Grantee Research Project Results
2007 Progress Report: Cyanobacteria and Cyanotoxins in Water Supply Reservoirs – to Develop and Validate a Microarray to Test for Cyanobacteria and Cyanotoxin Genes in Drinking Water Reservoirs as an Aid to Risk Assessment and Management of Water Supplies
EPA Grant Number: R831627Title: Cyanobacteria and Cyanotoxins in Water Supply Reservoirs – to Develop and Validate a Microarray to Test for Cyanobacteria and Cyanotoxin Genes in Drinking Water Reservoirs as an Aid to Risk Assessment and Management of Water Supplies
Investigators: Rublee, Parke , Henrich, Vincent C. , Burkholder, Joann M.
Current Investigators: Rublee, Parke , Henrich, Vincent C. , Burkholder, Joann M. , Glasgow, Howard
Institution: University of North Carolina at Greensboro , North Carolina State University
EPA Project Officer: Aja, Hayley
Project Period: November 1, 2004 through October 31, 2007 (Extended to April 30, 2008)
Project Period Covered by this Report: November 1, 2006 through October 31,2007
Project Amount: $594,982
RFA: Microbial Risk in Drinking Water (2003) RFA Text | Recipients Lists
Research Category: Drinking Water , Human Health , Water
Objective:
To develop and validate a gene microarray for the detection of cyanobacteria and cyanotoxin genes in drinking water reservoirs. The microarray can be used to monitor drinking water supplies as an aid to risk assessment and management.
Progress Summary:
Sampling
Intensive sampling of targeted water supply reservoirs as originally planned was completed in 2006. Eleven North Carolina water supply reservoirs (Badin, Falls, High Point , Jordan, Kerr-Scott, Michie, Oak Hollow, Tillery, Townsend, Tuckertown, and Wheeler) were sampled during summer and winter months, and 18 additional lakes (Apex, Brandt, Belews, Benson, Coachman’s Trail, Graham, Greenfield, Harris, High Rock, Hyco, Johnson, Kerr, Lucas, Mackintosh, Reese, Rhodhiss, University, and Waccamaw) were sampled on at least once and in some cases up to six times. Physical/chemical characterization of the samples included: temperature, suspended solids, pH, TKN, NH4-N, NO2 + NO3, total P, SRP, and chorophyll a.
During 2008 we were presented with a unique opportunity that added additional samples to this study. The UNC General Administration targeted funding toward projects at member institutions of the state university system that had potential for significant translational research, that is, research that could rapidly be turned toward practical use. This funding was at a significant level ($292,100) for short duration (1 year). We competed successfully for this funding based primarily on the previous results and ongoing effort of the current EPA funding. We used this award to sample three of our water supply reservoirs sampling sites (Falls Lake, City Lake, and Oak Hollow Lake) intensively – samples every 2-3 weeks over a 1 year period (Dec. 2007-Dec 2008). Additionally, this grant provided funds for purchase of in situ automated nutrient samplers, allowing for near real-time assessment of water quality that could be used in directed sampling or to indicate significant change that might trigger sampling around an event (e.g. algal bloom). These samples were targeted for assessment of the broader microbial community. Because of the short duration of the UNC GA funding period, it reduced the intensity of our effort on the current grant, but it also provided a unique opportunity to use the same samples for assessment of the yearly pattern of cyanobacterial taxa in the current study, along with the automated water chemistry data. In turn these allowed some assessment of the feasibility of the automated approach as applied to monitoring of water supply reservoir systems. A further component of this work included consultation with water supply management personnel (a municipal water quality laboratory supervisor) and commercial interests (an emerging biotech water quality assessment entrepreneur) in order to enhance the process of moving science from the laboratory to the end-user in the field. A direct outcome of the UNC GA grant funding and interaction with commercial and municipal end-user communities was an evaluation that the glass chip microarray approach was not “end-user friendly” - it would require major technological breakthroughs in speed and ease of use as well as a reduction in cost before it could find wide acceptance in the end-user community. During this same period, a different array – PCR arrays – have undergone rapid advancement in use and reduction in cost. Thus, toward the end of the current reporting period, we explored the substitution of PCR arrays for hybridization-type arrays, since they allow for more rapid assessment at a lower cost, require much reduced technical expertise, and can provide “real-time” quantitative estimates of the targets. The only significant trade-off is that the PCR array cannot currently handle as many targets in a single plate than a glass slide microarray, although advances in equipment and chemistries are rapidly increases the number of targets that can be processed in both single and multiple runs. By the end of the grant period we became convinced that there was more potential for rapid acceptance of a PCR array approach than the traditional glass slide hybridization approach. Both methods rely on oligonucleotide primers directed toward the targets of interest, and both methods can use taxon specific fluorescent probes, although the PCR Array can often be used with only the primers and without the probes. Finally, the preliminary analyses of data from the UNC GA grant, completed with the collaboration of a postdoctoral researcher and purchase of analytical software (both under separate funding), has improved our ability to analyze the data in a meaningful way for further applications. These include Non-metric Mutlidimensional Scaling (NMDS) approaches as well as Bioindicator Analyses that provide insight into the relationships of target microbial taxa and environmental variables. In turn, these provide predictive tools that can be of benefit to water quality managers.
