Grantee Research Project Results
Final Report: A Field-Deployable Droplet Digital PCR System for the Rapid Detection of Waterborne Bacterial Pathogens
EPA Grant Number: SU839880Title: A Field-Deployable Droplet Digital PCR System for the Rapid Detection of Waterborne Bacterial Pathogens
Investigators: Li, Yiyan , Ferguson, James , Duran, Jesse , Theobald, Nic , Kukula, Kaitlyn
Institution: Fort Lewis College
EPA Project Officer: Callan, Richard
Phase: I
Project Period: October 1, 2019 through September 30, 2020 (Extended to September 30, 2021)
Project Amount: $24,824
RFA: P3 Awards: A National Student Design Competition Focusing on People, Prosperity and the Planet (2019) RFA Text | Recipients Lists
Research Category: P3 Awards , P3 Challenge Area - Safe and Sustainable Water Resources
Objective:
Conventional waterborne bacteria monitoring is carried out by traditional culturing methods which suffer from long turnaround time, inaccurate bacteria count, and unclear genotype characterizations. The main goals of this proposed project are: 1) Develop a Field-Deployable Droplet Digital Polymerase Chain Reaction (ddPCR) system that can rapidly detect pathogenic Escherichia coli (E. coli) in the field. 2) Use this proposed system to test the water samples of Animas River in Colorado. 3) Educate communities (including the Native American tribes) in the four-corner area on the importance of water resource protection and the technologies for environmental protection.
Summary/Accomplishments (Outputs/Outcomes):
The main goal of this project is to develop a field-deployable Droplet Digital Polymerase Chain Reaction (ddPCR) system that can rapidly detect pathogenic Escherichia coli (E. coli) in the field. There are four technical objectives to be progressively completed during the funding cycle. The tasks include the ddPCR assay development, microfluidic device design and fabrication, design and test a customizable PCR thermocycler, system integration, and field tests. The team has received strong institutional and external support since this project was granted and by leveraging these supports the team expanded to 9 students and 3 faculty mentors (volunteered) in phase 1. Six of the students were from a senior design team who have graduated in April 2020 from the college. One student from the biology department was recruited in the summer of 2020 to train our engineering students on genetic engineering and DNA sequencing for bacterial samples. In addition to the funding provided by this award, the team has raised an extra $6,000 for lab materials and supplies, $9,711 for a VWR -80 ℃ freezer to permanently store the environmental bacterial samples, and $23,999 for a Raman spectrometer. The team has made significant progress toward the objectives of the proposal and contributed 2 peer-reviewed IEEE conference articles, 7 poster/oral presentations. The team developed a functioning PCR system and received good results of identifying environmental E coli in ddPCR trials. A droplet counting system is being developed by a senior seminar team in the 2021-2022 academic year. Integrate the developed PCR system and the droplet counting system will enable fast bacteria identification in the field for environmental researchers.
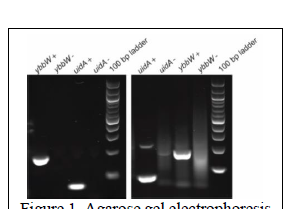
Objective 1: Development of the PCR assay to detect E. coli in microfluidic droplets. EPA method 1604 states that 100mL is the appropriate volume for surface water testing. However, a sample of this size would require an unrealistic amount of time and cost to process, so it must be concentrated. A back-flushing membrane filtration process is proposed to concentrate samples. Membrane filtration is characterized by the separation of particles via a porous membrane filter that allows fluid to flow through while collecting relatively larger particles on the top of the filter (EPA Method 1103.1 is observed in this study). We used two Nalgene™ reusable filter units (Thermo Fisher Scientific) and a mixed cellulose esters (MCE) membrane (MF-Millipore) of 47mm diameter and 0.45µm pore size. Filtration units are fully sterilized following the EPA 1604 testing procedure. The effectiveness of the PCR assay was verified in traditional PCR and tested in droplets. Figure 1 shows the PCR gel images of the products from both the Bio-Rad PCR and our FLC-PCR thermocyclers (developed during this funding cycle). In both thermocyclers, amplification of ybbW yielded a specific product of 177bp and amplification of uidA yielded a specific product of 70 bp. Smeared areas were observed in the gel that carries our FLC-PCR products. This may be attributed to the poor heat transfer rate from the heating block to the PCR tubes. Calibrations of the liquid temperature inside the tubes and the body of the tubes were performed using water-proof thermocouples to improve the performance of our FLC-PCR machine.
