Grantee Research Project Results
2009 Progress Report: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
EPA Grant Number: R833010Title: On-Chip PCR, Nanoparticles, and Virulence/Marker Genes for Simultaneous Detection of 20 Waterborne Pathogens and Potential Indicator Organisms
Investigators: Hashsham, Syed
Current Investigators: Hashsham, Syed , Tiedje, James M. , Tarabara, Volodymyr
Institution: Michigan State University
EPA Project Officer: Page, Angela
Project Period: September 1, 2006 through August 31, 2009 (Extended to August 31, 2011)
Project Period Covered by this Report: September 1, 2008 through August 31,2009
Project Amount: $600,000
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
The objective of this research is to develop and validate a highly parallel, sensitive, specific, and quantitative microchip with on-chip PCR and isothermal amplification for the simultaneous detection of multiple waterborne pathogens. Establishing highly parallel and specific methods are essential to reduce the health risk from microbial pathogens present in source and drinking waters.
Progress Summary:
1. Work Status and Progress, Preliminary Data, Results, and Evaluations
The main objectives of this research are to: i) develop low-cost microfluidic chips, ii) validate the primers targeting waterborne pathogens for parallel, sensitive, specific, and quantitative detection in microfluidic chips, and iii) develop an efficient sample concentration scheme for fast and efficient recovery of P22 bacteriophage. The progress related to these objectives made during the period Sep 1, 2008 to Aug 31, 2009 are described below.
1.1 Development of Microfluidic Chips

1.2 Validation of On-chip DNA Amplification for Waterborne Pathogens
To further simplify the assay and lower the cost, we have chosen an isothermal DNA amplification reaction named loop-mediated isothermal reaction (LAMP). Features of LAMP include, moderate reaction temperature between 63-65 oC, excellent specificity due to the use of four to six specific primers, need for only single enzyme, and superior tolerance to substances that typically inhibit PCR. Another advantage of LAMP is the large amount of DNA generated during amplification. This permits direct visualization of amplification product based on turbidity or fluorescence, either at endpoint or in real-time. We have designed LAMP primers for the VMGs associated with major waterborne pathogens (Table 1). We have used SYTO dyes to monitor real-time LAMP by using a fluorescence imaging system. Due to the isothermal nature of LAMP, time-to-positivity (TTP) value was calculated by applying the signal to noise ratio cutoff greater than 10. With the application of SYTO dyes, we found that fluorescence LAMP in microchips is faster than in real-time PCR machine (Chromo4TM). Three parameters were studied for the simultaneous screening of VMGs in microfluidic chips, sensitivity, specificity, and genotyping potential.
Table 1: Selected VMGs for major waterborne pathogens and their function
| Pathogen | Gene target(s) | Description |
| Cryptosporidium parvum | GP60 | 60 kDa glycoprotein |
| hsp70 | 70 kDa heat shock protein | |
| Giardia duodenalis | beta-giardin | conserved protein |
| Legionella pneumophila | dotA | integral cytoplasmic membrane protein |
| lepB | effector protein | |
| Vibrio cholerae | ctxA | cholera toxin |
| tcpA | toxin-coregulated pilus protein | |
| toxR | two-component regulator | |
| Shigella flexneri | ipaH | invasion plasmid antigen H |
| Campylobacter jejuni | 0414 | putative oxido reductase subunit |
| cdtA | cytolethal distending toxin A | |
| Escherichia coli O157:H7 | eaeA | intimin |
| stx1 | Shiga toxin 1 | |
| stx2 | Shiga-toxin 2 | |
| Salmonella enterica | invA | invasion protein |
| phoB | phosphate regulon |
Detection and quantitative sensitivity for 2 of the novel assays inside of the amplification chips was tested. This was done to test if the high surface to volume ratio or the hydrophobic polymeric chip did not inhibit the reaction with low amounts of starting copies. Inhibition at low copy numbers could influence both the limit of detection and quantification. Results showed high correlation between the starting copies and time to positive amplification. Detection limit was 100 copies for C. jejuni gene cdtA and was 10 copies for the other assay targeting C. jejuni gene 0414. In all cases, the standard deviation was low indicating good reproducibility and potential for quantification.
Specificity of real-time LAMP assays was tested from blind samples containing targeted genomic DNA mixed with background DNA from environmental samples. Detection sensitivity was in the range of 10-100 genomic DNA copies for these VMGs. For a total of 60 real-time LAMP reactions, only 4 false positive calls were observed. These false positives were only observed for less than 10 targeted genomic DNA copies. However, signal increased later in the reaction (over 20 min) indicating primer-dimer formation. In fact, these LAMP primers were insensitive to less than 10 genomic DNA copies as tested on real time PCR machine (Chromo4TM).
