Grantee Research Project Results
Final Report: Development of a larval fish neurobehavior adverse outcome pathway to predict effects of contaminants at the ecosystem level and across multiple ecologically relevant taxa
EPA Grant Number: R835798Title: Development of a larval fish neurobehavior adverse outcome pathway to predict effects of contaminants at the ecosystem level and across multiple ecologically relevant taxa
Investigators: Murphy, Cheryl A. , Garcia-Reyero, Natàlia , Carvan, Michael , Jones, Michael
Institution: Michigan State University , University of Wisconsin - Milwaukee , Mississippi State University
EPA Project Officer: Spatz, Kyle
Project Period: June 1, 2015 through May 31, 2018 (Extended to May 31, 2021)
Project Amount: $800,000
RFA: Systems-Based Research for Evaluating Ecological Impacts of Manufactured Chemicals (2014) RFA Text | Recipients Lists
Research Category: Chemical Safety for Sustainability
Objective:
Abnormal behavior after exposure to contaminants is a very good indicator and measure of sublethal effects. Behavior is a powerful bioassay because it integrates the internal physiological state of an animal with the animal’s response to external stimuli at the same time. However, embedding behavior into an AOP is challenging because it is difficult to interpret subtle changes in behavior in terms of population outcomes. We used directed laboratory studies that focus on behavior relevant to ecological processes such as foraging and predator avoidance, and used the data in an individual based model (IBM) calibrated for particular species and ecosystems. This project was focused on making use of valuable behavioral assays to create powerful AOPs that link the effects of manufactured chemicals at the molecular initiating level all the way to the population.
In this project, we placed the population outcomes of an AOP into the context of community structure to accommodate multiple stressors to the ecosystem (food web change as a result of climate change, invasive species or habitat loss). We also address the issue of error propagation as information is passed up through levels of biological organization and how this error influences estimation of risk – this is an issue that needs to be explored if the AOP framework is to be used as a viable risk assessment platform. This project was focused on two chemicals and three species of fish, but our ultimate goal was to develop methods that accelerate contaminant testing so that it can be simultaneously high throughput, it can predict the effects of thousands of contaminants on a wide array of taxa and it can extrapolates effects to individuals and populations in a cost efficient manner with acceptable levels of uncertainty
Main Objective: The overall objective of this project is to advance the adverse outcome pathway framework to predict effects of contaminants with different modes of action on the neurobehavior of larval fish from three different species and to determine what Adverse Outcome Pathways (AOPs) are common between species
Objective 1: Identify genes predictive of neurobehavior toxicity in response to exposure to two different chemicals with different molecular initiating events and modes of action in order to identify neurobehavior AOPs using a reverse engineering approach on zebrafish
Objective 2: Determine the effects of PCB126 and MeHg on gene expression and behavior of the larval stage of two species of ecological relevance (fathead minnow and yellow perch).
Objective 3: Incorporate behavioral effects and transcriptomics data from fathead minnow and yellow perch into an individual-based model (IBM) to predict changes in growth and survival to complete the neurobehavior AOP suitable for ecological risk assessment for MeHg and PCB126.
Objective 4: Define and compare neurobehavioral AOPs between species and contaminants to determine their similarities and to elucidate what kind of information is lost or gained by using a typical laboratory model to inform on environmentally relevant species at the population level.
Summary/Accomplishments (Outputs/Outcomes):
Objective 1
Identify genes predictive of neurobehavior toxicity in response to exposure to two different chemicals with different molecular initiating events and modes of action in order to identify neurobehavior AOPs using a reverse engineering approach on zebrafish
In the zebrafish we found significant differentially expressed genes (Table 1) and altered gene pathways. The behavior data has been collected from the videos, and we are currently running Bayesian models to determine significant behavioral effects that correlate to gene expression. The behavior and gene expression are being analyzed in the same way the yellow perch and killifish were completed.
Table 1. Total number of significantly differentially expressed genes (alpha = 0.05) found in the brains of zebrafish Danio rerio in this study.
| Treatment Comparison | Number of Differentially Expressed Genes |
| MeHg-Control vs MeHg-Low | 3 |
| MeHg-Low vs MeHg-High | 1349 |
| MeHg-Control vs MeHg-High | 1455 |
| PCB126-Control vs PCB126-Low | 162 |
| PCB126-Low vs PCB126-High | 1418 |
| PCB126-Control vs PCB126-High | 218 |
Objective 2
Determine the effects of PCB126 and MeHg on gene expression and behavior of the larval stage of two species of ecological relevance (fathead minnow and yellow perch). We added killifish to this Objective.