Figure 1. Agarose gel electrophoresis analysis of ybbW and uidA PCR products generated in a Bio-Rad T100 thermocyler (left panel) and our system (right panel). PCR reactions containing template are noted (+), no template controls are noted (-). Bright band in 100bp ladder is 500bp long.
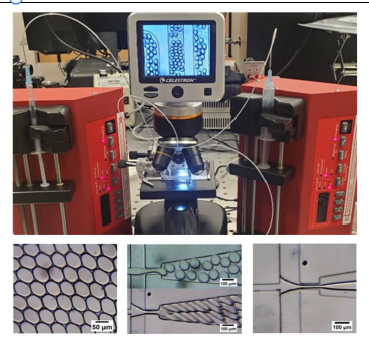
Objective 2: Design of the high-throughput microfluidic device. We have established the work flow for the batch fabrication of our microfluidic devices. A microfluidic device was necessary to discretize and emulsify the concentrated water sample into microscopic droplets. Figure 2 (top) shows the microscope-pump-microfluidics setup for droplet generation. Figure 2 (bottom left) shows droplets emulsified in oil, Figure 2 (bottom middle) shows droplet creation at various oil flow rates, and Figure 2 (bottom right) shows a T-junction experiencing chip failure due to wetting. The wetting issue can be solved by setting the microfluidic chip in a 65 ℃ oven for 3-7 days before use. Objective 3. Development of a portable PCR machine.
Figure 2. The pump-syringe-microfluidics-microscope setup for droplet generation (Top). Examples of droplet creation at the device T-junction Figure(droplets). (bottom-left) cell counting chamber slide. (bottom-middle) T-Junction droplet production. Channel wetting (bottom-right).
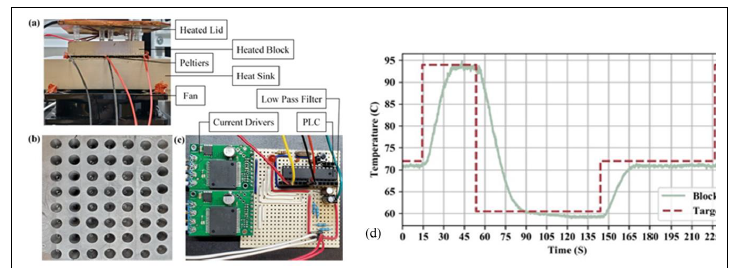
A thermocycler was designed and fabricated to replicate the target amplicon and amplify fluorescence released from droplets containing E. coli during the PCR process. Figure 3 (a-c) shows a diagram of the fabricated thermocycler. A peer-
reviewed conference paper that articulates the design was published in IEEE EMBC 2020. (James Ferguson, Jesse Duran, Wesley Killinen, Jason Wagner, Caroline Kulesza, Christie Chatterley, and Yiyan Li. Rapid Detection of E. Coli Using a Field Deployable and Low-Cost PCR (FLC-PCR) Thermocycler).
Figure 3. (a-c) The key components of the FLC-PCR prototype. (d) Thermocycler testing results.
Figure 3(d) shows the results of the thermocycler during operation. The dotted red line shows the target temperature of the device and the solid green block shows the actual temperature of the device over time. The user interface is an important part of any computer-based system that humans are expected to interact with. Graphical User Interfaces (GUI) are generally considered to be the best way to handle this type of interaction currently. The GUI for the FLC-PCR was written in C++ with SDL2. SDL2 provides access to a low-level rendering, user input, and window handling. While SDL2 requires more work than QT or TK to develop a GUI. SDL2 enables much more customization.
