Grantee Research Project Results
Final Report: ECOHAB: DNA-Based Molecular Diagnostics for Pfiesteria-Complex Organisms in Chesapeake Bay
EPA Grant Number: R826791Title: ECOHAB: DNA-Based Molecular Diagnostics for Pfiesteria-Complex Organisms in Chesapeake Bay
Investigators: Reece, Kimberly S. , Burreson, Eugene M. , Stokes, Nancy A.
Institution: Virginia Institute of Marine Science , College of William and Mary-VA
EPA Project Officer: Packard, Benjamin H
Project Period: October 1, 1998 through September 30, 2001
Project Amount: $279,202
RFA: Ecology and Oceanography of Harmful Algal Blooms (1998) RFA Text | Recipients Lists
Research Category: Aquatic Ecosystems , Harmful Algal Blooms , Water Quality , Water
Objective:
The overall objective of this research project was to develop complex and species-specific polymerase chain reaction (PCR) primers and DNA probes to facilitate identification of Pfiesteria-complex organisms in cultures, experimental aquaria, and environmental samples. DNA sequence data were obtained from clonal cultures of Pfiesteria spp. and Pfiesteria-like organisms (PLOs) found in the Chesapeake Bay and its tributaries for phylogenetic analysis and development of DNA-based molecular diagnostics.
Summary/Accomplishments (Outputs/Outcomes):
Sequence Analysis. DNA sequences of the 3' end (approx. 900 bp) of the small subunit ribosomal (SSU) RNA gene, the internal transcribed spacer (ITS) region and approximately 900 bp of the 5' end of the large subunit ribosomal (LSU) RNA gene were obtained from 8 cultures of Pfiesteria spp. and 15 other cultures of PLOs. This provided a comprehensive survey of DNA sequences for available clonal cultures. Sequences for each of the DNA regions were aligned and compared for designing DNA probes and PCR primers, and for phylogenetic analysis to examine the relationships among the Pfiesteria spp. and PLOs.
Until recently, the only reliable method for accurately identifying and distinguishing Pfiesteria (P) piscicida and other PLOs had been through the use of 3-D reconstructions of the thecal plate structures by scanning electron microscopy (SEM). This method is still considered to be the "gold" standard for accurate species identification. Sequence comparisons among the Pfiesteria spp. and PLO cultures, and to other dinoflagellates and protozoans allowed design of species-specific and genus (or group)-specific PCR primers and DNA probes. Clonal cultures of P. piscicida, Pfiesteria (P) shumwayae, Cryptoperidiniopsis spp., "Shepherd's crook" and "Lucy-like" PLOs, and three food source cultures of Rhodomonas spp. were first analyzed and identified by SEM and then obtained for DNA sequence analysis. DNA sequences were obtained from the food sources to ensure that dinoflagellate, rather than food source DNA clones, were selected for analysis. The SSU gene was highly conserved, while the ITS region (excluding the 5.8S portion), and portions of the LSU gene fragment, demonstrated considerable sequence variation. Comparison of DNA clone sequences obtained from clonal cultures demonstrated degenerated sites among replicate DNA clones amplified from clonal cultures in the ITS region and the LSU gene, and even a few degenerate sites in the SSU gene. These degeneracies demonstrated the importance of obtaining sequences from multiple clones before designing molecular diagnostics. A clonal culture isolated at VIMS, VIMS 1049 (CCMP 2089), is being used extensively for ongoing studies of Pfiesteria spp. pathogenicity. SSU sequence matched GenBank sequences for P. shumwayae. Overall, identifications made by comparing DNA sequences obtained from cultures, to those available in GenBank for Pfiesteria spp., were in agreement with identifications made by SEM. In addition, species of dinoflagellates were categorized based on SEM observations as well as on their DNA sequences.
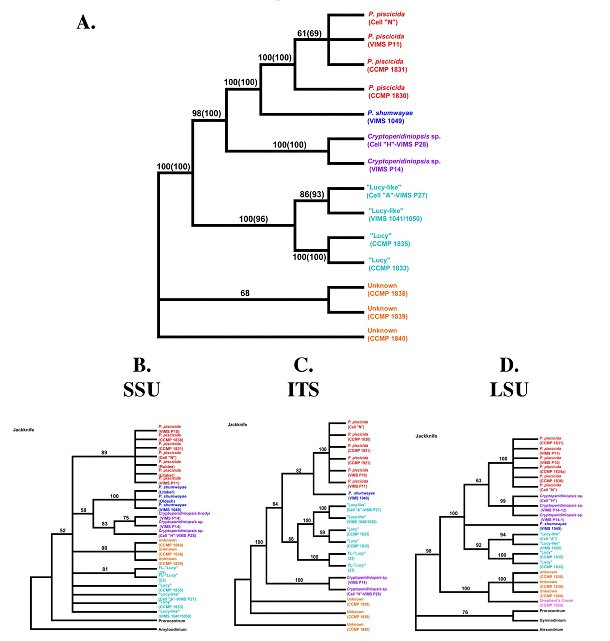
Phylogenetic Analysis. Phylogenetic analysis by maximum parsimony and maximum likelihood methods with all three data sets (SSU, ITS, and LSU), as well as with datasets combined, resulted in trees of identical topology (see Figure 1). In each analysis, cultures identified as P. piscicida grouped together. In addition, those cultures that were identified by SEM as Cryptoperidiniopsis spp. grouped together in the phylogenetic analysis, as did the cultures identified as "Lucy" or "Lucy-like."