-
Yellow perch demonstrated subtle behavioral alterations when exposed to environmentallyrelevant concentrations of PCB 126 and MeHg
-
Using Hidden Markov Chain models, we expanded the ability to quantify swimming alterations in larvae due to neurotoxicant exposure
-
More behavior endpoints were altered after MeHg exposure than PCB126 exposure.
-
The new HMM analytical techniques found more altered swimming characteristics than the traditional methods for both chemicals.
-
Many behavior endpoints did not exhibit a threshold response but a non-linear response which may be an indication of a compensatory mechanism that fish are known to have (e.g. neurogenesis)
-
HMM analytical techniques may assist with high-throughput analyses
Sequenced the yellow perch genome:
Yellow perch (Perca flavescens) are a native North American fish that is important commercially and ecologically. Yellow perch are prized as a recreational species and as a high-value food fish. In the United States, its native range is concentrated in the Midwestern U.S., particularly in the Great Lakes region. Habitat loss and alteration have resulted in declining populations and reduced commercial fisheries, which has spurred interest in commercial aquaculture production. However, there are significant production bottlenecks that continue to limit capacity for commercial production. As for production, key issues are slow growth to market size and the occurrence of sexually-dimorphic growth, wherein females grow faster and larger than males. To address this, recent efforts have shown promise using genetic improvement and production of all-female progeny to achieve faster growth rates, limited size variation and improved immune performance. Understanding sex-specific differences in growth and immunity, as it relates to genes, is of high interest because of potential for their use as markers to further optimize commercial production of this species. To address this need, we have developed a highquality reference genome sequence for yellow perch using a combination of short- and long-read technology coupled with Dovetail HiC scaffolding to produce chromosome scale sequences. The majority of the assembly is in 24 scaffolds, representing the 24 pairs of yellow perch chromosomes. Based upon ~100X sequencing coverage, we estimated the size of the yellow perch genome to be ~1.0 Gb. We have utilized internal transcriptome assemblies, and public resources, to perform gene annotation as a means to understand regions of biological significance related to sex-specific differences in growth and immune function.
Sequencing Methods
Standard Pacific Biosciences large insert library preparation was performed. DNA was fragmented to approximately 20kb by needle shearing. Fragmented DNA was enzymatically repaired and ligated to a PacBio adapter to form the SMRTbell Template. Templates larger than 6kb were BluePippin (Sage Science) size selected. Templates were annealed to sequencing primer, bound to polymerase (P6), and then bound to PacBio Mag - Beads and SMRTcell sequenced using C4 chemistry on a Pacific Biosciences RSII sequencer.
Bioinformatics Methods
Assembly of the yellow perch genome began with sequencing output from 64 SMRTcells generated by a Pacific Biosciences RS II instrument, whose gross total genomic yield surpassed 92.1 Gbp, with a nested median insert read length of 8.66 kilobases. Genome assembly was initially performed with release version 1.5 of the Canu whole genome assembler, a modern derivative of the Celera Assembler. The input gkpStore of Canu reported an estimated 59.2x coverage of the yellow perch genome, with the following metrics generated for the resulting contig assembly: 5,374 contigs spanning a 962 Mbp draft reference sequence, with a contig N50 of 993 Kbp. This genome was then re-assembled using the latest major release of Canu, version 1.6, which boasts an improved contig consensus-calling algorithm; the output assembly contains 9,074 contigs spanning a 1.04 Gbp draft reference sequence, with a contig N50 of 922 Kbp. Given a theoretical size of 1.2 Gbp for the yellow perch genome, the v1.6 reference increased contig-coverage from 80.1% to 86.7%, with only a minor drop in the N50 value; we therefore opted to employ the latter assembly downstream. Following genome polishing achieved through use of Bowtie [5], Pilon [6], and a short-read (e.g. Illumina) library, annotation of the yellow perch draft reference was performed through use of the MAKER pipeline , resulting in the identification of 31,306 candidate genes. Sequencing and assembly of the yellow perch genome was performed by the Great Lakes Genomics Center (GLGC), while genome polishing and annotation was performed by the Institute for Genomics, Biocomputing, and Biotechnology (IGBB).
