES, Hyde DM, Miller LA. Increased CCL24/eotaxin-2 with postnatal ozone exposure in allergen-sensitized infant monkeys is not associated with recruitment of eosinophils to airway mucosa. Toxicology and Applied Pharmacology 2011;257(3):309-318. |
R832947 (2007) R832947 (2009) R832947 (Final) |
Exit Exit Exit |
|
|
Clay CC, Reader JR, Gerriets JE, Wang TT, Harrod KS, Miller LA. Enhanced Viral Replication and Modulated Innate Immune Responses in Infant Airway Epithelium following H1N1 Infection. Journal of Virology 2014;88(13):7412-7425. |
R832947 (Final) |
Exit Exit |
|
|
Clay CC, Maniar-Hew K, Gerriets JE, Wang TT, Postlethwait EM, Evans MJ, Fontaine JH, Miller LA. Early Life Ozone Exposure Results in Dysregulated Innate Immune Function and Altered microRNA Expression in Airway Epithelium. PLoS ONE 2014;9(3):e90401. |
R832947 (Final) |
Exit Exit |
|
|
Maniar-Hew K, Clay CC, Postlethwait EM, Evans MJ, Fontaine JH, Miller LA. Innate Immune Response to LPS in Airway Epithelium Is Dependent on Chronological Age and Antecedent Exposures. American Journal of Respiratory Cell and Molecular Biology 2013;49(5):710-720. |
R832947 (Final) |
Exit Exit |
Supplemental Keywords:
Lymphocytes, cytokines, allergy, asthma, house dust mite, biomarkers of exposure and disease, ozone, air pollution, peripheral blood, RFA, Health, Air, Scientific Discipline, PHYSICAL ASPECTS, HUMAN HEALTH, Susceptibility/Sensitive Population/Genetic Susceptibility, Health Risk Assessment, Physical Processes, Risk Assessments, particulate matter, genetic susceptability, Environmental Chemistry, Allergens/Asthma, Health Effects, Environmental Monitoring, sensitive populations, health risks, chemical characteristics, airway inflammation, asthma indices, asthma triggers, asthma, aerosol composition, airborne particulate matter, human exposure, ambient air monitoring, environmental risks, second hand smoke, atmospheric particles, atmospheric aerosol particles, particulates, exposure, atmospheric particulate matter, airborne pollutants, allergic response, ambient air quality, air pollution, human health risk, human susceptibility, air toxics, inhalation, airway diseaseProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