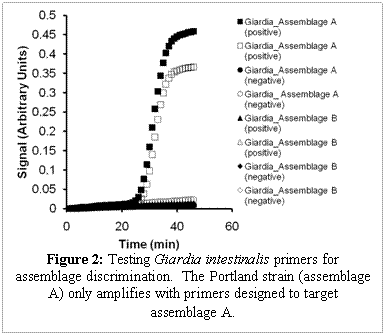
LAMP primers were designed to discriminate between alleles of a given gene (targeting single nucleotide polymorphisms) to genotype assemblages of Giardia. Primer sets (targeting the beta-giardin gene) to discriminate between assemblages of Giardia were designed with a mismatch on the 2nd, 5th, and 8th primer on the the 5’ end of the FIP, and the BIP primer has a mismatch on the 2nd and 5th base. An experiment with Giardia intestinalis (Portland-1 strain, assemblage A) mixed with primers targeting assemblage A and primers targeting assemblage B, only showed amplification with the assemblage A primer set (Figure 2). The success of this experiment will allow us to genotype Giardia.
1.3 Filtrations of P22 Bacteriophage with Calf serum Blocked Membrane at Various Pressures and Cross Flow Rates
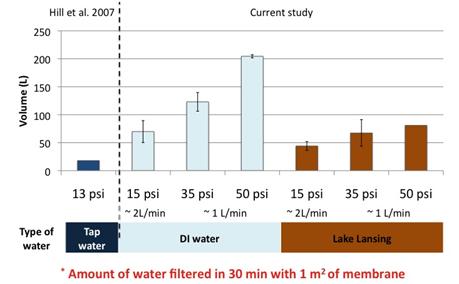
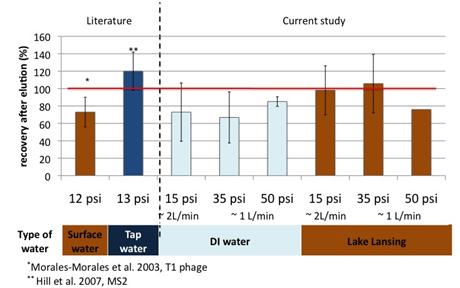
Result show that we were able to maintain recoveries observed in literature while increasing permeate flux (Figure 3). However poor reproducibility was observed. In addition the investigation of polyelectrolytes effect on plaque assay and qPCR showed that the presence of polyelectrolytes in samples could greatly affect plaque assay result; but their effect on qPCR results was limited and less than the one observed for calf serum. Because of these two factors we switched to qPCR analyses to compare PEM to calf serum blocked membrane.
Figure 3: Effect of the physiochemical factors and values on the filtration of P22 with calf serum blocked membrane in this study against the values reported in literature.
Future Activities:
- Validate the chips with a focus on increasing the well density of microchips.
- Optimize the filtration parameters like trans-membrane pressures and cross-flow rates and compare the performance for the concentration of P22 on calf-serum blocked and polyelectrolyte-coated membranes.
- Prepare publications/manuscripts focusing on: i) the correlation of potential indicators with antibiotic resistance genes; ii) the quantitative evaluation of bias during whole genome amplification; and iii) polyelectrolyte multilayer films as anti-adhesive membrane coatings. Pathogen concentration by ultrafiltration: effects of hydraulic conditions on virus rejection and recovery will be written in the next year. The manuscript related to the dye doped nanoparticle based detection of viability will be submitted to the journal Applied and Environmental Microbiology.
Journal Articles:
No journal articles submitted with this report: View all 16 publications for this projectSupplemental Keywords:
drinking water, genetics, innovative technology, bacteria, biology, measurement methods, human health, water sample concentration, Water, Scientific Discipline, RFA, Drinking Water, Environmental Engineering, Environmental Chemistry, Environmental Monitoring, biomarkers, drinking water contaminants, nanotechnology, polymerase chain reaction, analytical methods, monitoring, pathogens, biochip;, RFA, Scientific Discipline, Water, Environmental Chemistry, Drinking Water, Environmental Monitoring, monitoring, pathogens, polymerase chain reaction, analytical methods, drinking water contaminants, nanotechnology, biochip, drinking water monitoringProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.