Gene Expression and Behavior:
In the yellow perch larvae brains we found differentially expressed genes (Table 2) and altered pathways. We also found correlations between gene expression and behavior.
Table 2. Total number of significantly differentially expressed genes (α = 0.05) found in the brains of yellow perch Perca flavescens in this study.
| Treatment Comparison | Number of Differentially Expressed Genes |
| MeHg-Control vs MeHg-Low | 7 |
| MeHg-Low vs MeHg-High | 8 |
| MeHg-Control vs MeHg-High | 40 |
| PCB126-Control vs PCB126-Low | 24 |
| PCB126-Low vs PCB126-High | 9 |
| PCB126-Control vs PCB126-High | 17 |
To completely understand biological risk assessment of pollutants, direct connections between the exposure event, molecular, cellular, and organismal key events are needed. To understand these connections, we examined gene expression, behavior, and simulated population effects from the same cohort of embryonic exposed fish larvae to neurotoxicants. We exposed Atlantic killifish Fundulus heteroclitus to sublethal-environmentally relevant levels of mercury (Hg) and PCB126. At 16-24 days post fertilization, we collected brain gene expression and numerous behavior endpoints to assess impacts on individuals and projected populations. The reference killifish from Scorton Creek, MA were exposed to Hg and exhibited changes in many gene pathways covering brain metabolism and development that coincided with similar changes in feeding and swimming characteristics. After PCB126 exposure, these fish had many more altered gene expression pathways that pertained to another group of brain metabolic processes and were associated with swimming stamina behaviors. Both Hg and PCB126 exposed Scorton Creek larvae had altered ubiquitin gene pathways that coincided with increased prey capture attempts. Toxic adapted killifish from New Bedford Harbor, MA exhibited another suite of altered brain metabolic and developmental gene pathways that coincided with multiple changed swimming state characteristics. Ultimately, these changes resulted in decreases in predicted survival for both chemicals but the predicted survivor’s growth only decreased after exposure to PCB126.
In the fathead minnow larvae brains we found differentially expressed genes (Table 3) and altered pathways. We still plan to conduct behavior analyses and link to gene expression and population outcomes.
Table 3. Total number of significantly differentially expressed genes (α = 0.05) found in the brains of fathead minnow Pimephales promelas in this study.
| Treatment Comparison | Number of Differentially Expressed Genes |
| MeHg-Control vs MeHg-Low | 1 |
| MeHg-Low vs MeHg-High | 6 |
| MeHg-Control vs MeHg-High | 3713 |
| PCB126-Control vs PCB126-Low | 0 |
| PCB126-Low vs PCB126-High | 608 |
| PCB126-Control vs PCB126-High | 1572 |
Objective 3
Incorporate behavioral effects and transcriptomics data from fathead minnow and yellow perch into an individual-based model (IBM) to predict changes in growth and survival to complete the neurobehavior AOP suitable for ecological risk assessment for MeHg and PCB126. We used our generic larval fish model to incorporate behavioral responses into EC50s and varied the predator composition. We used our generic larval fish model, calibrated for yellow perch, to incorporate laboratory collected data into realistic environmental scenarios. We performed an extensive sensitivity analysis on our generic larval IBM parameterized for a realistic yellow perch population to explore impacts of MeHg and PCB 126 on population that accounts for uncertainty.
Objective 4
Define and compare neurobehavioral AOPs between species and contaminants to determine their similarities and to elucidate what kind of information is lost or gained by using a typical laboratory model to inform on environmentally relevant species at the population level.
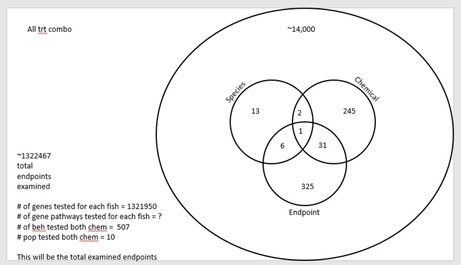
This section is still in development, but we have two papers planned. The first paper is to complete a meta analyses, that includes our data and depicts a dose response of behavioral impacts on larval fish exposed to MeHg and PCBs across species. We have conducted the literature review and standardized data. We are just developing a draft manuscript that will include data synthesized in Table 4. The second paper will compare all behavioral, gene expression and population endpoints across species, and chemical treatments. We are still finalizing some data, so this can’t be completed just yet, but we envision a summary that is idealized by Figure 1.
