Grantee Research Project Results
2008 Progress Report: Development of High-Throughput and Real-Time Methods for the Detection of Infective Enteric Viruses
EPA Grant Number: R833008Title: Development of High-Throughput and Real-Time Methods for the Detection of Infective Enteric Viruses
Investigators: Chen, Wilfred , Yates, Marylynn V. , Myung, Nosang V. , Mulchandani, Ashok
Current Investigators: Chen, Wilfred , Mulchandani, Ashok , Yates, Marylynn V. , Myung, Nosang V.
Institution: University of California - Riverside
EPA Project Officer: Page, Angela
Project Period: August 31, 2006 through August 30, 2009 (Extended to August 26, 2011)
Project Period Covered by this Report: August 31, 2007 through August 30,2008
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The main goal of this research is to improve on the current analytical methods for quantitative detection of infective enteric viruses, specifically the non-polio enteroviruses (NPEV), in drinking water. The overall objective of the research is to develop methods to provide high-throughput real-time detection and quantitation of infective enteric viruses from contaminated water. The specific objectives of this research are to:
1) Develop a new generation of molecular beacons (MBs) based on quantum dots as the fluorophore and gold nanoparticles as the quencher for improved sensitivity and multiplexing capability
2) Develop a real-time method to probe and quantify infective enteric viruses using TAT- or transferrin-modified nuclease-resistant molecular beacons (MBs) in infected cell lines without permeabilization
3) Develop a genetically engineered cell line to probe and quantify infective enteric viruses by generating a protease-sensitive FRET protein pair using an improved CFP-YFP pair
4) Evaluate the use of flow cytometry for high-throughput sample processing
5) Evaluate the above methods to rapidly detect and quantify the presence of infective NPEV in environmental water samples.
Progress Summary:
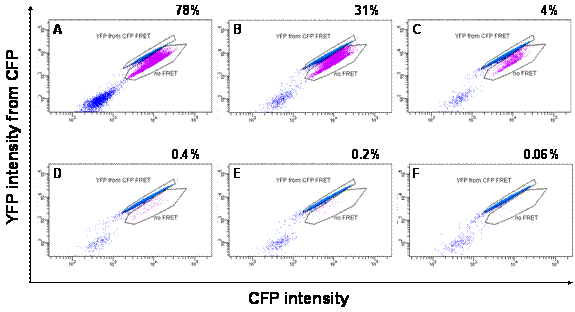
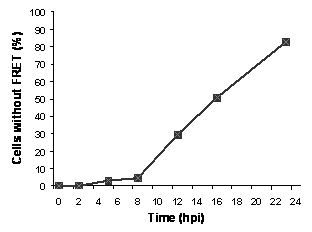

Fig. 3. Direct comparison of FACS and plaque assays. Confluent monolayer of BGMPV in 12-well plates were infected with PV1 (20 min absorption). For FACS assay, cells were harvested after 12 h as described, and then subjected to FACS. For plaque assay, infected cells were overlaid with 1% CMC in MEM (+ 2% FBS), incubated for 48 h, then fixed/stained with 1% crystal violet in 3.7% formaldehyde solution.
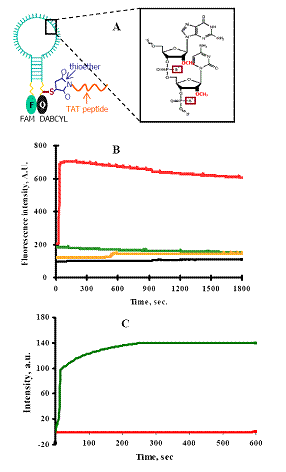
In this study, we describe the use of nuclease-resistant molecular beacons (MBs) for the real-time detection of coxsackievirus B6 (CVB6) replication in living Buffalo green monkey kidney (BGMK) cells via TAT peptide delivery. A nuclease-resistant MB containing 2'-O-methyl RNA bases with phosphorothioate internucleotide linkages was designed to specifically target an 18-bp 5' noncoding region of the viral genome. For intracellular delivery, a cell-penetrating TAT peptide was conjugated to the MB using a thiol-maleimide linkage (Fig. 4A). As expected, the modified MBs were highly resistant to nuclease cleavage by DNAase I (Fig. 4B). In contrast, an unmodified MB was susceptible to nuclease degradation, resulting in almost instantaneous increase in fluorescence. The dual modifications had no effect on the hybridization kinetics of the MB, as a rapid increase in fluorescence was observed in the presence of a complimentary target (Fig. 4C).