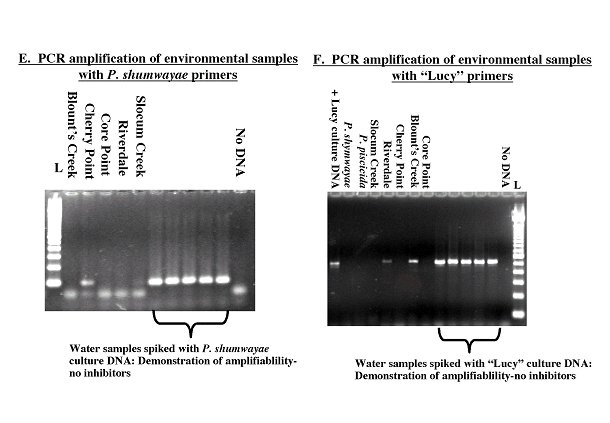
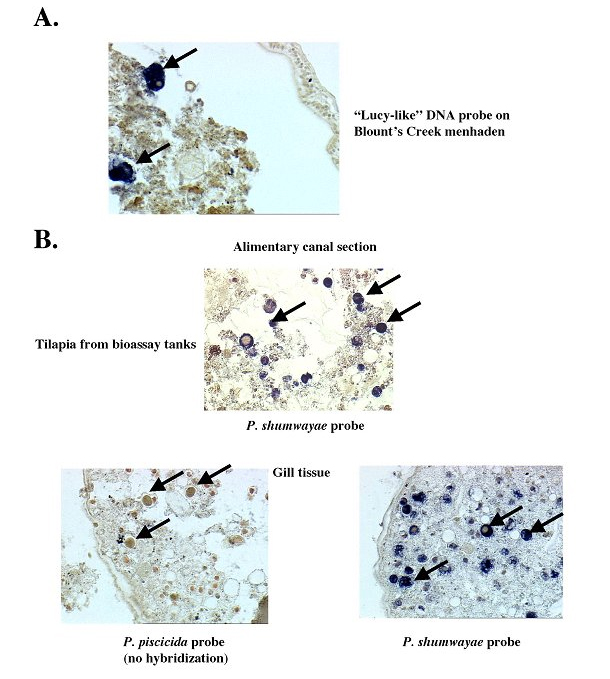
Molecular Diagnostics: PCR Primers and DNA Probes. Sequence alignments and comparisons were used to design several sets of PCR primers and DNA probes. Sequence variation within species, genera, and "groups" was examined to determine optimal regions for developing molecular diagnostics. Sequences unique to specific species, genera, or "groups" were identified for targeting by the molecular diagnostics. Primers were designed to specifically amplify P. piscicida, P. shumwayae, Cryptoperidiniopsis spp. or the so-called "Lucy" group of clonal culture DNAs. PCR primers were tested, and reaction conditions were initially optimized for specificity with clonal culture DNAs (see Figure 2 A-D). The primers were shown to specifically amplify the targeted DNA and not cross amplify the DNA of other PLO species. PCR primers also were tested on DNA isolated from toxic bioassay aquaria and from environmental samples, and were shown to specifically amplify the DNA of the targeted organisms (see Figure 2 E-F). In addition, specific DNA probes were designed for in situ hybridization. Water and fish samples from fish bioassay tanks and the environment were tested with the DNA probes (see Figure 3). Results of molecular diagnostic assays agreed with SEM identifications of clonal cultures and from bioassay tanks. Also, PCR assays confirmed the presence of PLO species detected by in situ hybridization. For example, the P. shumwayae probe hybridized to dinoflagellate cells found in the lateral line canal and alimentary tract of fish from an experimental tank with high cell counts of a PLO identified by PCR assay as P. shumwayae. DNA probe analysis indicated that dinoflagellates found in the alimentary tracts of menhaden collected from two sites in NC were "Lucy-like" cells (see Figure 3 A). PCR amplification of DNA from water samples collected at the same sites indicated the presence of "Lucy-like" DNA (see Figure 2 E-F). Amplicons obtained with P. piscicida, P. shumwayae, or "Lucy" group PCR primers from environmental samples were analyzed by DNA sequencing and digestion with several restriction endonucleases to confirm that the primers were specifically amplifying the targeted DNAs. The PCR assays have been shown to be specific for the target organisms and enabled rapid identification of PLOs from the environment. We have recently purchased a Cytospin™ centrifuge with funding from the Centers for Disease Control and Prevention (CDC) to facilitate the in situ hybridization assays with cultured cells and aquaria samples. We are continuing to analyze dinoflagellates in the alimentary canals, gills and lateral lines of fish and oysters, using in situ hybridization assays.
