Grantee Research Project Results
Final Report: Vascular MAPs: Vascular and Neurovascular Tissue Models
EPA Grant Number: R835737C004Subproject: this is subproject number 004 , established and managed by the Center Director under grant R835737
(EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
Center: Human Models for Analysis of Pathways (H MAPs) Center
Center Director: Murphy, William L
Title: Vascular MAPs: Vascular and Neurovascular Tissue Models
Investigators: Sheibani, Nader
Institution: University of Wisconsin - Madison
EPA Project Officer: Aja, Hayley
Project Period: December 1, 2014 through November 30, 2018 (Extended to November 30, 2019)
RFA: Organotypic Culture Models for Predictive Toxicology Center (2013) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
- Develop human models of vascular and neurovascular organotypic cultures.
- Identify adverse outcome pathways associated with environmental toxins.
- Define the critical elements for an organotypic response, critical cell types, and tissue architecture.
- Leverage synthetic matrices and microscale systems to achieve robust tissue assembly and enable high throughput analysis.
- Determine broad applicability of the model for translational research in predictive toxicology.
Summary/Accomplishments (Outputs/Outcomes):
1) Develop human models of vascular and neurovascular organotypic cultures.
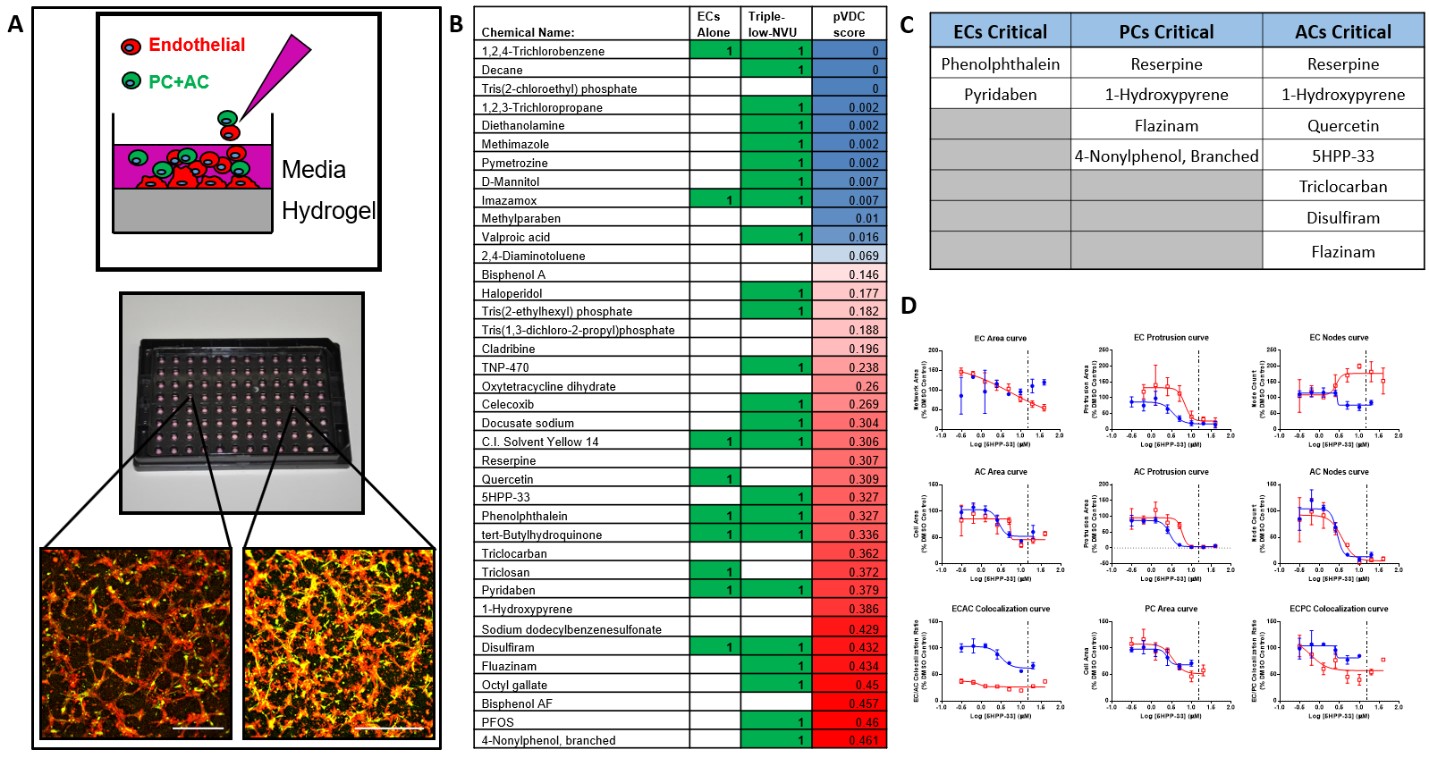
We have developed an in-vitro neurovascular unit (NVU) model by co-culturing human iPSC-derived endothelial cells (iPSC-EC) with human brain-derived pericytes (PC) and human iPSC-derived astrocytes (iPSC-AC) on synthetic poly(ethylene glycol) hydrogel surfaces (Figure 1A). We control the ratio of endothelial cells and perivascular support cells in culture, as well as the composition of the defined synthetic hydrogels to control vascular morphogenesis. Using this system, we have screened a panel of 38 putative vascular disrupting compounds, identified potential neurovascular disrupting compounds (Figure 1B), and prioritized hits based on the consistency of adverse effects on NVU morphology (Figure 1C). We then treated NVU models with the highest-priority chemicals detected in the screening experiments and generated dose response curves comparing chemical doses to the degree of vascular network disruption (Figure 1D). The dose response curves were generated using a NVU model where iPSC-EC were co-cultured with the lowest density of PC and iPSC-ACs that significantly increased the robustness of EC network formation compared to iPSC-EC-only controls. This perivascular cell density recapitulates situations in development or pathology where the NVU is particularly vulnerable to chemically-mediated disruption (Figure 1C). Additionally, we are currently able to quantify nine different morphological features of a NVU model a construct separate dose response curves for each metric (Figure 1D). Identical co-cultures were generated in culture environments based on synthetic hydrogels and Matrigel, and have compared dose response of the high-priority chemicals between the two environments.