Fig. 4. (A) A schematic representation of the TAT-modified nuclease-resistant MB. (B) Nuclease sensitivity assays utilizing ribonuclease-free DNase I. The fluorescence of the nuclease-resistant MB is shown in yellow and the fluorescence of an un-modified MB is shown in red. The background fluorescent signals (shown in black and green) without DNase I addition are also shown. (C) Kinetics of hybridization of TAT-modified MBs CVB6 with (green) or without (orange) complementary oligos.
The intracellular delivery efficiency was tested by incubating 0.5, 1, or 2 μM of MB-target hybrids with a monolayer of
Fig. 5. Intracellular delivery of (A) MB CBV6-TAT/target hybrids or (B) MB without TAT modification or without targets. BGMK cells were incubated with 1 µM MB for 12 h, and images were captured using a fluorescent microscope. Scale bar, 20 µm.
Fig. 6. (A) Visualization of BGMK cells infected with 0, 1 or 105 PFU at 2 hpi (B) The correlation between the number of PFU and fluorescent cells at 2 hpi Error bars represent the standard deviation of three replicate experiments.
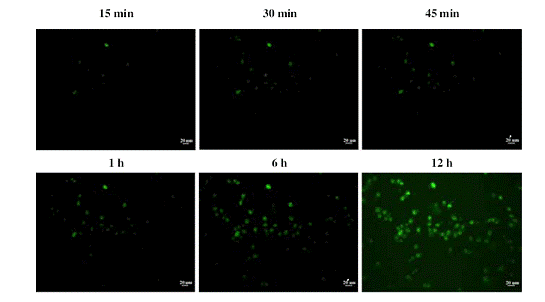
The ability to detect infected cells continuously should allow one to follow the spreading of infectious viruses on a real-time basis. To determine whether this was, indeed, possible, BGMK cells were infected at a very low infection dosage (MOI: 0.01 pfu/cell) and monitored continuously using a fluorescence microscope in a fixed area for 12 h. Fig. 7 shows the cell to cell progression of virus spreading at 6 representative time points. Several infected cells were observed at 15 min p.i., suggesting that the viruses entered the cells and started the uncoating process within 15 min. The number of fluorescent cells slowly increased with time suggesting continuous virus infection. By 6 h p.i., a further outward spread of fluorescent cells was observed, indicating the secondary spreading of infection from progeny virions to cells surrounding the initial infected cells. The number of fluorescent cells continued to increase with time and finally spread outward to the entire observation area by 12 h p.i. The majority of infected cells remained adherent and some fluorescence was visible outside the cells, indicating that the fluorescent hybrids with viral RNA entered the extracellular region as a result of the release of progeny virions during cell lysis.
Fig. 7. Real-time detection of viral spreading. BGMK cells were first incubated with 1 μM MB, infected with CVB6 at an M.O.I. of 0.01 pfu/cell, and monitored using a fluorescent microscope.
Future Activities:
Journal Articles on this Report : 3 Displayed | Download in RIS Format
| Other project views: | All 7 publications | 7 publications in selected types | All 7 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Hwang Y-C, Chu JJ-H, Yang PL, Chen W, Yate MV. Rapid identification of inhibitors that interfere with poliovirus replication using a cell-based assay. Antiviral Research 2008;77(3):232-236. |
R833008 (2008) R833008 (Final) |
Exit Exit |
|
|
Yeh H-Y, Hwang Y-C, Yates MV, Mulchandani A, Chen W. Detection of Hepatitis A virus using a combined cell culture-molecular beacon assay. Applied and Environmental Microbiology 2008;74(7):2239-2243. |
R833008 (2008) R833008 (Final) |
Exit Exit Exit |
|
|
Yeh H-Y, Yates MV, Mulchandani A, Chen W. Visualizing the dynamics of viral replication in living cells via TAt peptide delivery of nuclease-resistant molecular beacons. Proceedings of the National Academy of Sciences of the United States of America 2008;105(45):17522-17525. |
R833008 (2008) R833008 (Final) |
Exit Exit Exit |
Supplemental Keywords:
RFA, Scientific Discipline, PHYSICAL ASPECTS, INTERNATIONAL COOPERATION, Water, POLLUTANTS/TOXICS, Health Risk Assessment, Physical Processes, Microbiology, Microorganisms, Drinking Water, health effects, human health, measurement method, viruses, microbiological organisms, monitoring, pathogens, drinking water contaminants, enteric viruses, exposure, drinking water monitoring, human health effectsProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.