Grantee Research Project Results
2007 Progress Report: Simultaneous Concentration and Real-time Detection of Multiple Classes of Microbial Pathogens from Drinking Water
EPA Grant Number: R833006Title: Simultaneous Concentration and Real-time Detection of Multiple Classes of Microbial Pathogens from Drinking Water
Investigators: Sobsey, Mark D. , Simmons, Otto D.
Institution: University of North Carolina at Chapel Hill
EPA Project Officer: Aja, Hayley
Project Period: October 18, 2006 through October 17, 2009
Project Period Covered by this Report: October 18, 2006 through October 17,2007
Project Amount: $599,999
RFA: Development and Evaluation of Innovative Approaches for the Quantitative Assessment of Pathogens in Drinking Water (2005) RFA Text | Recipients Lists
Research Category: Drinking Water , Water
Objective:
Improved methods are needed for rapid recovery and molecular detection of viruses and other pathogens in water. The goal of this project is to develop a method and protocol with documented performance characteristics to rapidly and efficiently recover, extract and detect DNA and RNA from enteric viruses in drinking water samples. Original specific objectives are summarized as:
- refine and validate new and improved, rapid ultrafiltration methods to concentrate viruses and other pathogens in water; compare to the existing virus concentration methods;
- evaluate new electropositive filters to rapidly and efficiently concentrate enteric viruses by adsorption to and elution from water and compare to existing virus concentration methods;
- evaluate rapid PEG precipitation and large volume nucleic acid extraction as post-concentration sample preparation techniques to further concentrate viruses and remove inhibitors for real-time, molecular detection of viral nucleic acids;
- optimize direct detection of viral RNA/DNA by real-time molecular methods;
- develop complete protocols of the methods and provide them to a select number of other water virology laboratories to conduct a collaborative (round-robin) tests characterizes their performance;
- apply the new performance-validated method and the standard EPA method to selected field samples of raw water to document that the new methods give superior performance.
The decision was made to systematically test a new, commercially available electropositive filter instead of developing a new one. This decision was based on the effectiveness of the new commercial filter and difficulties in QA/QC and in designing and bringing to market our own new electropositive filter. Re-circulating hollow-fiber ultrafiltration was evaluated for primary virus concentration from water.
Overall, the project is on schedule, recovery methods for viruses and other microbes have been improved and the effective performance of rapid methods for molecular detection of adenoviruses has been characterized. QA/QC efforts and activities have been successful, and data quality is satisfactory.
Progress Summary:
Primary Microbial Concentration from Water
During the first research year, a commercially available charge-modified filter for adsorption-elution concentration of viruses and other microbes from large volume water samples was evaluated. Two systems using hollow-fiber ultrafiltration: one using single-pass filtration and one using recirculation were tested with two types of filters for recovery of viruses as well as for bacteria and bacterial spores.
Virus Recovery by Adsorption-Elution to Nanoceram Charge-modified Filters. In a series of trials, adenovirus adsorption from water ranged from >92% to 99.9%, but recoveries were low, ranging from 0.01-3.60% due to inefficient elution, regardless of elution medium. Norovirus recoveries were better, with >99.9% adsorption and recoveries of 3-39% using 3% beef extract elution medium.
Microbial concentration by re-circulating hollow-fiber ultrafiltration (HFUF). Two brands of HFUFs, the Fresenius F80A and the Hemocor HPH, were tested for recovery of two strains of E. coli bacteria (one type strain and an environmental isolate), two coliphages (one single-stranded RNA and the other double-stranded DNA), and bacterial spores from dechlorinated tapwater. For the Fresenius F80A, mean recoveries were 36 to 122% with mean processing times of 49 minutes. For the Hemocor HPH, mean recoveries were 63 to 208% with mean processing times of 42 minutes. Because the two HFUFs performed similarly they may be used interchangeably. In similar trials comparing these two HFUFs for recoveries from source water samples, mean recovery ranged from 12 to 85% for the Fresenius F80A and from 57 to 144% for the Hemocor HPH. Average processing times were 49 and 35 minutes, for the Fresenius F80A and Hemocor HFUFs, respectively. The Hemocor HPH gave significantly higher recovery efficiencies in source waters than did the Fresenius F80A. In trials to test the effects of HFUF pretreatment on method recovery efficiency as well as sample processing times, mean recovery efficiency ranged from 36 to 122% for the HFUFs without pretreatment and from 39 to 91% with pretreatment , which was not a significant difference. Pretreated HFUFs gave equivalent results to those that are not pretreated but took longer for sample processing because of the added pre-treatment steps. Therefore, pretreatment is not recommended based on our research.
To reduce sample processing times, the use of modified filter endcaps was evaluated to increase sample flux through the HFUF system. For source water, average recoveries ranged from 12 to 85% with the standard method and from 32 to 153% with the system with modified ends. For de-chlorinated drinking water, average recoveries ranged from 36% to 122% for the standard method and from 50% to 87% with the modified ends. Microbe recoveries were not significantly different from de-chlorinated water samples but they were higher using the modified ends for source waters. The modified system reduced sample processing times from 0:49 to 0:27 minutes and from 0:42 to 0:26 minutes in source and de-chlorinated drinking water, respectively. Modified filter endcaps improved the method, so will be used in future work.
