Grantee Research Project Results
2010 Progress Report: Differentiating biologically relevant from irrelevant IgE binding to food antigens for improved risk assessment and diagnostic studies using a humanized rat basophil cell line (RBL 30/25)
EPA Grant Number: R834065Title: Differentiating biologically relevant from irrelevant IgE binding to food antigens for improved risk assessment and diagnostic studies using a humanized rat basophil cell line (RBL 30/25)
Investigators: Goodman, Richard E.
Institution: University of Nebraska at Lincoln
EPA Project Officer: Aja, Hayley
Project Period: May 1, 2009 through March 31, 2011 (Extended to April 30, 2012)
Project Period Covered by this Report: June 1, 2009 through May 31,2011
Project Amount: $372,340
RFA: Exploratory Investigations in Food Allergy (2007) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
The overall purpose of this study is to develop standardized methods to use a humanized rat basophil leukemia (RBL) cell line that has been transformed with the alpha-chain of the human FceRI receptor, as a test system to evaluate potential biological significance of IgE binding from an allergic subject on any test protein/target. Evidence from many allergy studies demonstrates that some subjects have extremely high levels of IgE in their sera that binds specifically to extracts of proteins or purified proteins that the individual can tolerate without experiencing an allergic reaction. Possible reasons for the lack of response center on a lack of effective cross-linking of FceRI receptors on mast cells and basophils through IgE either because only one binding site (epitope) is recognized by that person’s IgE, possible instability of the antigen during cooking or digestion (food proteins), or other unknown reasons. The safety assessment of a new genetically modified organism may include serum IgE tests to evaluate potential risks. If such tests are used, it is important to have a functionality test method, such as basophil histamine release, something short of in vivo challenges in human subjects, to confirm safety or risk.
Progress Summary:
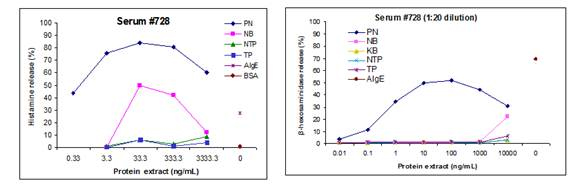
Figure 1. Comparison of the serum from peanut allergic donor 728 on activation using IgE stripped
human blood leukocytes by measuring histamine (left panel) and the humanized RBL cells 703/21
by measuring beta-hexosaminidase release (right panel). Extracts and proteins: PN, peanut extract;
NB, navy bean extract; NTP, non-transgenic pea extract; TP, transgenic pea extract; AIgE; pure
hIgE cross-lined with anti-IgE; BSA, bovine serum albumin.
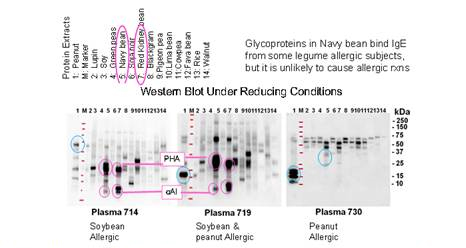
Figure 2. SDA-PAGE reducing Immunoblots with allergic sera, using hightly
specific mAb-IgE with chemiluminescent substrate. IgE binding to PHA and oIA
detection (serum samples 714 and 715, (circled in pink) was determined to be
carbohydrate-specific based on inhibition assays with protease digested PHA
compared to intact PHA. Protein-specific IgE binding shown in blue. This data
was presented at the EAACI and at the 5th International COwpea Confrence,
both in 2019 as part of two larger presentations on serum IgE binding and
basophil activation.
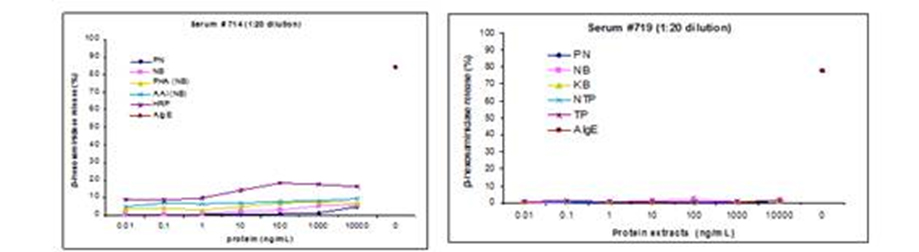
Figure 3. Beta-hexosaminidase release using hRBL [703/21] cells with diluted serum samples from subject 714 and 719, used in IgE western blots in Figure 2.
These two sera (714:719) were tested by commercial antigen-specific IgE tests [Immulite, DPC, reported in kU/L] against peanut [13kU/L;7.5kU/L] , CCD
(horseradish peroxidase = HRP [98 kU/L:98 kU/L] and one of the CCDs = MMXF [64 kU/L:48 kU/L]), soybean [22 kU/L:7.5 kU/L] and white bean (~navy
bean [kU/L:28 kU/L]). The angigens for hRBL assays include peanut extract (PN), navy bean extract (NB), kidney bean extract (KB), PHA purified from
navy bean, aAI from navy bean, horseradish peroxidase (HRP), non-transgenic pea (NTP), transgenic, alphy-amyalse-pea (TP), and cross-linking anti
IgE. This data was presented at the EAACI and at the 5th International Cowpea Confrence, both in 2010 as part of two larger presentations on serum
IgE binding and basophil activation.
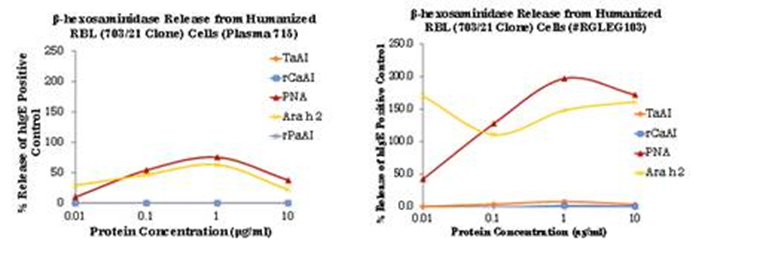
Figure 4, hRBL assay using plasma sample 715 and serum RGLEG 103. ZBoth released beta-hexosamindidase when
stimulated with Ara h 2 and also with PNA. No activity was measured for αAI. Examination of IgE immunoblots of PNA
demonstrated apparently higher IgE binding to Ara h 2 compared to PNA, although the Ara h 2 contamination of PNA
Future Activities:
Additional tests for the evaluation of other sera and plasma using other antigens will be performed during the next 5 months in the no-cost extension period. The impact of referencing activation with total release (detergent), pure IgE and anti-IgE and serum donor serum with anti-IgE will be explored. The utility of this test for evaluating heat processing effects on soybean flour products will be tested and possibly activation with maize lipid transfer protein will be added to support, or refute current results that IgE binding to CCD is unimportant, and that the impact of less potent allergens (e.g. PNA) can be over-estimated if contaminated with dominant allergens.
Journal Articles:
No journal articles submitted with this report: View all 7 publications for this projectSupplemental Keywords:
agricultural biotechnology, allergenicity, genetically modified (GM), basophil histamine release, cross-reactive carbohydrate determinants, IgE, FceRIProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