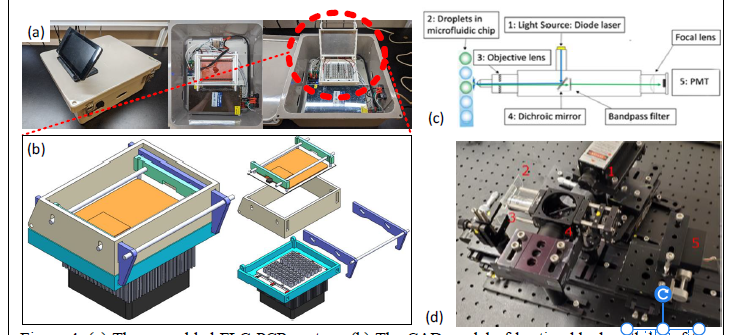
Objective 4 (phase I). System integration and automation. An enclosure was modeled in SOLIDWORKS that consists of four moving parts as shown in Figure 4. The cyan part is the base of the system containing the peltiers, heatsink, heating block and PCR tubes. The beige part forms the walls of the lid which is attached to the base by a hinge at the bottom. The orange and green part is the top of the lid, In the center is a model 16 engine preheater attached to a 2mm aluminum plate that prevents the samples from condensing at the top of a PCR tube. This plate is attached to the green brackets via springs that apply about 5lbs of pressure to the top of the tubes when closed, keeping the tubes sealed during a PCR experiment. The construction consists of a MBL-III-473-50mW laser, a LPlan 60x objective lens, 50mm Plano convex lens, 529/4 nm bandpass filter, Hamamatsu H9307 photomultiplier tube (PMT), a Thor Labs D= 25.4mm F = 75mm (AC245-075-A) achromatic lens, and a ZT405 dichroic mirror (Figure 4 (c-d)).
Figure 4: (a) The assembled FLC-PCR system. (b) The CAD model of heating block with lid of the
FLC-PCR machine. (c-d) Florescent microscope diagram and implementation.
Conclusions:
The team has made significant progress in the development of the field-deployable bacteria detection system and will be ready for massive field test in 2022. In addition to the funds requested from EPA, the team has raised an extra $39,710 to refresh lab equipment and purchase lab consumables. Four engineering students at FLC were hired to the team and a 6-student senior seminar team was involved for the first year of the project. Three faculty members served as mentors to these students and supervised the delivery of 2 publications and 7 presentations as of December 2021. The team developed a functioning PCR system and assay to identify environmental E coli samples on a gel, in an rtPCR machine, and in microfluidic droplets (Figure 5).
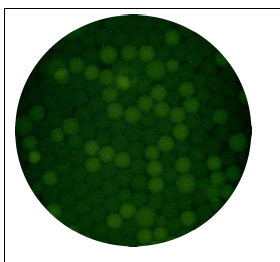
For water sampling and preparation, we found that the Elution fluid to be the most reliable and cost-effective method for concentering 100 mL of E. coli spiked water into volumes of approximately 0.1 mL. Our fabricated microfluidic device was able to create uniform droplets of the appropriate size (approximately 50 microns). Our optical detection setup was able to detect fluorescence down to 1:100,000 concentrations. False readings were found occasionally from the data acquisition system. In the next phase of this project, the entire system will be tested and timed in series, eventually using raw water samples during testing. After these tests have been completed, a field-deployable prototype will be manufactured.
Figure 5. A fluorescence microscopic image of a successful ddPCR trial for the detection of cultured environmental E. coli. Imaged at 200x.
Journal Articles:
No journal articles submitted with this report: View all 4 publications for this projectSupplemental Keywords:
Rapid, Bacteria Detection, ddPCR, Single CellProgress and Final Reports:
Original AbstractThe perspectives, information and conclusions conveyed in research project abstracts, progress reports, final reports, journal abstracts and journal publications convey the viewpoints of the principal investigator and may not represent the views and policies of ORD and EPA. Conclusions drawn by the principal investigators have not been reviewed by the Agency.