These molecular diagnostics currently are being used by Dr. Reece's laboratory for examining water samples collected in Virginia waters by the Department of Environmental Quality for a CDC-funded Cohort Study Project. In addition, the DNA probes are being used for identification of PLOs found in fish and oysters that filter water harboring dinoflagellates. In situ hybridizations to gut contents of menhaden and oysters collected from the field may provide a biological indication of PLO activity and abundance.
We have developed molecular detection techniques for P. piscicida, P. shumwayae, "Lucy-like" organisms, and Cryptoperidiniopsis spp. Primers for use in the PCR and DNA probes for in situ hybridizations were developed based on DNA sequences unique to each group of organisms. The primers were shown to specifically amplify the targeted DNA and did not cross amplify the DNA of other PLO species. In addition, the primers successfully amplified targeted organisms from environmental samples. DNA probes were developed and tested for specificity to P. piscicida, P. shumwayae, or "Lucy-like" PLOs.
Figure 1. Trees shown are he strict consensus of the most parsimonious trees generated by 10 random
addition sequence replecates using PAUP*4.0. Maximum likelihood analysis basded on the HKY model
(1985) using the combined dataset resulted in a tree of identical topology. Numbers on the branches outside
the parenteses give the parsimony jackknife values based on 100 replicates with 50% replacement.
Numbers in the parentheses represent the bootstrap support values for the maximum liklihood analysis.
Figure 2. PCR amplifications with specific primers. Primers were tested for specificity against DNA from
other PLOs (A-D) adn for the aability to amplify the targeted DNA from environmental samples (E-F). The
identity of the aplicons was confirmed by RFLP analysis and/or sequencing. L = 100 bp ladder.
Figure 3. In situ by hybridizations with DNA probes on field menhaden (A) and tilapia from bioassay tanks (B).
Journal Articles on this Report : 5 Displayed | Download in RIS Format
| Other project views: | All 20 publications | 5 publications in selected types | All 5 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Litaker RW, Vandersea MW, Kibler SR, Reece KS, Stokes NA, Steidinger KA, Millie DF, Bendis BJ, Pigg RJ, Tester PA. Identification of Pfiesteria piscicida (Dinophyceae) and Pfiesteria-like organisms using internal transcribed spacer-specific PCR assays. Journal of Phycology 2003;39(4):754-761. |
R826791 (Final) |
Exit Exit |
|
|
Lovko VJ, Vogelbein WK, Shields JD, Haas LW, Reece KS. A new larval fish bioassay for testing the pathogenicity of Pfiesteria spp. (Dinophyceae). Journal of Phycology 2003;39(3):600-609. |
R826791 (Final) R828225 (Final) |
Exit Exit Exit |
|
|
Steidinger K, Landsberg J, Richardson RW, Truby E, Blakesley B, Scott P, Tester P, Tengs T, Mason P, Morton S, Seaborn D, Litaker W, Reece K, Oldach D, Haas L, Vasta G. Classification and identification of Pfiesteria and Pfiesteria-like species. Environmental Health Perspectives 2001;109(Suppl 5):661-665. |
R826791 (Final) R827084 (2001) R827084 (Final) |
Exit |
|
|
Vogelbein WK, Lovko VJ, Shields JD, Reece KS, Mason PL, Haas LW, Walker CC. Pfiesteria shumwayae kills fish by micropredation not exotoxin secretion. Nature 2002;418(6901):967-970. |
R826791 (Final) R828225 (2000) R828225 (2001) R828225 (Final) |
Exit Exit |
|
|
Vogelbein WK, Shields JD, Haas LW, Reece KS, Zwerner DE. Skin ulcers in estuarine fishes:a comparative pathological evaluation of wild and laboratory-exposed fish. Environmental Health Perspectives 2001;109(Suppl 5):687-694. |
R826791 (Final) R828225 (2000) R828225 (2001) R828225 (Final) |
Exit Exit |
Supplemental Keywords:
genetics, Virginia, VA, North Carolina, NC., RFA, Ecosystem Protection/Environmental Exposure & Risk, Scientific Discipline, Geographic Area, Water, Waste, algal blooms, State, Oceanography, Contaminated Sediments, Environmental Microbiology, Ecology and Ecosystems, ECOHAB, phytoplankton, contaminated sediment, phylogenetic analyses, hybridization assays, Chesapeake Bay, polymerase chain reaction, Rhodomanas, dinoflagellates, DNA based molecular diagnostics, fish kills, pfiesteria, Maryland (MD)Progress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.