The optimized HFUF method was used to detect relatively low spike concentrations of the following microbial indicators and pathogens: E. coli O157:H7, Salmonella, Aeromonas, echovirus-12, adenovirus-41, Cryptosporidium, and Giardia at concentrations of 20-2835 organisms per liter. Average recoveries of indicators ranged from 51 to 495% for source waters and from 45 to 67% for de-chlorinated drinking waters. Average recoveries for pathogens ranged from 9 to 85% for source waters and from 7 to 117% for de-chlorinated drinking waters. These recoveries are satisfactory and more data at lower input levels will be collected in future studies. The schedule for primary recovery of microbes with filters is on track.
Secondary Concentration
Progress was also made for secondary concentration of adenovirus and noroviruses using polyethylene glycol (PEG) precipitation. For adenovirus 41 and Norovirus GII.4 and two concentrations of PEG (6% and 9%) and two concentrations of salt (NaCl, 0.1 M and 0.3 M), higher virus recoveries were achieved with the higher NaCl concentration of 0.3M and the higher PEG concentration of 9%. Virus recoveries ranged from 12.4% for adenovirus 41 with 6% PEG, 0.1 M NaCl to 115.0% with 9% PEG, 0.3 M NaCl. At optimum conditions recovery of Adenovirus at 115% was higher than Norovirus at 69%. PEG concentration trials were also conducted with mock filter eluents from de-chlorinated drinking water. Recoveries were 43 % for adenovirus 41 and 59% for Norovirus GII.4. Recoveries by PRG precipitation or satisfactory and this step in recovery will be used in future studies of the improved method.
Improved Molecular Detection of Adenoviruses
Adenovirus real time quantitative PCR (RTQ PCR) to assay Adenoviruses 2, 5, 8, 40 and 41
Improved molecular detection focused on real-time PCR for adenoviruses. Previously, real time quantitative PCR with primers and probe designed from the conserved hexon region of adenovirus genome could be used to quantify lower numbered, non-enteric adenoviruses (Ads). These primers and probes and amplification conditions are summarized in the following table.
Table 1. Primer, Probe and Reaction Conditions for Real Time or Conventional RT-PCR of Adenovirus
| Primers | Sequence | Total time: 90 mins RT for 30 min Denaturation: 15 min Total cycles: 45 Per Cycle: 95°: 5s; 55°: 5s; 72°: 10s |
| JHKXF | GGACGCCTCGGAGTACCTGA | |
| JHKXR | ACIGTGGGGTTTCTGAACTTGTT | |
| PROBE (JHKXP) | CTGGTGCAGTTCGCCCGTGCCA |
From: Ko, G., N. Jothikumar, et al. (2005). "Rapid detection of infectious adenoviruses by mRNA real-time RT-PCR." Journal of Virological Methods 127: 148-153
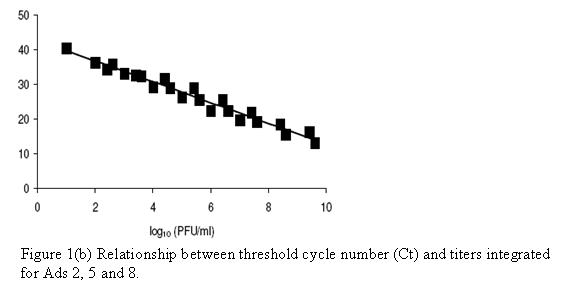
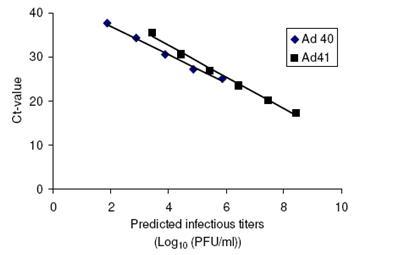
By concurrently measuring adenovirus titers by RTQ PCR and infectivity, the relationship between threshold cycle number (Ct-value) and adenovirus infectivity as PFU/ml was determined. For the quantification of Adenovirus 40, and 41, two assumptions were made. First, there might be a constant ratio of non-infectious virus particles to infectious virus particles within the general, lower numbered Ads, and second, real time quantitative PCR with primers and probe in the hexon region of Ads would give similar detection sensitivity as infectivity assays. Based on these assumptions, all data points for adenovirus 2, 5 and 8 titers and their corresponding Ct values were integrated. On the basis of regression analysis, Ad 40 and 41 titers for specific Ct values of 10-fold serially diluted samples also were estimated. These results are summarized in the Figure below. They show that there are predictable values between adenovirus concentration and Ct values for RTQ PCR, allowing for accurate estimations of virus concentrations based on this molecular assay procedure.
Figure 1(a). Quantification of Ad 2, 5, and 8 by real time RT-PCR
Figure 1(c) Predicted relationship between Ct value and Ad 40&41 titers
Future Activities:
Methods to simultaneously recover and detect viruses, bacteria and protozoan parasites in water using hollow fiber ultrafilters, PEG precipitation and (RT-)PCR will be further improved and evaluated for performance on experimentally contaminated samples of water. Protocols will be developed for the use of other labs and the developed methods will be collaboratively tested.
Journal Articles:
No journal articles submitted with this report: View all 4 publications for this projectSupplemental Keywords:
Viruses, Water, Filter, Concentration, Molecular Detection,, RFA, Scientific Discipline, Water, Environmental Chemistry, Drinking Water, Environmental Engineering, microbial contamination, ultrafiltration, contaminant candidate list, contaminant removal, pathogens, drinking water contaminants, drinking water treatment, drinking water monitoring, CCL, detectionProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.