Figure 1. A) Self-assembled vascular networks formed by iPSC-ECs (red), PCs, and iPSC-ACs (green) in 96-well plates. Scale bar: 0.5 mm B) Comparison of chemicals detected by NVU models consisting of iPSC-ECs only versus iPSC-ECs, PCs and iPSC-ACs in co-culture (Triple-Low-NVU). C) Highest-priority chemicals detected in a 38-compound chemical screen using the Triple-low NVU model constructed on hydrogel surfaces. D) Chemically-dependent changes in vascular network morphology are quantified through 9 metrics that are derived from fluorescent image analysis. Dose response to chemical treatments were compared between co-cultures generated in synthetic hydrogel (Blue) and Matrigel (Red) -based screening systems. Chemical: 5HPP-33.
2) Identify adverse outcome pathways associated with environmental toxins.
Our NVU model is capable of identifying differential effects of chemical exposure on the NVU based on the cell types present in co-culture. As a proof of concept, we previously exposed iPSC-EC, alone or in combination with PC and iPSC-AC, with high glucose representing diabetic hyperglycemia and associated osmotic stress, as well as VEGF inhibitor Sunitinib Malate. We observed differential responses to both inhibitors depending on cells present in the NVU model. We also demonstrated that a NVU model consisting of iPSC-EC, PC and iPSC-AC was able to detect the effects of chemical exposure with greater sensitivity than cultures consisting of iPSC-EC only (Figure 1B). To expand on the concept that individual cell types contribute to both tissue assembly and adverse outcomes to chemical insults, we developed image analysis methods to generate AC50 values based on chemically-mediated changes to the morphologies of individual cell types included in the NVU model. We were able to submit images and quantitative data on nine morphological metrics to the National Center for Computational Toxicology (NCCT) to aid in the generation of in-silico models of disrupted neurovascular development, and to relate known mechanisms of chemical action to specific changes in NVU morphology.
3) Define the critical elements for an organotypic response, critical cell types, and tissue architecture.
The NVU constructed on synthetic hydrogel surfaces is capable of identifying individual cell types that are affected by toxic insults. So far, we have determined chemicals, such as Pyridaben and Disulfiram, where iPSC-EC were critically affected by a chemical insult while PC and iPSC-AC were relatively unaffected, and Phenolphthalein, where iPSC-EC and iPSC-AC were more affected by chemical exposure compared to PC. This analysis promises to evaluate the effects of chemical exposure on all cell types included in the NVU model, inform investigators on which cell types are most critically affected by a chemical insult, and aid investigators on narrowing the pool of potential adverse outcome pathways that require further study. In addition, we are beginning to construct NVU models encapsulated in microfluidics channels to identify critical cell types and functional vascular changes involved in chemical exposure. Here, iPSC-ECs and PCs are encapsulated inside synthetic hydrogels to form perfusable vasculature, while iPSC-ACs will be added at a later time to stabilize the networks. The construction of these models enable the analysis of how NVU functions, such as fluid conduction and cytokine production, are affected by chemical insults.
4) Leverage synthetic matrices and microscale systems to achieve robust tissue assembly and enable high throughput analysis.