Clone libraries, Sequences
We completed sequencing of clone libraries from our targeted samples (selected samples from six reservoir samples with high chlorophyll a content indicative of high cyanobacterial populations). Sequences of 450 bp length from the beginning of the SSU rDNA were aligned and placed into operational taxonomic units (OTUs) based on a sequence similarity of ≥ 97.5%. Sequences from each OTU were compared to GenBank sequences (using Blastn) to identify OTUs. Design of primer pairs and probe sequences was initiated.
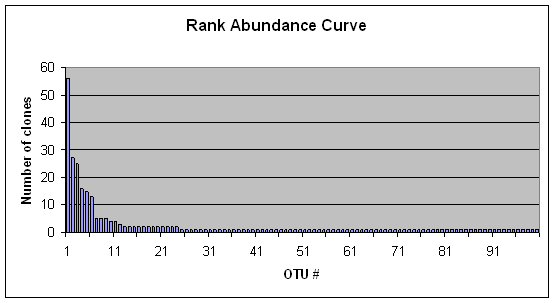
Overall, 278 clones were sequenced, and they aligned to 100 OTUs. The largest OTU grouping had 56 members, the second largest had 27 members, and 76 OTUs consisted of a single member (Fig. 1). Although most clones did not have 100% sequence identity with GenBank entries, most did have high similarity values such that at least some of the OTUs could be identified to a taxonomic category. For example, OTUs 1-4 mapped to Prochlorothrix spp., Cylindrospermopsis spp.; Cyanobium spp., and an uncultured cyanobacteria, respectively.
Figure 1. Rank abundance curve indicating number of clones in each operational taxonomic unit (OTU) found in this study.
An emerging outcome of this work has been the development of intellectual property and pursuit of patents by the University. One patent was issued based primarily on earlier work, but efforts and results from this research have been valuable in prosecution of the patent. Four additonal patent applications have been filed, each of which has been informed by the results of this research in combination with other efforts. We anticipate that final prosecution of these patent applications will still be years away, and it is likely that the final results from this grant will yield additional filings.
Finally, we have been communicating our results via presentations at local (including outreach to public schools), regional, and national levels. We have also had several publications due at least in part to this work. As noted later, several current patent applications have been enhanced by the results of this study to date.
Continuing Research Plan:
The major objectives for our final year are
- We will complete design of probe and primers sets to both known and novel cyanobacterial taxa from our samples. We will also continue to enhance our probe and primer collection from newly described taxa/sequences derived from recent literature.
- We will finalize a decision on whether to pursue real-time quantitative PCR arrays as the method of choice for a final array product. Continued consultation with “end-users” will be imperative for this decision.
- We will complete testing/validation of probe and primer sets.
- We will assay the field samples, including the seasonal samples derived from the UNC GA grant to characterize the patterns and geographical distribution of the cyanobacterial OTU targets generated by this study.
- We will prepare manuscripts for publication of results
- We will maintain collaboration with entrepreneurial and water quality laboratory manger end users in order to assure translation of our results for practical use.
As described in our proposal, development of the cyanobacterial array will provide: a research tool for aquatic scientists to identify factors that promote growth of different cyanobacterial species and their toxins; a prototype early warning system for the presence of potentially harmful cyanobacteria and their toxins which should be of use to water supply managers in order to reduce health risks; and the prototype will serve as the basis for development of a more “user-friendly” real-time (or near real-time) monitoring platform.
Journal Articles on this Report : 1 Displayed | Download in RIS Format
| Other project views: | All 33 publications | 6 publications in selected types | All 2 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Marshall MM, Amos RN, Henrich VC, Rublee PA. Developing SSU rDNA metagenomic profiles of aquatic microbial communities for environmental assessments. Ecological Indicators 2008;8(5):442-453. |
R831627 (2007) R831627 (Final) |
Exit Exit |
Supplemental Keywords:
RFA, Scientific Discipline, INTERNATIONAL COOPERATION, Water, Health Risk Assessment, Environmental Chemistry, Drinking Water, Environmental Engineering, Environmental Monitoring, microbial contamination, gene microarray assay, early warning, microbial risk assessment, monitoring, cyanobacteria, drinking water system, drinking water contaminants, aquatic organisms, other - risk assessmentRelevant Websites:
Real time physical and chemical data from some of our study sites can be found at the Center for Applied Aquatic Ecology website at NCSU: http://www.ncsu.edu/wq/ ExitProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.