Table 4. Comparison of dosing of PCB 126 and MeHg on yellow perch, zebrafish, killifish and fathead minnow, with an estimate of relative lethality.
| Chemical Treatment | Level of Treatment, Dose | Potential Treatment shift using tissue conc | Concentration in Solvent | Dry Weight Tissue Concentration (n=3) | We Weight Tissue Concentration (n=3) | Units for Tissue Concentration | Realtive Lethality Measure (LD50 ppb dw; LC50 ppb) | ||||
| (ppm) | (uM) | mean | SD | mean | SD | ||||||
| Yellow Perch | |||||||||||
| MeHgCl Treatments | |||||||||||
|
| Control | Control | 0 | 0 | 5.26 | 1.16 | 0.39 | 0.08 | ppb |
| |
|
| Middle | Middle | 0.00021 | 0.001 | 40.35 | 5.76 | 3.34 | 0.64 | ppb |
| |
|
| High | High | 0.02156 | 0.1 | 5251.49 | 1187.30 | 420.62 | 88.02 | ppb |
| |
| PCB126 Treatments | LD50 | 1000c | |||||||||
|
| Control | Control | 0 | NA | DND | NA | ND | NA | ppb |
| |
|
| Middle | Middle | 0.01 | NA | DND | NA | ND | NA | ppb |
| |
|
| High | High | 1 | NA | 109.51 | NA | 6.367 | 5.333 | ppb |
| |
| Zebrafish | |||||||||||
| MeHgCl Treatments | LC50 | 103.6f | |||||||||
|
| Control | Control | 0 | 0 | YTBD |
| YTBD |
| ppb |
| |
|
| Middle |
| 2.15625 | 0.25 | 0.25 |
| YTBD |
| ppb |
| |
|
| High | Middle | 215.625 | 0.1 | 25.1 |
| YTBD |
| ppb |
| |
| PCB126 Treatments | LD50 | 100c | |||||||||
|
| Control | Control | 0 | NA | DND | NA | ND | NA | ppb |
| |
|
| Middle |
| 0.0001 | NA | DND | NA | ND | NA | ppb |
| |
|
| High | Middle | 0.01 | NA | DND | NA | ND | NA | ppb |
| |
| Killifish | |||||||||||
| MeHgCl Treatments | LC50 | 72.7e | |||||||||
|
| SCO - Control daily dose | Control | 0.3 | 1.391 | 9.8007 | 2.48644 | 8.2 | 2.2689 | ppb |
| |
|
| SCO - Hg daily dose | Middle | 3.6 | 16.70 | 35.09223 | 17.05802 | 28.13 | 14.7057 | ppb |
| |
| PCB126 Treatments | LD50 | 10d | |||||||||
|
| SCO - PCB126 40 ng/Lb | High | 0.00004 | NA | 19a |
| 1.11 | NA | ppb |
| |
|
| SCO - PCB126 400 ng/Lb |
| 0.0004 | NA | 189a |
| 111 | NA | ppb |
| |
|
| NBH - controlb |
| 0 |
| DND |
| DND |
| NA |
| |
|
| NBH - PCB126 40 ng/Lb | High | 0.00004 | NA | 19a |
| 1.11 | NA | ppb |
| |
|
| NBH - PCB126 400 ng/Lb |
| 0.0004 | NA | 189a |
| 111 | NA | ppb |
| |
| Fathead Minnow | |||||||||||
| MeHgCl Treatments | LC50 | 120g | |||||||||
|
| Control |
| 0 | 0 | YTBD |
| YTBD |
| ppb |
| |
|
| Middle |
| 2.15625 | 0.001 | YTBD |
| YTBD |
| ppb |
| |
|
| High |
| 215.625 | 0.1 | YTBD |
| YTBD |
| ppb |
| |
| PCB126 Treatments | LD50 | 20c | |||||||||
|
| Control |
| 0 | NA | YTBD |
| YTBD |
|
|
| |
|
| Middle |
| 0.0001 | NA | YTBD |
| YTBD |
|
|
| |
|
| High |
| 0.01 | NA | YTBD |
| YTBD |
|
|
| |
a Estimate using previous experiments (Nacci et al 1999)
b Also exposed to ~300 ng tHg/g dw/day through salmon-based diet
c Estimate based on relative TCDD potentcy (Elonen et al. 1998, Toomey et al. 2001, Spitsbergen et al. 1988)
d D. Nacci unpublished data on fish from a relatively uncontaminated site
e Sharp and Neff 1982
f Selderslaghs et al. 2012
g Valenti et al. 2006
1 Estimate using a killifish embryo percent moisture of 94.186
Figure 1. Idealized way of looking for similar genes, pathways and behaviors between chemical, endpoints and species (still in development so this is not final).