We are assembling iPSC-ECs, PCs and iPSC-ACs into three-dimensional (3D) NVU models in microfluidic channels. We have utilized microfluidic devices fabricated by the Cancer MAPS center to generate vascular networks that are immediately amenable to 3D perfusion tests, as well as tests to quantify molecule secretion and physical interactions between differential cell types in a 3D space. Within the device, we are able to generate networks co-cultured of iPSC-EC and PC within the hydrogels 72 hours after cell encapsulation. We are working to assess the effects of adding iPSC-ACs after network formation in order to stabilize networks for long periods of time. We aim to test known compounds, particularly those detected using NVU models cultured on hydrogel surfaces in order to relate morphological changes seen in rapid screening formats to changes in NVU function in a 3D space. We are also determining methods to evaluate functionality of NVU models without the need to perform confocal microscopy on hundreds of samples per experiment.
5) Determine broad applicability of the model for translational research in predictive toxicology.
We have developed a workflow to build co-cultures of iPSC-derived cells that are representative of the NVU, screen compounds from chemical libraries to detect chemically-mediated disruption of neurovasculature, prioritize positive calls for further study, and generate dose response curves for the highest priority chemicals identified in the screen. Our approach relates the results of rapid chemical screens performed on hydrogel surfaces to changes in NVU function that can only be visualized using perfusable vascular networks constructed in 3D spaces. Once the workflow is validated, its capabilities of detecting and evaluating the effects of putative neurovascular disrupting compounds are expected to be utilized in various preclinical models of pathology, such as those of ocular angiogenesis. Relevant pathological neovascularization includes ischemia, diabetes, and inflammation. Identification of various classes of chemicals with potential antiangiogenic activity using our screening platform may have clinical translational value for many diseases associated with neurovasculature, and can be evaluated in these models. Additionally, we developed the capability to utilize cells from multiple sources, including patient-specific stem cell sources, to explore the impact of patient-patient variability on the effects of chemical exposure.
Conclusions:
The results of our studies demonstrate our ability to synthesize well-defined synthetic cell culture substrates which efficiently support capillary morphogenesis by iPSC-EC. These substrates are adaptable for use in high-throughput screening systems and are amenable to various modifications that impact angiogenesis. Exploration of co-culture protocols and adaptation to microfluidic systems will accelerate the elucidation of regulatory mechanisms that maintain vascular integrity and function, their disruption via chemical exposure, and the downstream adverse effects on vascular and neural dysfunction. Discoveries here may lead to preventative measures and therapeutic pathways that lead to reduced impact of neurological diseases such as Alzheimer's disease, Parkinson's disease, ALS, and Multiple Sclerosis. The NVU model lends itself to modelling vasculature in different regions of the central nervous system, and modelling neurovasculature during varying stages of development.
Our future plans beyond the HMAPs funding period will focus on comparing the system constructed here to simpler screening methods, including single-cell-type cultures and Matrigel-based tubulogenesis assays. This will clarify the benefits of using the NVU models developed here as opposed to prior technology. The ability to modify cell ratios in co-culture will expand the capabilities of the 2D vascular screen to model the effects of chemical toxins on different regions of the central nervous system and different stages of neurovascular development. We will also work to clarify how specific morphological changes seen in fluorescence microscopy, such as cell spreading or protrusion formation in EC networks, relate to changes in specific cell signaling pathways. To begin this effort, we are performing cytotoxicity testing in conjunction with morphological datamining to determine if changes in NVU morphology occur due to a mechanistic change in cell activity, or due to cell death. We expect that the work relating changes in NVU morphology to adverse outcome pathways will be done in collaboration with the NCCT. We will also apply gene expression analysis techniques in collaboration with the bioinformatics arm of the hMAPS center to identify active pathways using the neurovascular model. We will also continue optimizing 3D encapsulation techniques to use fluid perfusion and co-culture assembly in 3D as readouts of neurovascular integrity and function. Observations here will be followed by RNA-seq analysis by the bioinformatics core of the hMAPs center, as well as further studies to relate specific changes in NVU morphology, such as iPSC-EC spreading or protrusion formation, to defined adverse outcome pathways.
Journal Articles:
No journal articles submitted with this report: View all 21 publications for this subprojectProgress and Final Reports:
Original AbstractMain Center Abstract and Reports:
R835737 Human Models for Analysis of Pathways (H MAPs) Center Subprojects under this Center: (EPA does not fund or establish subprojects; EPA awards and manages the overall grant for this center).
R835737C001 Liver MAPs
R835737C002 Brain MAPs
R835737C003 Cancer MAPs: A 3D Organotypic Microfluidic Culture System to
Identify Chemicals that Impact Progression and Development of Breast Cancer
R835737C004 Vascular MAPs: Vascular and Neurovascular Tissue Models
R835737C005 Pathway Analysis Core
The perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
17 journal articles for this subproject
Main Center: R835737
215 publications for this center
81 journal articles for this center