Finally we will also use the developed IBM is to assess the practice of using model species as a proxy for the impacts of toxicants on ecologically relevant species. To show this potentially future use of the model, we used the calibrated yellow perch model but substituted the multipliers developed for killifish. The additional scenarios we ran included using the perch model with killifish multipliers for the Scorton population for the baseline, mercury and PCB scenarios. We then ran each simulation 20 times to account for stochasticity. Finally, we compared model results for the perch control with perch and killifish multipliers, the perch low mercury with killifish and perch multipliers, and for the perch high PCB with perch and killifish multipliers.
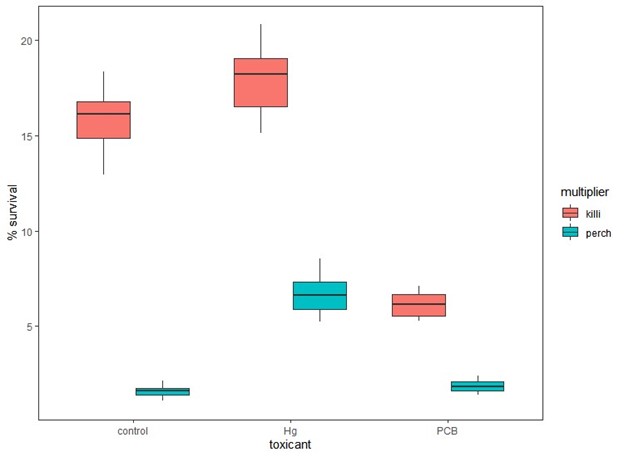
Figure 2. Percent survival of simulated yellow perch cohorts using multipliers derived from perch (perch) or killifish (killi) for control, low dose mercury (Hg) and high dose PCB exposures.
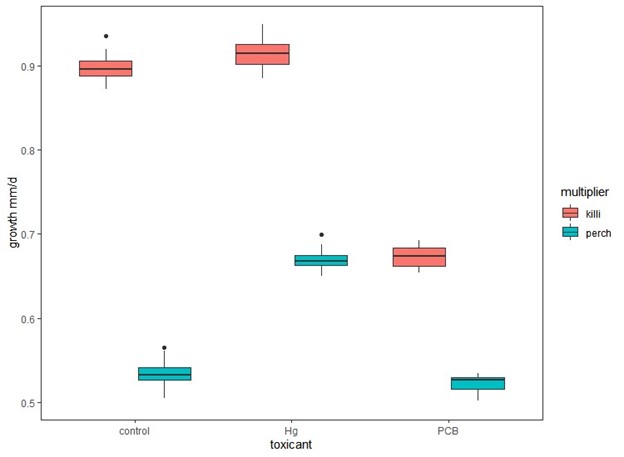
Figure 3. Growth (mm/d) of simulated yellow perch cohorts using multipliers derived from perch (perch) or killifish (killi) for control, low dose mercury (Hg) and high dose PCB exposures.
Using multipliers derived from killifish experiments rather than perch for all exposures resulted in significantly higher estimates of cohort survival and growth (Fig. 2 & 3) when compared to those derived from perch. In addition, the observed patterns would change when using killifish versus perch multipliers. When using perch multipliers, survival and growth estimates are similar between control and high PCB exposures and significantly higher for low mercury exposures (Fig. 2 & 3). However, the cohort survival and mean growth rates are greater for control and low mercury than PCB survival estimates when using the killifish-derived multipliers (Fig. 2 & 3).
This exercise shows that it may not be reasonable to use information derived for one species to estimate the impacts of a toxicant on another species, although we will explore this further once our data collection is complete.
Conclusions:
Understanding the ecologically relevant sublethal impacts on fish populations from human pollution involves interpreting complex biological information collected from suborganismal and organismal levels of biological organization. The Adverse Outcome Pathway framework (AOP) is proposed as a way to organize and predict impacts on suborganismal processes, ecologically relevant species from model species and different chemicals. AOPs could potentially expand the application of current studies that would ultimately decrease the amount of laboratory testing performed in toxicology. AOP assumptions were examined in this study; specifically, whether AOPs could predict adverse effects just from perturbation of biologically processes, if AOPs could extrapolate to other species from laboratory species, and to simplify toxicity endpoints by using measurable correlates in key events. Our study collected endpoints at multiple biological levels on three fish species after expose to two neurotoxicants and determined altered brain gene expression, fish behavior, and modeled YOY cohort survival and growth. In conducting this study, we discovered that what is needed is critical attention to chemical exposures regimes, animal husbandry, sequenced genomes, and analytical techniques that combine a diverse suite of biological endpoints. Preliminary results suggest little common responses for both chemicals and all fish species. However, there may be consistency with some gene expression and behavior endpoints within species and chemicals. Results from this study suggest limitations to predicting sublethal impacts across the species and chemicals in this study using the AOP framework, but overcoming identified limitations and further development in critical areas may help.
References:
Selderslaghs, I. W. T., R. Blust, and H. E. Witters. 2012. Feasibility study of the zebrafish assay as an alternative method to screen for developmental toxicity and embryotoxicity using a training set of 27 compounds. Reproductive Toxicology 33(2):142–154.
Sharp, J. R., and J. M. Neff. 1982. The toxicity of mercuric chloride and methylmercuric chloride to Fundulus heteroclitus embryos in relation to exposure conditions. Environmental Biology of Fishes 7(3):277–284.
Spitsbergen, J. M., J. M. Kleeman, and R. E. Peterson. 1988. 2,3,7,8-Tetrachlorodibenzo- p -dioxin toxicity in yellow perch (Perca flavescens). Journal of Toxicology and Environmental Health 23(3):359–383.
Toomey, B. H., S. Bello, M. E. Hahn, S. Cantrell, P. Wright, D. E. Tillit, and R. T. Di Giulio. 2001. 2,3,7,8-Tetrachlorodibenzodibenzo-p-dioxin induces apoptotic cell death and cytochrome P4501A expression in developing Fundulus heteroclitus embryos. Aquatic Toxicology 53:127–138.
Valenti, T. W., D. S. Cherry, R. J. Neves, B. A. Locke, and J. J. Schmerfeld. 2006. Case Study: Sensitivity of Mussel Glochidia and Regulatory Test Organisms to Mercury and a Reference Toxicant. Pages 351–367 in J. L. Farris and J. H. Hassel, editors. Freshwater Bivalve Ecotoxicology. SETAC, Pensacola, FL.
Journal Articles on this Report : 2 Displayed | Download in RIS Format
| Other project views: | All 23 publications | 8 publications in selected types | All 6 journal articles |
|---|
| Type | Citation | ||
|---|---|---|---|
|
|
Albers J, Ivan L, Clark B, Nacci D, Klinger R, Thrash A, Steibel J, Vinas N, Carvan M, Murphy C. Impacts on Atlantic Killifish from Neurotoxicants: Genes, Behavior, and Population-Relevant Outcomes. ENVIRONMENTAL SCIENCE & TECHNOLOGY 2024; |
R835798 (Final) R835872 (Final) |
Exit |
|
|
Ivan L, Jones M, Albers J, Carvan M, Garcia-Reyero N, Nacci D, Clark B, Klinger R, Murphy C. How Model Organisms and Model Uncertainty Impact Our Understanding of the Risk of Sublethal Impacts of Toxicants to Survival and Growth of Ecologically Relevant Species. ENVIRONMENTAL TOXICOLOGY AND CHEMISTRY 2024; |
R835798 (Final) |
Exit |
Supplemental Keywords:
transcriptomics, larval fish, fathead minnow, yellow perch, zebrafish, neurobehavior, MeHg, PCBs, Adverse outcome pathways, individual-based models, ecological risk assessment, uncertainty, riskProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.
Project Research Results
- 2019 Progress Report
- 2018 Progress Report
- 2017 Progress Report
- 2016 Progress Report
- 2015 Progress Report
- Original Abstract
6 journal articles for this project